- Last 7 days
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Hall et al describe the superiority of ONT sequencing and deep learning-based variant callers to deliver higher SNP and Indel accuracy compared to previous gold-standard Illumina short-read sequencing. Furthermore, they provide recommendations for read sequencing depth and computational requirements when performing variant calling.
Strengths:
The study describes compelling data showing ONT superiority when using deep learning-based variant callers, such as Clair3, compared to Illumina sequencing. This challenges the paradigm that Illumina sequencing is the gold standard for variant calling in bacterial genomes. The authors provide evidence that homopolymeric regions, a systematic and problematic issue with ONT data, are no longer a concern in ONT sequencing.
Weaknesses:
(1) The inclusion of a larger number of reference genomes would have strengthened the study to accommodate larger variability (a limitation mentioned by the authors).
(2) In Figure 2, there are clearly one or two samples that perform worse than others in all combinations (are always below the box plots). No information about species-specific variant calls is provided by the authors but one would like to know if those are recurrently associated with one or two species. Species-specific recommendations could also help the scientific community to choose the best sequencing/variant calling approaches.
(3) The authors support that a read depth of 10x is sufficient to achieve variant calls that match or exceed Illumina sequencing. However, the standard here should be the optimal discriminatory power for clinical and public health utility (namely outbreak analysis). In such scenarios, the highest discriminatory power is always desirable and as such an F1 score, Recall and Precision that is as close to 100% as possible should be maintained (which changes the minimum read sequencing depth to at least 25x, which is the inflection point).
(4) The sequencing of the samples was not performed with the same Illumina and ONT method/equipment, which could have introduced specific equipment/preparation artefacts that were not considered in the study. See for example https://academic.oup.com/nargab/article/3/1/lqab019/6193612.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This is a tour de force study that aims to understand the genetic basis of male germ cell development across three animal species (human, mouse, and flies) by performing a genetic program conservation analysis (using phylostratigraphy and network science) with a special emphasis on genes that peak or decline during mitosis-to-meiosis. This analysis, in agreement with previous findings, reveals that several genes active during and before meiosis are deeply conserved across species, suggesting ancient regulatory mechanisms. To identify critical genes in germ cell development, the investigators integrated clinical genetics data, performing gene knockdown and knockout experiments in both mice and flies. Specifically, over 900 conserved genes were investigated in flies, with three of these genes further studied in mice. Of the 900 genes in flies, ~250 RNAi knockdowns had fertility phenotypes. The fertility phenotypes for the fly data can be viewed using the following browser link: https://pages.igc.pt/meionav. The scope of target gene validation is impressive. Below are a few minor comments.
(1) In Supplemental Figure 2, it is notable that enterocyte transcriptomes are predominantly composed of younger genes, contrasting with the genetic age profile observed in brain and muscle cells. This difference is an intriguing observation and it would be curious to hear the author's comments.
(2) Regarding the document, the figures provided only include supplemental data; none of the main text figures are in the full PDF.
(3) Lastly, it would be great to section and stain mouse testis to classify the different stages of arrest during meiosis for each of the mouse mutants in order to compare more precisely to flies.
This paper serves as a vital resource, emphasizing that only through the analysis of hundreds of genes can we prioritize essential genes for germ cell development. its remarkable that about 60% of conserved genes have no apparent phenotype during germ cell development.
Strengths:
The high-throughput screening was conducted on a conserved network of 920 genes expressed during the mitosis-to-meiosis transition. Approximately 250 of these genes were associated with fertility phenotypes. Notably, mutations in 5 of the 250 genes have been identified in human male infertility patients. Furthermore, 3 of these genes were modeled in mice, where they were also linked to infertility. This study establishes a crucial groundwork for future investigations into germ cell development genes, aiming to delineate their essential roles and functions.
Weaknesses:
The fertility phenotyping in this study is limited, yet dissecting the mechanistic roles of these proteins falls beyond its scope. Nevertheless, this work serves as an invaluable resource for further exploration of specific genes of interest.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
A large number of ovarian experiments have been conducted - especially in morphological and molecular biology studies - specifically removing the ovarian membrane. This experiment is a good supplement to existing knowledge and plays an important role in early ovarian development and the regulation of ovarian homeostasis during the estrous cycle. There are also innovations in research ideas and methods, which will meet the requirements of experimental design and provide inspiration for other researchers.
This reviewer did not identify any major issues with the article. However, the following points could be further clarified:
(1) Is there any comparative data on the proteomics of RO and rete testis in early development? With some molecular markers also derived from rete testis, it would be better to provide the data or references.
(2) Although the size of RO and its components is quite small and difficult to operate, the researchers in this article had already been able to perform intracavitary injection of EOR and extract EOR or CR for mass spectrometry analysis. Therefore, can EOR, CR, or IOR be damaged or removed, providing further strong evidence of ovarian development function?
(3) Although IOR is shown on the schematic diagram, it cannot be observed in the immunohistochemistry pictures in Figure 1 and Figure 3. The authors should provide a detailed explanation.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript described the second earliest known winged ovule without a capule in the Famennian of Late Devonian. Using Mathematical analysis, the authors suggest that the integuments of the earliest ovules without a cupule, as in the new taxon and Guazia, evolved functions in wind dispersal.
Strengths:
The new ovule taxon's morphological part is convincing. It provides additional evidence for the earliest winged ovules, and the mathematical analysis helps to understand their function.
Weaknesses:
The discussion should be enhanced to clarify the significance of this finding. What is the new advance compared with the Guazia finding? The authors can illustrate the character transformations using a simplified cladogram. The present version of the main text looks flat.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This paper develops an under-flow migration tracker to evaluate all the steps of the extravasation cascade of immune cells across the BBB. The algorithm is useful and has important applications.
Strengths:
Algorithm is almost as accurate as manual tracking and importantly saves time for researchers.
Weaknesses:
Applicability can be questioned because the device used is 2D and physiological biology is in 3D. Comparisons to other automated tools was not performed by the authors.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The document "Mapping spatial patterns to energetic benefits in groups of flow-coupled swimmers" by Heydari et al. uses several types of simulations and models to address aspects of stability of position and power consumption in few-body groups of pitching foils. I think the work has the potential to be a valuable and timely contribution to an important subject area. The supporting evidence is largely quite convincing, though some details could raise questions, and there is room for improvement in the presentation. My recommendations are focused on clarifying the presentation and perhaps spurring the authors to assess additional aspects:
(1) Why do the authors choose to set the swimmers free only in the propulsion direction? I can understand constraining all the positions/orientations for investigating the resulting forces and power, and I can also understand the value of allowing the bodies to be fully free in x, y, and their orientation angle to see if possible configurations spontaneously emerge from the flow interactions. But why constrain some degrees of freedom and not others? What's the motivation, and what's the relevance to animals, which are fully free?
(2) The model description in Eq. (1) and the surrounding text is confusing. Aren't the authors computing forces via CFD or the VS method and then simply driving the propulsive dynamics according to the net horizontal force? It seems then irrelevant to decompose things into thrust and drag, and it seems irrelevant to claim that the thrust comes from pressure and the drag from viscous effects. The latter claim may in fact be incorrect since the body has a shape and the normal and tangential components of the surface stress along the body may be complex.
(3) The parameter taudiss in the VS simulations takes on unusual values such as 2.45T, making it seem like this value is somehow very special, and perhaps 2.44 or 2.46 would lead to significantly different results. If the value is special, the authors should discuss and assess it. Otherwise, I recommend picking a round value, like 2 or 3, which would avoid distraction.
(4) Some of the COT plots/information were difficult to interpret because the correspondence of beneficial with the mathematical sign was changing. For example, DeltaCOT as introduced on p. 5 is such that negative indicates bad energetics as compared to a solo swimmer. But elsewhere, lower or more negative COT is good in terms of savings. Given the many plots, large amounts of data, and many quantities being assessed, the paper needs a highly uniform presentation to aid the reader.
(5) I didn't understand the value of the "flow agreement parameter," and I didn't understand the authors' interpretation of its significance. Firstly, it would help if this and all other quantities were given explicit definitions as complete equations (including normalization). As I understand it, the quantity indicates the match of the flow velocity at some location with the flapping velocity of a "ghost swimmer" at that location. This does not seem to be exactly relevant to the equilibrium locations. In particular, if the match were perfect, then the swimmer would generate no relative flow and thus no thrust, meaning such a location could not be an equilibrium. So, some degree of mismatch seems necessary. I believe such a mismatch is indeed present, but the plots such as those in Figure 4 may disguise the effect. The color bar is saturated to the point of essentially being three tones (blue, white, red), so we cannot see that the observed equilibria are likely between the max and min values of this parameter.
(6) More generally, and related to the above, I am favorable towards the authors' attempts to find approximate flow metrics that could be used to predict the equilibrium positions and their stability, but I think the reasoning needs to be more solid. It seems the authors are seeking a parameter that can indicate equilibrium and another that can indicate stability. Can they clearly lay out the motivation behind any proposed metrics, and clearly present complete equations for their definitions? Further, is there a related power metric that can be appropriately defined and which proves to be useful?
(7) Why do the authors not carry out CFD simulations on the larger groups? Some explanations should be given, or some corresponding CFD simulations should be carried out. It would be interesting if CFD simulations were done and included, especially for the in-line case of many swimmers. This is because the results seem to be quite nuanced and dependent on many-body effects beyond nearest-neighbor interactions. It would certainly be comforting to see something similar happen in CFD.
(8) Related to the above, the authors should discuss seemingly significant differences in their results for long in-line formations as compared to the CFD work of Peng et al. [48]. That work showed apparently stable groups for numbers of swimmers quite larger than that studied here. Why such a qualitatively different result, and how should we interpret these differences regarding the more general issue of the stability of tandem groups?
(9) The authors seem to have all the tools needed to address the general question about how dynamically stable configurations relate to those that are energetically optimal. Are stable solutions optimal, or not? This would seem to have very important implications for animal groups, and the work addresses closely related topics but seems to miss the opportunity to give a definitive answer to this big question.
(10) Time-delay particle model: This model seems to construct a simplified wake flow. But does the constructed flow satisfy basic properties that we demand of any flow, such as being divergence-free? If not, then the formulation may be troublesome.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Summary:
The receptor tyrosine kinase Anaplastic Lymphoma Kinase (ALK) in humans is nervous system expressed and plays an important role as an oncogene. A number of groups have been studying ALK signalling in flies to gain mechanistic insight into its various roles. In flies, ALK plays a critical role in development, particularly embryonic development and axon targeting. In addition, ALK was also shown to regulate adult functions including sleep and memory. In this manuscript, Sukumar et al., used a suite of molecular techniques to identify downstream targets of ALK signalling. They first used targeted DamID, a technique that involves a DNA methylase to RNA polymerase II, so that GATC sites in close proximity to PolII binding sites are marked. They performed these experiments in wild type and ALK loss of function mutants (using an Alk dominant negative ALkDN), to identify Alk responsive loci. Comparing these loci with a larval single cell RNAseq dataset identified neuroendocrine cells as an important site of Alk action. They further combined these TaDa hits with data from RNA seq in Alk Loss and Gain of Function manipulations to identify a single novel target of Alk signalling - a neuropeptide precursor they named Sparkly (Spar) for its expression pattern. They generated a mutant allele of Spar, raised an antibody against Spar, and characterised its expression pattern and mutant behavioural phenotypes including defects in sleep and circadian function.
Strengths:
The molecular biology experiments using TaDa and RNAseq were elegant and very convincing. The authors identified a novel gene they named Spar. They also generated a mutant allele of Spar (using CrisprCas technology) and raised an antibody against Spar. These experiments are lovely, and the reagents will be useful to the community. The paper is also well written, and the figures are very nicely laid out making the manuscript a pleasure to read.
Weaknesses:
The manuscript has improved very substantially in revision. The authors have clearly taken the comments on board in good faith.
Editors' note: The authors have satisfactorily addressed the concerns raised in the previous rounds of review. These were related to the unconventional analysis of the TaDa data, the addition of other means of down regulated gene function, and the nature of analyses of behavioural data.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Chang et al. investigated neuronal activity firing patterns across various cortical regions in an interesting context-dependent tactile vs visual detection task, developed previously by the authors (Chevee et al., 2021; doi: 10.1016/j.neuron.2021.11.013). The authors report the important involvement of a medial frontal cortical region (MM, probably a similar location to wM2 as described in Esmaeili et al., 2021 & 2022; doi: 10.1016/j.neuron.2021.05.005; doi: 10.1371/journal.pbio.3001667) in mice for determining task rules.
Strengths:
The experiments appear to have been well carried out and the data well analysed. The manuscript clearly describes the motivation for the analyses and reaches clear and well-justified conclusions. I find the manuscript interesting and exciting!
Weaknesses:
I did not find any major weaknesses.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
This article is the first report to study the effects of T. pallidum on the neural development of an iSPC-derived brain organoid model. The study indicates that T. pallidum inhibits the differentiation of subNPC1B neurons into hindbrain neurons, hence affecting brain organoid neurodevelopment. Additionally, the TCF3 and notch signaling pathways may be involved in the inhibition of the subNPC1B-hindbrain neuron differentiation axis. While the majority of the data in this study support the conclusions, there are still some questions that need to be addressed and data quality needs to be improved. The study provides valuable insights for future investigations into the mechanisms underlying congenital neurodevelopment disability.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors study through theory and simulations the diffusion of microscopic particles, and aim to account for the effects of inhomogeneous viscosity and diffusion - in particular regarding the intracellular environment. They propose a mechanism, termed "Diffusive lensing", by which particles are attracted towards low-diffusivity regions where they remain trapped. To obtain these results, the authors rely on agent-based simulations using custom rules performed within the Ito stochastic calculus convention, without drift. They acknowledge the fact that this convention does not describe equilibrium systems, and that their results would not hold at equilibrium - and discard these facts by invoking the facts that cells are out-of-equilibrium. Finally, they show some applications of their findings, in particular enhanced clustering in the low-diffusivity regions. The authors conclude that as inhomogeneous diffusion is ubiquitous in life, so must their mechanism be, and hence it must be important.
Strengths:
The article is well-written, clearly intelligible, its hypotheses are stated relatively clearly and the models and mathematical derivations are compatible with these hypotheses. In the appendices, the authors connect their findings to known results for classic stochastic differential equation formalisms.
Weaknesses:
This study is, in my opinion, deeply flawed. The main problem lies in the hypotheses, in particular the choice of considering drift-less dynamics in the Ito convention. It is regrettable that the authors choose to use agent-based custom simulations with little physical motivation, rather than a well-established stochastic differential equations framework.
Indeed, stochastic conventions are a notoriously tricky business, but they are both mathematically and physically well-understood and do not result in any "dilemma" [some citations in the article, such as (Lau and Lubensky) and (Volpe and Wehr), make an unambiguous resolution of these]. In the continuous-time limit, conventions are not an intrinsic, fixed property of a system, but a choice of writing; however, whenever going from one to another, one must include a corresponding "spurious drift" that compensates the effect of this change - a mathematical subtlety that is omitted in the article (except in a quick note in the appendix): in the presence of diffusive gradients, if the drift is zero in one convention, it will thus be non-zero in another. It is well established that for equilibrium systems obeying fluctuation-dissipation, the spurious drift vanishes in the anti-Ito stochastic convention; more precisely one can write in the anti-Ito convention
dx/dt = - D(x)/kT grad U(x) + sqrt(2D(x)) dW
with D(x) the diffusion, kT the thermal energy (which is space-independent at equilibrium), and dW a d-dimensional Wiener process. Equivalently one can write in the Ito convention:
dx/dt = - D(x)/kT grad U(x) + sqrt(2D(x)) dW + div D(x) (*)
where the latter term is the spurious drift arising from convention change. This ensures that the diffusion gradients do not induce currents and probability gradients, and thus that the steady-state PDF is the Gibbs measure (this form has been confirmed experimentally, for instance, for colloidal particles near walls, that have strong diffusivity gradients despite not having significant forces). It generalizes to near-equilibrium systems with non-conservative forces and/or temperature gradient in the form:
dx/dt = F(x) + sqrt(2D(x)) dW + div D(x) (**)
where the drift field F(x) encodes these forces. In some cases, it has been shown through careful microscopic analysis that one can have effectively a different form for the last term, namely
dx/dt = F(x) + sqrt(2D(x)) dW + alpha div D(x)
where alpha is a "convention parameter" that would be =1 at equilibrium. For instance, in the Volpe and Wehr review this can occur through memory effects in robotic dynamics, or through strong fluctuation-dissipation breakdown. In a near-equilibrium system, this should be strongly justified, as the continuous-time dynamics with alpha \neq 1 and drift F would be indistinguishable from one with alpha = 1 and drift F + (1-alpha) div D: the authors would have the burden of proving that the observed (absence of) drift is indeed due to alpha\neq 1, rather than to much more common force fields F(x).
Here, without further motivation than the statement that cells are out-of-equilibrium, drifts are arbitrarily set to zero in the Ito convention, which is in (**) the equivalent to adding a force with drift $-div D$ exactly compensating the spurious drift. It is the effects of this arbitrary force that are studied in the article. The fact that it results in probability gradients is trivial once formulated this way (and in no way is this new - many of the references, for instance Volpe and Wehr, mention this). Enhanced clustering is also a trivial effect of this probability gradient (the local concentration is increased by this force field, so phase separation can occur). As a side note the "neighbor sensing" scheme to describe interactions is itself very peculiar and not physically motivated - it violates stochastic thermodynamics laws too, as detailed balance is apparently not respected. There again, the authors have chosen to disregard a century of stochastic thermodynamics in favor of a non-justified unphysical custom rule.
The authors make no further justification of their choice of driftless Ito simulations than the fact that cells are out-of-equilibrium, leaving the feeling that this is a detail. They make mentions of systems (eg glycogen, prebiotic environment) for which (near-)equilibrium physics should mostly prevail, and of fluctuation dissipation ("Diffusivity varies inversely with viscosity", in the introduction). Yet the "phenomenon" they discuss is entirely reliant on an undiscussed mechanism by which these assumptions would be completely violated (the citations they make for this - Gnesotto '18 and Phillips '12 - are simply discussions of the fact that cells are out-of-equilibrium, not on any consequences on the convention).
Finally, while inhomogeneous diffusion is ubiquitous, the strength of this effect in realistic conditions is not discussed. Even in the most "optimistic" case where alpha=0 would make sense (knowing that in the cellular context we are discussing thermal systems immersed in water and if energy consumption and metabolism were stopped alpha would relax back to 1), the equation (*) above shows that having zero ito drift is equivalent to having a potential countering the spurious drift, with value
U(x) = kT log(D(x) / D0 )
[I have assumed isotropic diffusion for simplicity here, so the div is replaced by a grad]. This means that the diffusion contrasts logarithmically compare to the chemical potential ones -- for instance a major diffusion difference of 100x is equivalent to 4.6kT in potential energy, a relatively modest effect. To prove that the authors' effect of "diffusive lensing" is involved in such a system, one would thus have to<br /> 1) observe strong spatial variations of the diffusion coefficient (this is doable, and was done before), AND<br /> 2) show that there is an enrichment of the diffusing species in the low-diffusion region inversely proportional to the diffusion, AND<br /> 3) show that this enrichment cannot be attributed to mild differences in potential energy, for instance by showing that if nonequilibrium energy consumption stops, the concentration fully homogenizes while the diffusion gradients remain.
If the authors were to successfully show all that in an experimental system, or design a theoretical framework where these effects convincingly emerge from physically realistic microscopic dynamical rules, they would have indeed discovered a new phenomenon. In contrast, the current article only demonstrates the well-known fact that when using arbitrary dynamical rules in heterogeneous diffusion simulations, one can get concentration gradients.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This work aggregates data across 5 openly available stopping studies (3 at 7 tesla and 2 at 3 tesla) to evaluate activity patterns across the common contrasts of Failed Stop (FS) > Go, FS > stop success (SS), and SS > Go. Previous work has implicated a set of regions that tend to be positively active in one or more of these contrasts, including the bilateral inferior frontal gyrus, preSMA, and multiple basal ganglia structures. However, the authors argue that upon closer examination, many previous papers have not found subcortical structures to be more active on SS than FS trials, bringing into question whether they play an essential role in (successful) inhibition. In order to evaluate this with more data and power, the authors aggregate across five datasets and find many areas that are *more* active for FS than SS, including bilateral preSMA, GPE, thalamus, and VTA. They argue that this brings into question the role of these areas in inhibition, based upon the assumption that areas involved in inhibition should be more active on successful stop than failed stop trials, not the opposite as they observed.
Since the initial submission, the authors have improved their theoretical synthesis and changed their SSRT calculation method to the more appropriate integration method with replacement for go omissions. They have also done a better job of explaining how these fMRI results situate within the broader response inhibition literature including work using other neuroscience methods.
They have also included a new Bayes Factor analysis. In the process of evaluating this new analysis, I recognized the following comments that I believe justify additional analyses and discussion:
First, if I understand the author's pipeline, for the ROI analyses it is not appropriate to run FSL's FILM method on the data that were generated by repeating the same time series across all voxels of an ROI. FSL's FILM uses neighboring voxels in parts of the estimation to stabilize temporal correlation and variance estimates and was intended and evaluated for use on voxelwise data. Instead, I believe it would be more appropriate to average the level 1 contrast estimates over the voxels of each ROI to serve as the dependent variables in the ROI analysis.
Second, for the group-level ROI analyses there seems to be inconsistencies when comparing the z-statistics (Figure 3) to the Bayes Factors (Figure 4) in that very similar z-statistics have very different Bayes Factors within the same contrast across different brain areas, which seemed surprising (e.g., a z of 6.64 has a BF of .858 while another with a z of 6.76 has a BF of 3.18). The authors do briefly discuss some instances in the frequentist and Bayesian results differ, but they do not ever explain by similar z-stats yield very different bayes factors for a given contrast across different brain areas. I believe a discussion of this would be useful.
Third, since the Bayes Factor analysis appears to be based on repeated measures ANOVA and the z-statistics are from Flame1+2, the BayesFactor analysis model does not pair with the frequentist analysis model very cleanly. To facilitate comparison, I would recommend that the same repeated measures ANOVA model should be used in both cases. My reading of the literature is that there is no need to be concerned about any benefits of using Flame being lost, since heteroscedasticity does not impact type I errors and will only potentially impact power (Mumford & Nichols, 2009 NeuroImage).
Fourth, though frequentist statistics suggest that many basal ganglia structures are significantly more active in the FS > SS contrast (see 2nd row of Figure 3), the Bayesian analyses are much more equivocal, with no basal ganglia areas showing Log10BF > 1 (which would be indicative of strong evidence). The authors suggest that "the frequentist and Bayesian analyses are monst in line with one another", but in my view, this frequentist vs. Bayesian analysis for the FS > SS contrast seems to suggest substantially different conclusions. More specifically, the frequentist analyses suggest greater activity in FS than SS in most basal ganglia ROIs (all but 2), but the Bayesian analysis did not find *any* basal ganglia ROIs with strong evidence for the alternative hypothesis (or a difference), and several with more evidence for the null than the alternative hypothesis. This difference between the frequentist and Bayesian analyses seems to warrant discussion, but unless I overlooked it, the Bayesian analyses are not mentioned in the Discussion at all. In my view, the frequentist analyses are treated as the results, and the Bayesian analyses were largely ignored.
Overall, I think this paper makes a useful and mostly solid contribution to the literature. I have made some suggestions for adjustments and clarification of the neuroimaging pipeline and Bayesian analyses that I believe would strengthen the work further.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This work clarifies neural mechanisms that can lead to a phenomenology consistent with motor preparation in its broader sense. In this context, motor preparation refers to activity that occurs before the corresponding movement. Another property often associated with preparatory activity is a correlation with global movement characteristics such as reach speed (Churchland et al., Neuron 2006), reach angle (Sun et al., Nature 2022), or grasp type (Meirhaeghe et al., Cell Reports 2023). Such activity has notably been observed in premotor and primary motor cortices, and it has been hypothesized to serve as an input to a motor execution circuit. The timing and mechanisms by which such 'preparatory' inputs are made available to motor execution circuits remain however unclear in general, especially in light of the presence of a 'trigger-like' signal that appears to relate to the transition from preparatory dynamics to execution activity (Kaufman et al. eNeuron 2016, Iganaki et al., Cell 2022, Zimnik and Churchland, Nature Neuroscience 2021).
The preparatory inputs have been hypothesized to fulfill one or several (non-mutually-exclusive) possible objectives. Two notable hypotheses are that these inputs could be shaped to maximize output accuracy under regularization of the input magnitude; or that they may help the flexible re-use of the neural machinery involved in the control of movements in different contexts.
Here, the authors investigate in detail how the former hypothesis may be compatible with the presence of early inputs in recurrent network models driving arm movements, and compare models to data.
Strengths:
The authors are able to deploy an in-depth evaluation of inputs that are optimized for producing an accurate output at a pre-defined time while using a regularization term on the input magnitude, in the case of movements that are thought to be controlled in a quasi-open loop fashion such as reaches.
First, the authors have identified that optimal control theory is a great framework to study this question as it provides methods to find and analyze exact solutions to this cost function in the case of models with linear dynamics. The authors not only use this framework to get an exact assessment of how much pre-movement input arises in large recurrent networks, but also give insight into the mechanisms by which it happens by dissecting in detail low-dimensional networks. The authors find that two key network properties - observability of the readout's nullspace and limited controllability - give rise to optimal inputs that are large before the start of the movement (while the corresponding network activity lies in the nullspace of the readout). Further, the authors numerically investigate the timing of optimized inputs in models with nonlinear dynamics, and find that pre-movement inputs can also arise in these more general networks. The authors also explore how some variations on their model's constraints - such as penalizing the input roughness or changing task contingencies about the go cue timing - affect their results. Finally, the authors point out some coarse-grained similarities between the pre-movement activity driven by the optimized inputs in some of the models they studied, and the phenomenology of preparation observed in the brain during single reaches and reach sequences. Overall, the authors deploy an impressive arsenal of tools and a very in-depth analysis of their models.
Limitations:
(1) Though the optimal control theory framework is ideal to determine inputs that minimize output error while regularizing the input norm or other simple input features, it cannot easily account for some other varied types of objectives - especially those that may lead to a complex optimization landscape. For instance, the reusability of parts of the circuit, sparse use of additional neurons when learning many movements, and ease of planning (especially under uncertainty about when to start the movement), may be alternative or additional reasons that could help explain the preparatory activity observed in the brain. It is interesting to note that inputs that optimize the objective chosen by the authors arguably lead to a trade-off in terms of other desirable objectives. Specifically, the inputs the authors derive are time-dependent, so a recurrent network would be needed to produce them and it may not be easy to interpolate between them to drive new movement variants. In addition, these inputs depend on the desired time of output and therefore make it difficult to plan, e.g. in circumstances when timing should be decided depending on sensory signals. Finally, these inputs are specific to the full movement chain that will unfold, so they do not permit reuse of the inputs e.g. in movement sequences of different orders. Of note, the authors have pointed out in the discussion how their framework may be extended in future work to account for some additional objectives, such as inputs' temporal smoothness or some strategies for dealing with go cue timing uncertainty.
(2) Relatedly, if the motor circuits were to balance different types of objectives, the activity and inputs occurring before each movement may be broken down into different categories that may each specialize into their own objective. For instance, previous work (Kaufman et al. eNeuron 2016, Iganaki et al., Cell 2022, Zimnik and Churchland, Nature Neuroscience 2021) has suggested that inputs occurring before the movement could be broken down into preparatory inputs 'stricto sensu' - relating to the planned characteristics of the movement - and a trigger signal, relating to the transition from planning to execution - irrespective of whether the movement is internally timed or triggered by an external event. The current work does not address which type(s) of early input may be labeled as 'preparatory' or may be thought of as a part of 'planning' computations, or whether these inputs may come from several different source circuits.
(3) While the authors rightly point out some similarities between the inputs that they derive and observed preparatory activity in the brain, notably during motor sequences, there are also some differences. For instance, while both the derived inputs and the data show two peaks during sequences, the data reproduced from Zimnik and Churchland show preparatory inputs that have a very asymmetric shape that really plummets before the start of the next movement, whereas the derived inputs have larger amplitude during the movement period - especially for the second movement of the sequence. In addition, the data show trigger-like signals before each of the two reaches. Finally, while the data show a very high correlation between the pattern of preparatory activity of the second reach in the double reach and compound reach conditions, the derived inputs appear to be more different between the two conditions. Note that the data would be consistent with separate planning of the two reaches even in the compound reach condition, as well as the re-use of the preparatory input between the compound and double reach conditions. Therefore, different motor sequence datasets - notably, those that would show even more coarticulation between submovements - may be more promising to find a tight match between the data and the author's inputs. Further analyses in these datasets could help determine whether the coarticulation could be due to simple filtering by the circuits and muscles downstream of M1, planning of movements with adjusted curvature to mitigate the work performed by the muscles while permitting some amount of re-use across different sequences, or - as suggested by the authors - inputs fully tailored to one specific movement sequence that maximize accuracy and minimize the M1 input magnitude.
(4) Though iLQR is a powerful optimization method to find inputs optimizing the author's cost function, it also has some limitations. First, given that it relies on a linearization of the dynamics at each timestep, it has a limited ability to leverage potential advantages of nonlinearities in the dynamics. Second, the iLQR algorithm is not a biologically plausible learning rule and therefore it might be difficult for the brain to learn to produce the inputs that it finds. Therefore, when observing differences between model and data, this can confound the question of whether it comes from a difference of assumed objective or a difference of optimization procedure. It remains unclear whether using alternative algorithms with different limitations - for instance, using variants of BPTT to train a separate RNN to produce the inputs in question - could impact some of the results.
(5) Under the objective considered by the authors, the amount of input occurring before the movement might be impacted by the presence of online sensory signals for closed-loop control. Even if considering that the inputs could include some sensory activity and/or that the RNN activity could represent general variables whose states can be decoded from M1, the model would not include mechanisms that process imperfect (delayed, noisy) sensory feedback to adapt the output in a trial-specific manner. It is therefore an open question whether the objective and network characteristics suggested by the authors could also explain the presence of preparatory activity before e.g. grasping movements that are thought to be more sensory-driven (Meirhaeghe et al., Cell Reports 2023).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
In this work, Dasgupta et al. investigate the role of Sema7a in the formation of peripheral sensory circuit in the lateral line system of zebrafish. They show that Sema7a protein is present during neuromast maturation and localized, in part, to the base of hair cells (HCs). This would be consistent with pre-synaptic Sema7a mediating formation and/or stabilization of the synapse. They use sema7a loss-of-function strain to show that lateral line sensory terminals display abnormal arborization. They provide highly quantitative analysis of the lateral line terminal arborization to show that a number of specific topological parameters are affected in mutants. Next, they ectopically express a secreted form of Sema7a to show that lateral line terminals can be ectopically attracted to the source. Finally, they also demonstrate that the synaptic assembly is impaired in the sema7a mutant. Overall, the data are of high quality and properly controlled. The availability of Sema7a antibody is a big plus, as it allows to address the endogenous protein localization as well to show the signal absence in the sema7a mutant. The quantification of the arbor topology should be useful to people in the field who are looking at the lateral line as well as other axonal terminals.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
In this paper, Boi et al. thoroughly classified the electrophysiological and morphological characteristics of serotonergic and dopaminergic neurons in the DRN and examined the alterations of these neurons in the 6-OHDA-induced mouse PD model. Using whole-cell patch clamp recording, they found that 5-HT and dopamine (DA) neurons in the DRN are electrophysiologically distinct from each other. Additionally, they characterized distinct morphological features of 5-HT and DA neurons in the DRN. Notably, these specific features of 5-HT and DA neurons in the DRN exhibited different changes in the 6-OHDA-induced PD model. Then the authors utilized desipramine (DMI) to separate the effects of nigrostriatal DA depletion and noradrenaline (NA) depletion induced by 6-OHDA. Interestingly, protection from NA depletion by DMI pretreatment reversed the changes in 5-HT neurons, while having a minor impact on the changes in DA neurons in the DRN. These data indicate that the role of NA lesion in the altered properties of DRN 5-HT neurons by 6-OHDA is more critical than that of DA lesions.
Overall, this study provides foundational data on the 5-HT and DA neurons in the DRN and their potential involvement in PD symptoms. Given the deficits of the DRN in PD, this paper may offer insights into the cellular mechanisms underlying non-motor symptoms associated with PD.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
The authors presented point light displays of human walkers to children (mean = 9 years) with and without ADHD to compare their biological motion perception abilities, and relate them to IQ, social responsiveness scale (SRS) scores and age. They report that children with ADHD were worse at all three biological motion tasks, but that those loading more heavily on local processing related to social interaction skills and global processing to age. The valuable and solid findings are informative for understanding this complex condition, as well as biological motion processing mechanisms in general. However, the correlations present a pattern that needs further examination in future studies because many of the differences between correlations are not significant.
Strengths:
The authors present differences between ADHD and TD children in biological motion processing, and this question has not received as much attention as equivalent processing capabilities in autism. They use a task that appears well controlled. They raise some interesting mechanistic possibilities for differences in local and global motion processing, which are distinctions worth exploring. The group differences will therefore be of interest to those studying ADHD, as well as other developmental conditions, and those examining biological motion processing mechanisms in general.
Weaknesses:
The data are not strong enough to support claims about differences between global and lobal processing wrt social communication skills and age. The mechanistic possibilities for why these abilities may dissociate in such a way are interesting, but the crucial tests of differences between correlations do not present a clear picture. Further empirical work would be needed to test this further. Specifics:
The authors state frequently that it was the local BM task that related to social communication skills (SRS) and not the global tasks. However, the results section shows a correlation between SRS and all three tasks. The only difference is that when looking specifically within the ADHD group, the correlation is only significant for the local task. The supplementary materials demonstrate that tests of differences between correlations present an incomplete picture. Currently they have small samples for correlations, so this is unsurprising.
Theoretical assumptions. The authors make some statements about local vs global biological motion processing that may have been made in previous studies, but would appear controversial and not definitive. E.g., that local BM processing does not improve with age and is uninfluenced by attention.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The paper presents a novel approach to expand iPSC-derived pdx1+/nkx6.1+ pancreas progenitors, making them potentially suitable for GMP-compatible protocols. This advancement represents a significant breakthrough for diabetes cell replacement therapies, as one of the current bottlenecks is the inability of expanding PP without compromising their differentiation potential. The study employs a robust dataset and state-of-the-art methodology, unveiling crucial signaling pathways (eg TGF, Notch...) responsible for sustaining pancreas progenitors while preserving their differentiation potential in vitro.
The current version of the paper has improved, increasing the clarity and providing clear explanations to the comments raised regarding quantifications, functionality of the cells in vivo etc...
The discussion on challenges adds depth to the study and encourages future research to build upon these important findings
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Goldstein et al. provide a thorough characterization of the interaction of attention and eye movement planning. These processes have been thought to be intertwined since at least the development of the Premotor Theory of Attention in 1987, and their relationship has been a continual source of debate and research for decades. Here, Goldstein et al. capitalize on their novel urgent saccade task to dissociate the effects of endogenous and exogenous attention on saccades towards and away from the cue. They find that attention and eye movements are, to some extent, linked to one another but that this link is transient and depends on the nature of the task. A primary strength of the work is that the researchers are able to carefully measure the timecourse of the interaction between attention and eye movements in various well-controlled experimental conditions. As a result, the behavioral interplay of two forms of attention (endogenous and exogenous) is illustrated at the level of tens of milliseconds as they interact with the planning and execution of saccades towards and away from the cued location. Overall, the results allow the authors to make meaningful claims about the time course of visual behavior, attention, and the potential neural mechanisms at a timescale relevant to everyday human behavior.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Nagy et al investigated the role of volume increase and swelling in neutrophils in response to the chemoattractant. Authors show that following chemoattractant response cells lose their volume slightly owing to the cell spreading phase and then have a relatively rapid increase in the cell volume that is concomitant with cell migration. Authors performed an impressive genome-wide CRISPR screen and buoyant density assay to identify the regulators of neutrophil swelling. This assay showed that stimulating cells with chemoattractant fMLP lead to an increase in the cell volume that was abrogated with the FPR1 receptor knockout. The screen revealed a cascade that could potentially be involved cell swelling including NHE1 (sodium-proton antiporter) and PI3K. NHE1 and PI3K is required for chemoattractant-induced swelling in human primary neutrophils. Authors also suggest slightly different functions of NHE1 and PI3K activity where PI3K is also required for maintain chemoattractant-induced cell shape changes. Authors convincingly show that chemoattractant induced cell swelling is linked to cell migration and NHE1 is required for swelling at the later stages of swelling since the cells at the early point work on low-volume and low-velocity regime. Interesting authors also show that lack of swelling in NHE1 inhibited cells could be rescued by mild hypo-osmotic swelling strengthening the argument that water influx followed chemoattractant stimulation is important for potentiation for migration.
The conclusions of this paper are mostly well supported by data and is pretty convincing
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript seeks to reconcile observations in multisensory perception - from behavior and neural responses. It is intuitively obvious that perceiving a stimulus via two senses results in better performance than one alone. In fact, it is not uncommon to observe that for a perceptual task, the percentage of correct responses seen with two senses is higher than the sum of the percentage correct obtained with each modality individually. i.e. the gains are "superadditive". The gains of adding a second sense are typically larger when the performance with the first sense is relatively poor - this effect is often called the principle of inverse effectiveness. More generally, what this tells us is that performance in a multisensory perceptual task is a non-linear sum of performance for each sensory modality alone.
Despite this abundant evidence of behavioral non-linearity in multisensory integration, evoked responses (EEG) to such sensory stimuli often show little evidence of it - and this is the problem this manuscript tackles. The key assertion made is that univariate analysis of the EEG signal is likely to average out the non-linear effects of integration. This is a reasonable assertion, and their analysis does indeed provide evidence that a multivariate approach can reveal non-linear interactions in the evoked responses.
Strengths:
It is of great value to understand how the process of multisensory integration occurs, and despite a wealth of observations of the benefits of perceiving the world with multiple senses, we still lack a reasonable understanding of how the brain integrates information. For example - what underlies the large individual differences in the benefits of two senses over one? One way to tackle this is via brain imaging, but this is problematic if important features of the processing - such as non-linear interactions are obscured by the lack of specificity of the measurements. The approach they take to the analysis of the EEG data allows the authors to look in more detail at the variation in activity across EEG electrodes, which averaging across electrodes cannot.
This version of the manuscript is well-written and for the most part clear. It shows a good understanding of the non-linear effects described above (where many studies show a poor understanding of "superadditivity" of perceptual performance) and the report of non-linear summation of neural responses is convincing.
A particular strength of the paper is their use of a statistical model of multisensory integration as their "null" model of neural responses, and the "inverted-encoder" which infers an internal representation of the stimulus which can explain the EEG responses. This encoder generates a prediction of decoding performance, which can be used to generate predictions of multisensory decoding from unisensory decoding, or from a sum of the unisensory internal representations.
In behavioural performance, it is frequently observed that the performance increase from two senses is close to what is expected from the optimal integration of information across the senses, in a statistical sense. It can be plausibly explained by assuming that people are able to weigh sensory inputs according to their reliability - and somewhat optimally. Critically the apparent "superadditive" effect on performance described above does not require any non-linearity in the sum of information across the senses but can arise from correctly weighting the information according to reliability.
The authors apply a similar model to predict the neural responses expected to audiovisual stimuli from the neural responses to audio and visual stimuli alone, assuming optimal statistical integration of information. The neural responses to audiovisual stimuli exceed the predictions of this model and this is the main evidence supporting their conclusion, and it is convincing.
Weaknesses:
The main weakness of the manuscript is that their behavioural data show no evidence of performance that exceeds the predictions of these statistical models. In fact, the models predict multisensory performance from unisensory performance pretty well. So this manuscript presents the opposite problem to that which motivated the study - neural interactions across the senses which appear to be more non-linear than perception. This makes it hard to interpret their results, as surely if these nonlinear neural interactions underlie the behaviour, then we should be able to see evidence of it in the behaviour? I cannot offer an easy explanation for this.
Overall, therefore, I applaud the motivation and the sophistication of the analysis method and think it shows great promise for tackling these problems, but the manuscript unfortunately brushes over an important problem specific to the results. It appeals to the higher-level reasoning - that non-linearity is a behavioural hallmark of integration and therefore we should see it in neural responses. Yet it ignores the fact that the behaviour observed here does not exceed the predictions of the "null" model applied to the neural response.
Part of the problem, I think, is that the authors never explain the difference between superadditivity of perceptual performance (proportion correct) and superadditivity of the underlying processing, which is implied by the EEG results but not their behavior. This is of course a difficult matter to describe succinctly or clearly (I somehow doubt I have). It is however worth addressing. The literature is full of confusing claims of superadditivity. I believe these authors understand this distinction and have an opportunity to represent it clearly for the benefit of all.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this work, Duan and Curtis addressed an important issue related to the nature of working memory representations. This work is motivated by findings illustrating that orientation decoding performance for perceptual representations can be biased by the stimulus aperture (modulator). Here, the authors examined whether the decoding performance for working memory representations is similarly influenced by these aperture biases. The results provide convincing evidence that working memory representations have a different representational structure, as the decoding performance was not influenced by the type of stimulus aperture.
Strengths:
The strength of this work lies in the direct comparison of decoding performance for perceptual representations with working memory representations. The authors take well-motivated approach and illustrate that perceptual and working memory representations do not share a similar representational structure. The authors test a clear question, with a rigorous approach and provide compelling evidence. First, the presented oriented stimuli are carefully manipulated to create orthogonal biases introduced by the stimulus aperture (radial or angular modulator), regardless of the stimulus carrier orientation. Second, the authors implement advanced methods to decode the orientation information, in visual and parietal cortical regions, when directly perceiving or holding an oriented stimulus in memory. The data illustrates that working memory decoding is not influenced by the type of aperture, while this is the case in perception. In sum, the main claims are important and shed light on the nature of working memory representations.
Weaknesses:
After the authors revised the original manuscript, a few of my initial concerns remain.
(1) Theoretical framing in the introduction. The introduction proposes that decoding of orientation information during perception does not reflect orientation selectivity, and it is instead driven by coarse scale biases. This is an overstatement. Recent work shows that orientation decoding is indeed influenced by coarse biases, but also reflects orientation selectivity (Roth, Kay & Merriam, 2022).
(2) The description of the image computable V1 model remains incomplete. The steerable pyramid is a model that simulates the responses of V1 neurons. To do so, it incorporates a set of linear receptive fields with varying orientation and spatial frequency tuning. However, the information that is lacking in the Methods is whether the implemented pyramid also included two quadrature phase pairs (odd and even phase Gabor filters making the output phase invariant). The sum of the squares of the responses to these offset phase filters computes the stimulus energy within each orientation and spatial frequency channel. Without this description, it is unclear what the model output represents.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, Alam et al. sought to understand how memory interacts with incoming visual information to effectively guide human behavior by using a task that combines spatial contexts (houses) with objects of one or multiple semantic categories. Three additional datasets (all from separate participants) were also employed: one that functionally localized regions of interest (ROIs) based on subtractions of different visually presented category types (in this case, scenes, objects, and scrambled objects); another consisting of resting-state functional connectivity scans, and a section of the Human Connectome Project that employed DTI data for structural connectivity analysis. Across multiple analyses, the authors identify dissociations between regions preferentially activated during scene or object judgments, between the functional connectivity of regions demonstrating such preferences, and in the anatomical connectivity of these same regions. The authors conclude that the processing streams that take in visual information and support semantic or spatial processing are largely parallel and distinct.
Strengths:
(1) Recent work has reconceptualized the classic default mode network as two parallel and interdigitated systems (e.g., Braga & Buckner, 2017; DiNicola et al., 2021). The current manuscript is timely in that it attempts to describe how information is differentially processed by two streams that appear to begin in visual cortex and connect to different default subnetworks. Even at a group level where neuroanatomy is necessarily blurred across individuals, these results provide clear evidence of stimulus-based dissociation.
(2) The manuscript contains a large number of analyses across multiple independent datasets. It is therefore unlikely that a single experimenter choice in any given analysis would spuriously produce the overall pattern of results reported in this work.
Weaknesses:
(1) Throughout the manuscript, a strong distinction is drawn between semantic and spatial processing. However, given that only objects and spatial contexts were employed in the primary experiment, it is not clear that a broader conceptual distinction is warranted between "semantic" and "spatial" cognition. There are multiple grounds for concern regarding this basic premise of the manuscript.<br /> a. One can have conceptual knowledge of different types of scenes or spatial contexts. A city street will consistently differ from a beach in predictable ways, and a kitchen context provides different expectations than a living room. Such distinctions reflect semantic knowledge of scene-related concepts, but in the present work spatial and "all other" semantic information are considered and discussed as distinct and separate.<br /> b. As a related question, are scenes uniquely different from all other types of semantic/category information? If faces were used instead of scenes, could one expect to see different regions of the visual cortex coupling with task-defined face > object ROIs? The current data do not speak to this possibility, but as written the manuscript suggests that all (non-spatial) semantic knowledge should be processed by the FT-DMN.<br /> c. Recent precision fMRI studies characterizing networks corresponding to the FT-DMN and MTL-DMN have associated the former with social cognition and the latter with scene construction/spatial processing (DiNicola et al., 2020; 2021; 2023). This is only briefly mentioned by the authors in the current manuscript (p. 28), and when discussed, the authors draw a distinction between semantic and social or emotional "codes" when noting that future work is necessary to support the generality of the current claims. However, if generality is a concern, then emphasizing the distinction between object-centric and spatial cognition, rather than semantic and spatial cognition, would represent a more conservative and better-supported theoretical point in the current manuscript.
(2) Both the retrosplenial/parieto-occipital sulcus and parahippocampal regions are adjacent to the visual network as defined using the Yeo et al. atlas, and spatial smoothness of the data could be impacting connectivity metrics here in a way that qualitatively differs from the (non-adjacent) FT-DMN ROIs. Although this proximity is a basic property of network locations on the cortical surface, the authors have several tools at their disposal that could be employed to help rule out this possibility. They might, for instance, reduce the smoothing in their multi-echo data, as the current 5 mm kernel is larger than the kernel used in Experiment 2's single-echo resting-state data. Spatial smoothing is less necessary in multi-echo data, as thermal noise can be attenuated by averaging over time (echoes) instead of space (see Gonzalez-Castillo et al., 2016 for discussion). Some multi-echo users have eschewed explicit spatial smoothing entirely (e.g., Ramot et al., 2021), just as the authors of the current paper did for their RSA analysis. Less smoothing of E1 data, combined with a local erosion of either the MTL-DMN and VIS masks (or both) near their points of overlap in the RSFC data, would improve confidence that the current results are not driven, at least in part, by spatial mixing of otherwise distinct network signals.
(3) The authors identify a region of the right angular gyrus as demonstrating a "potential role in integrating the visual-to-DMN pathways." This would seem to imply that lesion damage to right AG should produce difficulties in integrating "semantic" and "spatial" knowledge. Are the authors aware of such a literature? If so, this would be an important point to make in the manuscript as it would tie in yet another independent source of information relevant to the framework being presented. The closest of which I am aware involves deficits in cued recall performance when associates consisted of auditory-visual pairings (Ben-Zvi et al., 2015), but that form of multi-modal pairing is distinct from the "spatial-semantic" integration forwarded in the current manuscript.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The authors attempt to establish presaccadic pupil size as an index of 'saccade effort' and propose this index as one new predictor of saccade target selection. They only partially achieved their aim: When choosing between two saccade directions, the less costly direction, according to preceding pupil size, is preferred. However, the claim that with increased cognitive demand participants would especially cut costly directions is not supported by the data. I would have expected to see a negative correlation between saccade effort and saccade direction 'change' under increased load. Yet participants mostly cut upwards saccades, but not other directions that, according to pupil size, are equally or even more costly (e.g. oblique saccades).
Strengths:
The paper is well-written, easy to understand, and nicely illustrated.
The sample size seems appropriate, and the data were collected and analyzed using solid and validated methodology.
Overall, I find the topic of investigating factors that drive saccade choices highly interesting and relevant.
Weaknesses:
The authors obtain pupil size and saccade preference measures in two separate tasks. Relating these two measures is problematic because the computations that underly saccade preparation differ. In Experiment 1, the saccade is cued centrally, and has to be delayed until a "go-signal" is presented; In Experiment 2, an immediate saccade is executed to an exogenously cued peripheral target. The 'costs' in Experiment 1 (computing the saccade target location from a central cue; withholding the saccade) do not relate to Experiment 2. It is unfortunate, that measuring presaccadic pupil size directly in the comparatively more 'natural' Experiment 2 (where saccades did not have to be artificially withheld) does not seem to be possible. This questions the practical application of pupil size as an index of saccade effort
The authors claim that the observed direction-specific 'saccade costs' obtained in Experiment 1 "were not mediated by differences in saccade properties, such as duration, amplitude, peak velocity, and landing precision (Figure 1e,f)". Saccade latency, however, was not taken into account here but is discussed for Experiment 2.
The apparent similarity of saccade latencies and pupil size, however, is striking. Previous work shows shorter latencies for cardinal than oblique saccades, and shorter latencies for horizontal and upward saccades than downward saccades - directly reflecting the pupil sizes obtained in Experiment 1 as well as in the authors' previous study (Koevoet et al., 2023, PsychScience).
-
The authors state that "from a costs-perspective, it should be efficient to not only adjust the number of saccades (non-specific), but also by cutting especially expensive directions the most (specific)". However, saccade targets should be selected based on the maximum expected information gain. If cognitive load increases (due to an additional task) an effective strategy seems to be to perform less - but still meaningful - saccades. How would it help natural orienting to selectively cut saccades in certain (effortful) directions? Choosing saccade targets based on comfort, over information gain, would result in overall more saccades to be made - which is non-optimal, also from a cost perspective.
Overall, I am not sure what practical relevance the relation between pupil size (measured in a separate experiment) and saccade decisions has for eye movement research/vision science. Pupil size does not seem to be a straightforward measure of saccade effort. Saccade latency, instead, can be easily extracted in any eye movement experiment (no need to conduct a separate, delayed saccade task to measure pupil dilation), and seems to be an equally good index.
-
-
-
Reviewer #2 (Public Review):
Summary:
In this study, Vicaro et al. aimed to quantify and characterize mosaic mutations in human sporadic Alzheimer's disease (AD) brain samples. They focused on three broad classes of brain cells, neurons that express the marker NeuN, microglia that express the marker PU.1, and double-negative cells that presumably comprise all other brain cell types, including astrocytes, oligodendrocytes, oligodendrocyte progenitor cells, and endothelial cells. The authors find an enrichment of potentially pathogenic somatic mutations in AD microglia compared to controls, with MAPK pathway genes being particularly enriched for somatic mutations in those cells. The authors report a striking enrichment for mutations in the gene CBL and use in vitro functional assays to show that these mutations indeed induce MAPK pathway activation.
The current state of the AD and somatic mutation fields puts this work into context. First, AD is a devastating disease whose prevalence is only increasing as the population of the U.S. is aging, necessitating the investigation of novel features of AD to identify new therapeutic opportunities. Second, microglia have recently come into focus as important players in AD pathogenesis. Many AD risk genes are selectively expressed in microglia, and microglia from AD brain samples show a distinct transcriptional profile indicating an inflammatory phenotype. The authors' previous work shows that a genetic mouse model of mosaic BRAF activation in macrophages (including microglia) displays a neurodegenerative phenotype similar to AD (Mass et al., 2017, doi:10.1038/nature23672). Third, new technological developments have allowed for identifying mosaic mutations present in only a small fraction of or even single cells. Together, these data form a rationale for studying mosaic mutations in microglia in AD. In light of the authors' findings regarding MAPK pathway gene somatic mutations, it is also important to note that MAPK has previously been implicated in AD neuroinflammation in the literature.
Strengths:
The study demonstrated several strengths.
Firstly, the authors used two methods to identify mosaic mutations:<br /> (1) deep (~1,100x) DNA sequencing of a targeted panel of 716 genes they hypothesized might, if mutated somatically, play a role in AD, and<br /> (2) deep (400x) whole-exome sequencing (WES) to identify clonal mosaics outside of those 716 genes.
A second strength is the agreement between these experiments, where WES found many variants identified in the panel experiment, and both experiments revealed somatic mutations in MAPK pathway genes.
Third, the authors demonstrated in several in vitro systems that many mutations they identified in MAPK genes activate MAPK signaling. Finally, the authors showed that in some human brain samples, single-cell gene expression analysis revealed that cells bearing a mosaic MAPK pathway mutation displayed dysregulated inflammatory signaling and dysregulation in other pathways. This single-cell analysis was in agreement with their in vitro analyses.
Weaknesses:
The study also showed some weaknesses. The sample size (45 AD donors and 44 controls) is small, reflected in the relatively modest effect sizes and p-values observed. This weakness is partially ameliorated by the authors' extensive molecular and functional validation of mutation candidates. Another weakness is the lack of discussion of whether the genes found to be mutated somatically in AD show any AD-risk alleles in the population. If they did, it would further support the authors' conclusions that they are playing a role in AD. Finally, as the authors point out, this study cannot conclude whether microglial mosaic mutations cause AD or are an effect of AD. Future studies may shed more light on this important question.
Conclusions and Impact:
Considering the study's aims, strengths, and weaknesses, I conclude that the authors achieved their goal of characterizing the role of mosaic mutations in human AD. Their data strongly suggest that mosaic MAPK mutations in microglia are associated with AD. The impacts of this study remain to be seen, but they could include attempts to target CBL or other mutated genes in the treatment of AD. This work also suggests a similar approach to identifying potentially causative somatic mutations in other neurodegenerative diseases.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This is a very interesting paper that leveraged several publicly available datasets: invasive cortical recording in epilepsy patients, functional and structural connectomic data, and PET data related to dopaminergic and gaba-ergic synapses. These were combined to create a unified hypothesis of beta band oscillatory activity in the human brain. They show that beta frequency activity is ubiquitous, not just in sensorimotor areas, and cortical regions where beta predominated had high connectivity to regions high in dopamine re-update.
Strengths:
The authors leverage and integrate three publicly available human brain datasets in a creative way. While these public datasets are powerful tools for human neuroscience, it is innovative to combine these three types of data into a common brain space to generate novel findings and hypotheses. Findings are nicely controlled by separately examining cortical regions where alpha predominates (which have a different connectivity pattern). GABA uptake from PET studies is used as a control for the specificity of the relationship between beta activity and dopamine uptake. There is much interest in synchronized oscillatory activity as a mechanism of brain function and dysfunction, but the field is short on unifying hypotheses of why particular rhythms predominate in particular regions. This paper contributes nicely to that gap. It is ambitious in generating hypotheses, particularly that modulation of beta activity may be used as a "proxy" for modulating phasic dopamine release.
Weaknesses:
As the authors point out, the use of normative data is excellent for exploring hypotheses but does not address or explore individual variations which could lead to other insights. It is also biased to resting state activity; maps of task-related activity (if they were available) might show different findings.
The figures, results, introduction, and methods are admirably clear and succinct but the discussion could be both shorter and more convincing.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Recent studies have identified specific regions within the occipito-temporal cortex as part of a broader fronto-parietal, domain-general, or "multiple-demand" (MD) network that mediates fluid intelligence (gF). According to the abstract, the authors aim to explore the mechanistic roles of these occipito-temporal regions by examining GABA/glutamate concentrations. However, the introduction presents a different rationale: investigating whether area MT+ specifically, could be a core component of the MD network.
Strengths:
The authors provide evidence that GABA concentrations in MT+ and its functional connectivity with frontal areas significantly correlate with visuo-spatial intelligence performance. Additionally, serial mediation analysis suggests that inhibitory mechanisms in MT+ contribute to individual differences in a specific subtest of the Wechsler Adult Intelligence Scale, which assesses visuo-spatial aspects of gF.
Weaknesses:
While the findings are compelling and the analyses robust, the study's rationale and interpretations need strengthening. For instance, Assem et al. (2020) have previously defined the core and extended MD networks, identifying the occipito-temporal regions as TE1m and TE1p, which are located more rostrally than MT+. Area MT+ might overlap with brain regions identified previously in Fedorenko et al., 2013, however the authors attribute these activations to attentional enhancement of visual representations in the more difficult conditions of their tasks. For the aforementioned reasons, It is unclear why the authors chose MT+ as their focus. A stronger rationale for this selection is necessary and how it fits with the core/extended MD networks.
Moreover, although the study links MT+ inhibitory mechanisms to a visuo-spatial component of gF, this evidence alone may not suffice to position MT+ as a new core of the MD network. The MD network's definition typically encompasses a range of cognitive domains, including working memory, mathematics, language, and relational reasoning. Therefore, the claim that MT+ represents a new core of MD needs to be supported by more comprehensive evidence.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This study takes advantage of multiple methodological advances to perform layer-specific staining of cortical neurons and tracking of their axons to identify the pattern of their projections. This publication offers a mesoscale view of the projection patterns of neurons in the whisker primary and secondary somatosensory cortex. The authors report that, consistent with the literature, the pattern of projection is highly different across cortical layers and subtype, with targets being located around the whole brain. This was tested across 6 different mouse types that expressed a marker in layer 2/3, layer 4, layer 5 (3 sub-types) and layer 6.<br /> Looking more closely at the projections from primary somatosensory cortex into the primary motor cortex, they found that there was a significant spatial clustering of projections from topographically separated neurons across the primary somatosensory cortex. This was true for neurons with cell bodies located across all tested layers/types.
Strengths:
This study successfully looks at the relevant scale to study projection patterns, which is the whole brain. This is achieved thanks to an ambitious combination of mouse lines, immuno-histochemistry, imaging and image processing, which results in a standardized histological pipeline that processes the whole-brain projection patterns of layer-selected neurons of the primary and secondary somatosensory cortex.<br /> This standardization means that comparisons between cell-types projection patterns are possible and that both the large-scale structure of the pattern and the minute details of the intra-areas pattern are available.<br /> This reference dataset and the corresponding analysis code are made available to the research community.
Weaknesses:
One major question raised by this dataset is the risk of missing axons during the post-processing step. Indeed, it appears that the control and training efforts have focused on the risk of false positives (see Figure 1 supplementary panels). And indeed, the risk of overlooking existing axons in the raw fluorescence data id discussed in the article.
Based on the data reported in the article, this is more than a risk. In particular, Figure 2 shows an example Rbp4-L5 mouse where axonal spread seems massive in Hippocampus, while there is no mention of this area in the processed projection data for this mouse line.
Similarily, the Ntsr1-L6CT example shows a striking level of fluorescence in Striatum, that does not reflect in the amount of axons that are detected by the algorithms in the next figures.<br /> These apparent discrepancies may be due to non axonal-specific fluorescence in the samples. In any case, further analysis of such anatomical areas would be useful to consolidate the valuable dataset provided by the article.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors re-analyse MEG data from a speech production and perception study and extend their previous Granger causality analysis to a larger number of cortical-cortical and in particular cortical-subcortical connections. Regions of interest were defined by means of a meta-analysis using Neurosynth.org and connectivity patterns were determined by calculating directed influence asymmetry indices from the Granger causality analysis results for each pair of brain regions. Abbasi et al. report feedforward signals communicated via fast rhythms and feedback signals via slow rhythms below 40 Hz, particularly during speaking. The authors highlight one of these connections between the right cerebellum lobule VI and auditory association area A5, where in addition the connection strength correlates negatively with the strength of speech tracking in the theta band during speaking (significant before multiple comparison correction). Results are interpreted within a framework of active inference by minimising prediction errors.
While I find investigating the role of cortical-subcortical connections in speech production and perception interesting and relevant to the field, I am not yet convinced that the methods employed are fully suitable to this endeavour or that the results provide sufficient evidence to make the strong claim of dissociation of bottom-up and top-down information flow during speaking in distinct frequency bands.
Strengths:
The investigation of electrophysiological cortical-subcortical connections in speech production and perception is interesting and relevant to the field. The authors analyse a valuable dataset, where they spent a considerable amount of effort to correct for speech production-related artefacts. Overall, the manuscript is well-written and clearly structured.
Weaknesses:
The description of the multivariate Granger causality analysis did not allow me to fully grasp how the analysis was performed and I hence struggled to evaluate its appropriateness.<br /> Knowing that (1) filtered Granger causality is prone to false positives and (2) recent work demonstrates that significant Granger causality can simply arise from frequency-specific activity being present in the source but not the target area without functional relevance for communication (Schneider et al. 2021) raises doubts about the validity of the results, in particular with respect to their frequency specificity. These doubts are reinforced by what I perceive as an overemphasis on results that support the assumption of specific frequencies for feedforward and top-down connections, while findings not aligning with this hypothesis appear to be underreported. Furthermore, the authors report some main findings that I found difficult to reconcile with the data presented in the figures. Overall, I feel the conclusions with respect to frequency-specific bottom-up and top-down information flow need to be moderated and that some of the reported findings need to be checked and if necessary corrected.
Major points
(1) I think more details on the multivariate GC approach are needed. I found the reference to Schaum et al., 2021 not sufficient to understand what has been done in this paper. Some questions that remained for me are:
(i) Does multivariate here refer to the use of the authors' three components per parcel or to the conditioning on the remaining twelve sources? I think the latter is implied when citing Schaum et al., but I'm not sure this is what was done here?
If it was not: how can we account for spurious results based on indirect effects?
(ii) Did the authors check whether the GC of the course-target pairs was reliably above the bias level (as Schaum et. al. did for each condition separately)? If not, can they argue why they think that their results would still be valid? Does it make sense to compute DAIs on connections that were below the bias level? Should the data be re-analysed to take this concern into account?
(iii) You may consider citing the paper that introduced the non-parametric GC analysis (which Schaum et al. then went on to apply): Dhamala M, Rangarajan G, Ding M. Analyzing Information Flow in Brain Networks with Nonparametric Granger Causality. Neuroimage. 2008; 41(2):354-362. https://doi.org/10.1016/j.neuroimage.2008.02. 020
(2) GC has been discouraged for filtered data as it gives rise to false positives due to phase distortions and the ineffectiveness of filtering in the information-theoretic setting as reducing the power of a signal does not reduce the information contained in it (Florin et al., 2010; Barnett and Seth, 2011; Weber et al. 2017; Pinzuti et al., 2020 - who also suggest an approach that would circumvent those filter-related issues). With this in mind, I am wondering whether the strong frequency-specific claims in this work still hold.
(3) I found it difficult to reconcile some statements in the manuscript with the data presented in the figures:
(i) Most notably, the considerable number of feedforward connections from A5 and STS that project to areas further up the hierarchy at slower rhythms (e.g. L-A5 to R-PEF, R-Crus2, L CB6 L-Tha, L-FOP and L-STS to R-PEF, L-FOP, L-TOPJ or R-A5 as well as R-STS both to R-Crus2, L-CB6, L-Th) contradict the authors' main message that 'feedback signals were communicated via slow rhythms below 40 Hz, whereas feedforward signals were communicated via faster rhythms'. I struggled to recognise a principled approach that determined which connections were highlighted and reported and which ones were not.
(ii) "Our analysis also revealed robust connectivity between the right cerebellum and the left parietal cortex, evident in both speaking and listening conditions, with stronger connectivity observed during speaking. Notably, Figure 4 depicts a prominent frequency peak in the alpha band, illustrating the specific frequency range through which information flows from the cerebellum to the parietal areas." There are two peaks discernible in Figure 4, one notably lower than the alpha band (rather theta or even delta), the other at around 30 Hz. Nevertheless, the authors report and discuss a peak in the alpha band.
(iii) In the abstract: "Notably, high-frequency connectivity was absent during the listening condition." and p.9 "In contrast with what we reported for the speaking condition, during listening, there is only a significant connectivity in low frequency to the left temporal area but not a reverse connection in the high frequencies."<br /> While Fig. 4 shows significant connectivity from R-CB6 to A5 in the gamma frequency range for the speaking, but not for the listening condition, interpreting comparisons between two effects without directly comparing them is a common statistical mistake (Makin and Orban de Xivry). The spectrally-resolved connectivity in the two conditions actually look remarkably similar and I would thus refrain from highlighting this statement and indicate clearly that there were no significant differences between the two conditions.
(iv) "This result indicates that in low frequencies, the sensory-motor area and cerebellum predominantly transmit information, while in higher frequencies, they are more involved in receiving it."<br /> I don't think that this statement holds in its generality: L-CB6 and R-3b both show strong output at high frequencies, particularly in the speaking condition. While they seem to transmit information mainly to areas outside A5 and STS these effects are strong and should be discussed.
(4) "However, definitive conclusions should be drawn with caution given recent studies raising concerns about the notion that top-down and bottom-up signals can only be transmitted via separate frequency channels (Ferro et al., 2021; Schneider et al., 2021; Vinck et al., 2023)."
I appreciate this note of caution and think it would be useful if it were spelled out to the reader why this is the case so that they would be better able to grasp the main concerns here. For example, Schneider et al. make a strong point that we expect to find Granger-causality with a peak in a specific frequency band for areas that are anatomically connected when the sending area shows stronger activity in that band than the receiving one, simply because of the coherence of a signal with its own linear projection onto the other area. The direction of a Granger causal connection would in that case only indicate that one area shows stronger activity than the other in the given frequency band. I am wondering to what degree the reported connectivity pattern can be traced back to regional differences in frequency-specific source strength or to differences in source strength across the two conditions.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Han et al. present a manuscript focusing on difference metabolism and the regulatory circuits controlling it in C. elegans fed two bacterial diets. In the first three figures and a half figures, using a combination of methods, they investigate lipid levels, changes in gene expression and genetic assays to come to the conclusion that vitamin B12 acts through the S-adenosylmethioine synthase sams-1 to perturb phosphatidylcholine levels, which in turn stimulate the C. elegans ortholog of the SREBP transcription factors to activate fatty acid synthesis genes such as fat-7/SCD1. Thus, while connections between diet, metabolic pathways and gene regulation is of general interest, this study largely confirms the work of others without direct credit in many instances, then fails to develop a more novel cell non-autonomous link between the pathways in the last two figures. Thus, this study would be expected to have a useful impact on the field, if it can be placed in context of previously published work.
Strengths:
(1) Connections between diet, metabolic pathways and gene regulation is of general interest<br /> (2) Figures 1-4 confirm data/observations from previously published work from MacNeil, et al. Cell 2015; Walker, et al. Cell 2011; Svensk, et al. PLoS Genetics 2013; Smulan, et al. Cell Reports, 2016; Giese, et al. eLife 2020 and Qin, et al. Cell Reports 2022..<br /> (3) The data in figures 5 and 6 showing importance of non-cell autonomous effects on metabolism.
Weaknesses:
(1) In order to differentiate their study from previous work, it seems that the authors try to make the argument that PC is higher in Comomonas than E. coli, therefore they are looking at repression of SBP-1-dependent function, however, the pairing of the diets is arbitrary, and the comparisons could easily be reversed. They are simply comparing a higher to a lower level of PC, rather than a basal to a lower, thus the concepts are the same. In addition, they fail to cite the larger body of literature linking phospholipid balance to SREBP function. For example, multiple studies in mammalian models link phospholipid balance, not just lowered PC, to SREBP function: Lim, Genes and Dev 2011; Wang, et al. Cell Stem Cell, 2018; Rong, et al. J Clin Invest 2017; Smulan et al, Cell Reports, 2016; Dobrosotskaya, Science. 2002 and recently, Rong, et al. Cell Met 2024.
(2) Figure 1: For example, the data in figure 1, shows measures of lipid content, RNA seq showing changes in metabolic enzymes such as fat-7/SCD-1 and lipid levels have already been shown in MacNeil, et al. Cell 2013 (lipid levels and gene expression changes) and the lipid levels in Comomonas vs E. coli were published in Ditot, et al. Nature Communications 2022 by Dr. Marian Walhout's lab.
(3) Figure 2/3: In Figure 2 and 3, they use a genetic screen to find regulators of fat-7/scd1 expression, and unsurprisingly, pull out genes with known to regulate this pathway. The authors go on to show that changes in SAM lead to changes in PC, and affect SBP-1/SREBP-1-dependent lipogenesis. This is a well described pathway from publications by the Walhout lab, Dr. Amy Walker's lab and Dr. Marc Pilon's lab (Walker, et al. Cell 2011; Svensk, et al. PLoS Genetics 2013; Smulan, et al. Cell Reports, 2016; Giese, et al. eLife 2020) in addition to a recent publication, Qin, et al. Cell Reports 2022. While some of these studies are cited in other places in the manuscript, the authors describe their results as "discovery", then fail to cite the relevant studies at those points (selected examples below
(4) Selected examples of citation issues:
a) Selected example: pg 6: "To understand the mechanism underlying the regulation of host lipid content triggered by DA, we examined the gene expression changes elicited by the two different bacterial diets in young adult animals by RNA-seq...In particular, genes related to the biosynthesis of unsaturated fatty acids showed a significant decrease in expression in DA-fed worms. For example, the delta-(9) fatty acid desaturases, fat-5 and fat-7, (which convert fatty acids 16:0 to 16:1n7 and 18:0 to 18:1n9, respectively32) decreased"
MacNeil et al Cell 2013 published a transcriptomics comparing young adult DA and Op50, which demonstrated decreases in fat-5 and fat-7. While MacNeil is cited in other parts of the paper, since the authors have performed a highly similar experiment and obtained similar results, this should be described as confirming the MacNeil study rather than as new data.
b) Selected Example: pg 10: "To determine whether PC levels have a causal effect on organismal lipid content, we supplemented worm diets with choline, the PC precursor, and uncovered a dose-dependent decrease in lipid content as measured by O.R.O staining (Figure 3B)."
Addition of choline to supplement defects in PC synthesis was first shown by Brendza, et al. Biochem J 2007. It was confirmed in Walker, et al. 2011, and further confirmation of PC rescue show in Ding, et al. 2015. The Brendza study is not cited at all and while studies from the Walker lab are cited in other places, the authors omit that changes in the DA diet are the same as changes seen when choline rescues PC loss from other perturbations.
c) Selected Example: pg 9: "Notably, DA has been reported as a B12-rich bacterium compared to OP16, hinting at the possibility that the DA diet might boost dietary B12 levels."
Reference 16 is Watson, et al. Cell 2015 where the Walhout lab demonstrates that DA does in fact act through the diet to alter the Met/SAM cycle and other B12 dependent processes in C. elegans. This paper, along with MacNeil above broke ground in linking B12 and the Met/SAM cycle to specific phenotypes in C. elegans, which was followed up by extensive work from the Walhout lab on this cycle, thus, it seems odd that the authors describe their own data as "hinting" at this connection.
d) Selected example: pg 17: "Indeed, this is further supported by our observation that mutants of histone methyltransferases SET-2 and SET-30 (which install H3K4me1 and H3K4me2, respectively) exhibited elevated lipid content on DA diet (data not shown). Notably, while both set-2 and set-30 mutants had this effect, only set-2 appears to control fat-7 expression (data not shown)". Extensive work from Dr. Anne Brunet's lab (Greer, et al. Nature 2010; Greer, et al. Nature 2011; Han, et al. Nature 2017) link set-2 and H3K4 methylation to lipid accumulation and fat-7. The authors fail to cite these studies.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The study by Li et al. aimed to demonstrate the role of the G𝛾13-mediated signal transduction pathway in tuft cell-driven inflammation resolution and repairing injured lung tissue. The authors showed the reduced number of tuft cells in the parenchyma of G𝛾13 null lungs following viral infection. Mice with a G𝛾13 null mutation showed increased lung damage and heightened macrophage infiltration when exposed to the H1N1 virus. Their further findings suggested that lung inflammation resolution, epithelial barrier and fibrosis were worsen in G𝛾13 null mutants.
Strengths:
The revised study carefully analyzed phenotypes in mice lacking G𝛾13 in response to viral infection, providing further support that G𝛾13+ tuft cells play a role in the resolution of inflammation and injury repair.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript shows detailed evidence about the role of cohesin regulator in rice meiosis and mitosis
Strengths:
There is a very clear mechanism for its role during replication
Weaknesses:
The authors did not consider to create heterozygous mutants for the replication fork.
April 15. Revisions read.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The authors identify a third component in the interaction between myosin Va and melanophilin- an interaction between a 32-residue sequence encoded by exon-g in myosin Va and melanophilin's actin binding domain. This interaction has implications for how melanosome motility may be regulated.
The authors have now included some necessary controls that were requested. In terms of adding new information to increase the significance and impact of the paper, they added a single affinity measurement. Unfortunately, it did not involve Exon G specifically. Moreover, they did not add any new mechanistic or functional data to provide a more conceptual advance. For example, is the Exon G interaction regulated by phosphorylation? Is this what dictates the choice between Mlph's actin binding domain (ABD) binding to actin or to exon-G. How does local actin concentration influence this decision. What changes regarding melanosome dynamics in cells between these two alternatives? Do in vitro reconstitution assays show that binding to Exon-G instead of actin affects the processivity of a Rab27a/Myosin 5a/Mlph transport complex? Finally, while the authors make clear in the abstract and text that they are just identifying a third component that mediates the Melanophilin-dependent association of myosin-5a with melanosomes, the title gives the impression that they identified all three in this manuscript. I really think the title should be changed to something like Identification of a third component that mediates the Melanophilin-dependent association of myosin-5a with melanosomes, as this accurately reflects what is new in this work.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this new paper, the authors used biochemical, structural, and biophysical methods to elucidate the mechanisms by which IP4, the PIP3 headgroup, can induce an autoinhibit form of P-Rex1 and propose a model of how PIP3 can trigger long-range conformational changes of P-Rex1 to relieve this autoinhibition. The main findings of this study are that a new P-Rex1 autoinhibition is driven by an IP4-induced binding of the PH domain to the DH domain active site and that this autoinhibit form stabilized by two key interactions between DEP1 and DH and between PH and IP4P 4-helix bundle (4HB) subdomain. Moreover, they found that the binding of phospholipid PIP3 to the PH domain can disrupt these interactions to relieve P-Rex1 autoinhibition.
Strengths:
The study provides good evidence that binding of IP4 to the P-Rex1 PH domain can make the two long-range interactions between the catalytic DH domain and the first DEP domain, and between the PH domain and the C-terminal IP4P 4HB subdomain that generate a novel P-Rex1 autoinhibition mechanism. This valuable finding adds an extra layer of P-Rex1 regulation (perhaps in the cytoplasm) to the synergistic activation by phospholipid PIP3 and the heterotrimeric Gβγ subunits at the plasma membrane. Overall, this manuscript's goal sounds interesting, the experimental data were carried out carefully and reliably.
Weakness:
The set of experiments with the disulfide bond S235C/M244C caused a bit of confusion for interpretation, it should be moved into the supplement, and the text and Figure 4 were altered accordingly.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This study represents an ambitious endeavor to comprehensively analyze the role of miR-199a/b-5p and its networks in cartilage formation. By conducting experiments that go beyond in vitro MSC differentiation models, more robust conclusions can be achieved.
Strengths:
This research investigates the role of miR-199a/b-5p during chondrogenesis using bioinformatics and in vitro experimental systems. The significance of miRNAs in chondrogenesis and OA is crucial, warranting further research, and this study contributes novel insights.
Weaknesses:
While miR-140 and miR-455 are used as controls, these miRNAs have been demonstrated to be more relevant to Cartilage Homeostasis than chondrogenesis itself. Their deficiency has been genetically proven to induce Osteoarthritis in mice. Therefore, the results of this study should be considered in comparison with these existing findings.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript mainly studied the biological effect of tenascin XB (TNXB) on hemophilic arthropathy (HA) progression. Using bioinformatic and histopathological approaches, the authors identified the novel candidate gene TNXB for HA. Next, authors showed that TNXB knockdown lead to chondrocyte apoptosis, matrix degeneration and subchondral bone loss in vivo/vitro. Furthermore, AKT agonist promoted extracellular matrix synthesis and prevented apoptosis in TNXB knockdown chondrocytes.
Strengths:
In general, this study significantly advances our understanding of HA pathogenesis. The authors utilize comprehensive experimental strategies to demonstrate the role of TNXB in cartilage degeneration associated with HA. The results are clearly presented, and the conclusions appear appropriate.
Weaknesses:
Additional clarification is required regarding the gender of the F8-/- mouse in the study. Is the mouse male or female?
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This study takes a new approach to studying the role of corticofugal projections from auditory cortex to inferior colliculus. The authors performed two-photon imaging of cortico-recipient IC neurons during a click detection task in mice with and without lesions of auditory cortex. In both groups of animals, they observed similar task performance and relatively small differences in the encoding of task-response variables in the IC population. They conclude that non-cortical inputs to the IC provide can substantial task-related modulation, at least when AC is absent.
Strengths:
This study provides valuable new insight into big and challenging questions around top-down modulation of activity in the IC. The approach here is novel and appears to have been executed thoughtfully. Thus, it should be of interest to the community.
Weaknesses:
There are however, substantial concerns about the interpretation of the findings and limitations to the current analysis. In particular, Analysis of single unit activity is absent, making interpretation of population clusters and decoding less interpretable. These concerns should be addressed to make sure that the results can be interpreted clearly in an active field that already contains a number of confusing and possibly contradictory findings.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors' report describes a novel vaccine platform derived from a newly discovered organelle called a migrasome. First, the authors address a technical hurdle in using migrasomes as a vaccine platform. Natural migrasome formation occurs at low levels and is labor intensive, however, by understanding the molecular underpinning of migrasome formation, the authors have designed a method to make engineered migrasomes from cultured, cells at higher yields utilizing a robust process. These engineered migrasomes behave like natural migrasomes. Next, the authors immunized mice with migrasomes that either expressed a model peptide or the SARS-CoV-2 spike protein. Antibodies against the spike protein were raised that could be boosted by a 2nd vaccination and these antibodies were functional as assessed by an in vitro pseudoviral assay. This new vaccine platform has the potential to overcome obstacles such as cold chain issues for vaccines like messenger RNA that require very stringent storage conditions.
Strengths:
The authors present very robust studies detailing the biology behind migrasome formation and this fundamental understanding was used to form engineered migrasomes, which makes it possible to utilize migrasomes as a vaccine platform. The characterization of engineered migrasomes is thorough and establishes comparability with naturally occurring migrasomes. The biophysical characterization of the migrasomes is well done including thermal stability and characterization of the particle size (important characterizations for a good vaccine).
Weaknesses:
With a new vaccine platform technology, it would be nice to compare them head-to-head against a proven technology. The authors would improve the manuscript if they made some comparisons to other vaccine platforms such as a SARS-CoV-2 mRNA vaccine or even an adjuvanted recombinant spike protein. This would demonstrate a migrasome-based vaccine could elicit responses comparable to a proven vaccine technology. Additionally, understanding the integrity of the antigens expressed in their migrasomes could be useful. This could be done by looking at functional monoclonal antibodies binding to their migrasomes in a confocal microscopy experiment.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The aim of the study was to understand how cells of the skin communicate across dermal layers. The research group has previously demonstrated that cellular connections called airinemes contribute to this communication. The current work builds upon this knowledge by showing that differentiated keratinocytes also use cytonemes, specialized signaling filopodia, to communicate with undifferentiated keratinocytes. They show that cytonemes are the more abundant type of cellular extension used for communication between the differentiated keratinocyte layer and the undifferentiated keratinocytes. Disruption of cytoneme formation led to the expansion of the undifferentiated keratinocytes into the periderm, mimicking skin diseases like psoriasis. The authors go on to show that disruption of cytonemes results in perturbations in Notch signaling between the differentiated keratinocytes of the periderm and the underlying proliferating undifferentiated keratinocytes. Further, the authors show that Interleukin-17, also known to drive psoriasis, can restrict the formation of periderm cytonemes, possibly through the inhibition of Cdc42 expression. This work suggests that cytoneme-mediated Notch signaling plays a central role in normal epidermal regulation. The authors propose that disruption of cytoneme function may be an underlying cause of various human skin diseases.
Strengths:
The authors provide strong evidence that periderm keratinocytes cytonemes contain the notch ligand DeltaC to promote Notch activation in the underlying intermediate layer to regulate accurate epidermal maintenance.
Weaknesses:
The impact of the study would be increased if the mechanism by which Interlukin-17 and Cdc42 collaborate to regulate cytonemes was defined. Experiments measuring Cdc42 activity, rather than just measuring expression, would strengthen the conclusions.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Juvenile hormone (JH) is a pleiotropic terpenoid hormone in insects that mainly regulates their development and reproduction. In particular, its developmental functions are described as the "status quo" action, as its presence in the hemolymph (the insect blood) prevents metamorphosis-initiating effects of ecdysone, another important hormone in insect development, and maintains the juvenile status of insects.
While such canonical functions of JH are known to be mediated by its intracellular receptor complex composed of Met and Tai, there have been multiple reports suggesting the presence of cell membrane receptor(s) for JH, which mediate non-genomic effects of this terpenoid hormone. In particular, the presence of receptor tyrosine kinase(s) that phosphorylate Met/Tai in response to JH and thus indirectly affect the canonical JH signaling pathway has been strongly suggested. Given the importance of JH in insect physiology and the fact that the JH signaling pathway is a major target of insect growth regulators, elucidating the identification and functions of putative JH membrane receptors is of great significance from both basic and applied perspectives.
In the present study, the authors identified candidate receptors for such cell membrane JH receptors, CAD96CA and FGFR1, in the cotton bollworm Helicoverpa armigera.
Strengths:
Their in vitro analyses are conducted thoroughly using multiple methods, which overall supports their claim that these receptors can bind to JH and mediate their non-genomic effects.
Weaknesses:
Results of their in vivo experiments, particularly those of their loss-of-function analyses using CRISPR mutants are still preliminary, and the results rather indicate that these membrane receptors do not have any physiologically significant roles in vivo. More specifically, previous studies in lepidopteran species have clearly and repeatedly shown that precocious metamorphosis is the hallmark phenotype for all JH signaling-deficient larvae. In contrast, the present study showed that Cad96ca and Fgfr1 G0 mutants only showed a slight acceleration in their pupation timing, which is not a typical phenotype one would expect from JH signaling deficiency. This is inconsistent with their working model provided in Figure 6, which indicates that these cell membrane JH receptors promote the canonical JH signaling by phosphorylating Met/Tai.
If the authors argue that this slight acceleration of pupation is indeed a major JH signaling-deficient phenotype in Helicoverpa, they need to provide more data to support their claim by analyzing CRISPR mutants of other genes involved in JH signaling, such as Jhamt and Met. An alternative explanation is that there is functional redundancy between CAD96CA and FGFR1 in mediating phosphorylation of Met/Tai. This possibility can be tested by analyzing double knockouts of these two receptors.
Currently, the validity of their calcium imaging analysis in Figure 5 is also questionable. When performing calcium imaging in cultured cells, it is critically important to treat all the cells at the end of each experiment with a hormone or other chemical reagents that universally induce calcium increase in each particular cell line. Without such positive control, the validity of calcium imaging data remains unknown, and readers cannot properly evaluate their results.
-
-
www.ncbi.nlm.nih.gov www.ncbi.nlm.nih.gov
-
Bloomington, 16671
DOI: 10.1038/s41467-023-43550-2
Resource: BDSC_16671
Curator: @maulamb
SciCrunch record: RRID:BDSC_16671
-
Bloomington, 9575
DOI: 10.1038/s41467-023-43550-2
Resource: (BDSC Cat# 9575,RRID:BDSC_9575)
Curator: @maulamb
SciCrunch record: RRID:BDSC_9575
-
BDSC, 1309
DOI: 10.1038/s41467-023-43550-2
Resource: (BDSC Cat# 1309,RRID:BDSC_1309)
Curator: @maulamb
SciCrunch record: RRID:BDSC_1309
-
BDSC, 27656
DOI: 10.1038/s41467-023-43550-2
Resource: (BDSC Cat# 27656,RRID:BDSC_27656)
Curator: @maulamb
SciCrunch record: RRID:BDSC_27656
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors carry out a careful and rigorous quantitative analysis of RecB transcript and protein levels at baseline and in response to DNA damage. Using single-molecule FISH and Halo-tagging in order to achieve sensitive measurements, they provide evidence that enhanced RecB protein levels in response to DNA damage are achieved through a post-transcriptional mechanism mediated by the Sm-like RNA binding protein, Hfq. In terms of biological relevance, the authors suggest that this mechanism provides a way to control the optimum level of RecB expression as both deletion and over-expression are deleterious. In addition, the proposed mechanism provides a new framework for understanding how transcriptional noise can be suppressed at the protein level.
Strengths:
Strengths of the manuscript include the rigorous approaches and orthogonal evidence to support the core conclusions, for example, the evidence that altering either Hhq or its recognition sequence on the RNA similarly enhance the protein to RNA ratio of RecB. The writing is clear and the experiments are well-controlled. The modeling approaches provide essential context to interpret the data, particularly given the small numbers of molecules per cell. The interpretations are careful and well supported.
Weaknesses:
The authors make a compelling case for the biological need to exquisitely control RecB levels, which they suggest is achieved by the pathway they have uncovered and described in this work. However, this conclusion is largely inferred as the authors only investigate the effect on cell survival in response to (high levels of) DNA damage and in response to two perturbations - genetic knock-out or over-expression, both of which are likely more dramatic than the range of expression levels observed in unstimulated and DNA damage conditions.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The authors have done well to address the points raised in my previous review.
The updated version of this manuscript retains the technical competence of the first, but with important changes that make the analysis more legible and results better contextualized. Specifically, the discussion is richer, the interpretation of the results is more nuanced, the terminology is more precise, and issues of clarity related to the methodology and results have been resolved.
Broad caveats remain about the nature of authorship, and who we should expect to be quoted in science journalism. Namely, who is the lead author? Ideally, the corresponding author would be included as well, or else some bibliometric definition of the most senior author on the byline. However, the authors' approach here is certainly adequate, and they did well to incorporate discussion of authorship and the scholarly division of labour in their discussion.
In sum, I find the article greatly improved and a competent analysis into the unequal use of quotations in scientific journalism.
-
-
www.derstandard.at www.derstandard.at
-
Investigativer Hintergrundbericht zum Aufbau von LNG-Kapazitäten durch Russland in der Arktis. Das Projekt Arctic LNG 2 ist nicht nur klimaschädlich, sondern auch katastrophal für die lokalen Ökosysteme. Europa bezieht mehr LNG aus Russland als vor der Invasion der kompletten Ukraine. Russland nutzte auch 2023 westliches Knowhow und trotz der Sanktionen gelieferte Ausrüstung vor allem der deutschen Firma Linde. https://www.derstandard.at/story/3000000191618/wie-westliche-konzerne-ein-russisches-prestigeprojekt-auch-nach-kriegsbeginn-ermoeglichten
-
-
www.nytimes.com www.nytimes.com
-
Ausführlicher Bericht über die Konflikte um das neue LNG-Terminal CP2 an der Küste von Louisiana. https://www.nytimes.com/2023/12/26/climate/cp2-natural-gas-export-louisiana.html
Tags
- project: Willow
- Bill McKibben
- fossil expansion
- natural gas
- 2023-12-26
- Jane Fonda
- Manish Bapna
- Louisiana
- Charif Souki
- Venture Global LNG
- CO2
- USA
- LNG expansion
- Cheniere
- Healthy Gulf
- Naomi Yoder
- Calcasieu Pass 2
- Fossil lobbying
- Natural Resources Defense Council
- Michael Sabel
- CP2
- LNG
- Greenpeace
- Biden Administration
Annotators
URL
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Liang and colleagues set out to test whether the human brain uses distance and grid-like codes in social knowledge using a design where participants had to navigate in a two-dimensional social space based on competence and warmth during an fMRI scan. They showed that participants were able to navigate the social space and found distance-based codes as well as grid-like codes in various brain regions, and the grid-like code correlated with behavior (reaction times).
On the whole, the experiment is designed appropriately for testing for distant-based and grid-like codes, and is relatively well powered for this type of study, with a large amount of behavioral training per participant. They revealed that a number of brain regions correlated positively or negatively with distance in the social space, and found grid-like codes in the frontal polar cortex and posterior medial entorhinal cortex, the latter in line with prior findings on grid-like activity in entorhinal cortex. The current paper seems quite similar conceptually and in design to previous work, most notably Park et al., 2021, Nature Neuroscience.
(1) The authors claim that this study provides evidence that humans use a spatial / grid code for abstract knowledge like social knowledge.
This data does specifically not add anything new to this argument. As with almost all studies that test for a grid code in a similar "conceptual" space (not only the current study), the problem is that, when the space is not a uniform, square/circular space, and 2-dimensional then there is no reason the code will be perfectly grid like, i.e., show six-fold symmetry. In real world scenarios of social space (as well as navigation, semantic concepts), it must be higher dimensional - or at least more than two dimensional. It is unclear if this generalizes to larger spaces where not all part of the space is relevant. Modelling work from Tim Behrens' lab (e.g., Whittington et al., 2020) and Bradley Love's lab (e.g., Mok & Love, 2019) have shown/argued this to be the case. In experimental work, like in mazes from the Mosers' labs (e.g., Derdikman et al., 2009), or trapezoid environments from the O'Keefe lab (Krupic et al., 2015), there are distortions in mEC cells, and would not pass as grid cells in terms of the six-fold symmetry criterion.
After revision, the authors now discuss some of this and the limitations and notes that future work is required to address the problem.
-
-
www.medrxiv.org www.medrxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this study, the authors hypothesized that individuals with diabetes have elevated blood CTSL levels, which facilitates SARS-CoV-2 infection. The authors conducted in vitro experiments, revealing that elevated glucose levels promote SARS-CoV-2 infection in wild-type cells. In contrast, CTSL knockout cells show reduced susceptibility to high glucose-promoted effects. Additionally, the authors utilized lung tissue samples obtained from both diabetic and non-diabetic patients, along with db/db diabetic and control mice. Their findings indicate that diabetic conditions lead to an elevation in CTSL activity in both human and mice.
Strengths:
The authors have effectively met their research objectives, and their conclusions are supported by the data presented. Their findings suggest that high glucose levels promote CTSL maturation and translocation from the endoplasmic reticulum to the lysosome, potentially contributing to diabetic comorbidities and complications.
Weaknesses:
(1) In Figure 1e, the authors measured plasma levels of COVID-19 related proteins, including ACE2, CTSL, and CTSB, in both diabetic and non-diabetic COVID-19 patients. Notably, only CTSL levels exhibited a significant increase in diabetic patients compared to non-diabetic patients, and these levels varied throughout the course of COVID-19. Given that the diabetes groups encompass both male and female patients, it is essential to ascertain whether the authors considered the potential impact of gender on CTSL levels. The diabetes groups comprised a higher percentage of male patients (61.3%) compared to the non-diabetes group, where males constituted only 38.7%.
(2) lines145-149: "The results showed that WT Huh7 cell cultured in high glucose medium exhibited a much higher infective rate than those in low glucose medium. However, CTSL KO Huh7 cells maintained a low infective rate of SARS-CoV-2 regardless of glucose or insulin levels (Fig. 3f-h). Therefore, hyperglycemia enhanced SARS-CoV-2 infection dependent on CTSL." However, this evidence may be insufficient to support the claim that hyperglycemia enhances SARS-CoV-2 infection dependent on CTSL. The human hepatoma cell line Huh7 might not be an ideal model to validate the authors' hypothesis regarding high blood glucose promoting SARS-CoV-2 infection through CTSL.
(3) The Abstract and Introduction sections lack effective organization.
In this revised version of the study, the authors have addressed my concerns by providing additional experiments, references and discussing further the points of controversy. I think that the authors have made improvements to the manuscript.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Liu and colleagues describe the transcriptional changes observed during chloramphenicol-induced surface mobility of Bacillus subtilis. Practically, they describe that numerous transcriptional regulatory pathways are influenced by the subinhibitory concentration of a translational inhibitor and some of these regulatory changes might contribute to the induction of sliding. Nevertheless, how such translational stress is translated to induction of sliding remains undetermined. The authors clearly describe their aim (line 457): "Our goal for this study was to gain insight into how B. subtilis mobilizes a colony in response to subinhibitory exposure to translation inhibitors.", this is unfortunately not solved here, only the authors characterize the transcriptional landscape differences.
Strengths:
The very thorough analysis of transcriptional changes in the wild type and codY mutant strains is appreciated, and there are definitely a plethora of changes observed related to several global transcriptional regulators in B. subtilis. I compliment the authors for this very detailed and thorough description of transcriptional changes.
Weaknesses:
While the transcriptional changes are well and carefully described, the discussion practically interprets the correlations as causations. I am not disputing that the authors are not on the correct path with their assumptions, but their conclusions are not supported by direct experimental data, especially on (1) translational stress directly inducing mobility and (2) division of labor.
Major 1:
The authors conclude that their results point towards a putative mechanism, e.g. line 460 "which suggests translation stress is a trigger for colony mobilization"; however, no experiment demonstrates this aspect. The authors do not test ppGpp-related stress (mutants in ppGpp-related genes, or mutating the functional domain of CodY), nor do they directly connect ppGpp levels dynamics with induction of subsequent pathways. Again, I understand that the authors are on the right path to connect these pathways and identify what is causing mobility induction, but no direct data is represented, solely the transcriptional changes, therefore remains slightly descriptive.
The statement in the chapter title (line 474) is not demonstrated directly and should be revised. Similarly, in line 476, the authors claim that their "data supports a model", but "support" would require direct experimental data on this aspect.
The authors even clearly indicate in lines 504-506 that they do not reveal the direct mechanism, but the rest of the discussion delivers statements that do not consider the lack of direct data.
Major 2:
Line 427: "The results are consistent with a division of metabolic labor among cells in the expanding population" - the data shows heterogeneity, but the direct division of labor is not demonstrated.
Line 442: So in this case, the proposed division of labor is disrupted in the codY mutant (no inner localisation), and hence expansion appears, suggesting a lack of a putative division of labor is not necessary for induced mobility. On the contrary, there could be heterogeneous gene expression, division of labor requires demonstration of fitness benefit from such interaction.
Division of labor assumes that a mixture of mutants would complement full sliding dynamics, and this could be easily demonstrated by fluorescent labeled cells that should be organized in a similar fashion to those observed with luciferase reporters (pucA mutant on the outer ring, while pdhA mutant interior colony part). Without such experimental demonstration, the authors can only conclude spatially heterogeneous gene expresstion without clear functional contribution to subinhibitory chrolamphenociol-induced surface mobility.
Again, the authors' statement in line 472 "reveal a regulated, spatiotemporal division of metabolism" is not demonstrated by experiment, but spatial heterogeneity is revealed here.<br /> The statement in the Discussion chapter (line 499) is also not demonstrated by experimental data: "Metabolic coordination enables surface expansion of mobilitzed B. subtilis"
Line 550: while I agree with the authors' statement that these functions work cooperatively as demonstrated by van Gestel and colleagues (2015 PloS Biol), the exploitation of these shared goods is not quantitatively equivalent, see Jautzus et al 2022 ISME J (DOI: 10.1038/s41396-022-01279-8).
In summary: the two major conclusions of the manuscript are unfortunately not demonstrated, the presented transcriptional data delivers suggestions, supported with specific mutants displaying certain phenotypes (lack of mobility induction or constitutive mobility without inducer), but it remains unclear how translational stress induces mobility and whether the transcriptional heterogeneity detected directly contributes to metabolic division of labor.
The authors should present direct evidence on the major concerns: how translational stress induces surface mobility (using ppGpp synthesis and turnover mutants and specific CdoY mutant lacking ppGpp sensing) and whether the metabolic division of labor contributes to induced surface mobility (mixing mutants and following their distribution).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Secretion of the prototypical F-associated filamentous phage (Ff) of E. coli depends on the selective binding of a hairpin (the packaging signal, PS) by two phage encoded protein, pVII and pIX. PVII and pIX target the PS to IM channels formed by pI and pIV. However, integrative filamentous phages lack a homologue of pIX and pIV, and many of them also lack a homologue of pVII, raising questions on the assembly and secretion of new phages. In the manuscript, Yueh et al. present the identification of a phage-encoded protein, PSB15, which binds to the PS signal of a Xanthomonas integrative filamentous phage, ΦLf-UK. They showed that PSB15 is required for viral assembly and is conserved in several other integrative filamentous phages. They further analyzed how PSB15 binds to PS and demonstrated that it associates to the IM, which targets phage DNA to it. Finally, they show that thioredoxin, the only host protein that was found to be essential for Ff secretion, interacts with PSB15 and releases the PSB15-PS complex from the IM. These results are important because they elucidate a major step in the secretion of integrative filamentous phage, and the role of thioredoxin on filamentous phage secretion in general.
I found the data and interpretation convincing. However, the presentation and description are confusing in places because the reader has to juggle between figures. A scheme depicting what is known and unknown in the integration of Ff phages and interactive filamentous phages in the introduction would be useful to the general reader.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
In this study, Ruan et al. investigate the role of the IQCH gene in spermatogenesis, focusing on its interaction with calmodulin and its regulation of RNA-binding proteins. The authors examined sperm from a male infertility patient with an inherited IQCH mutation as well as Iqch CRISPR knockout mice. The authors found that both human and mouse sperm exhibited structural and morphogenetic defects in multiple structures, leading to reduced fertility in Ichq-knockout male mice. Molecular analyses such as mass spectrometry and immunoprecipitation indicated that RNA-binding proteins are likely targets of IQCH, with the authors focusing on the RNA-binding protein HNRPAB as a critical regulator of testicular mRNAs. The authors used in vitro cell culture models to demonstrate an interaction between IQCH and calmodulin, in addition to showing that this interaction via the IQ motif of IQCH is required for IQCH's function in promoting HNRPAB expression. In sum, the authors concluded that IQCH promotes male fertility by binding to calmodulin and controlling HNRPAB expression to regulate the expression of essential mRNAs for spermatogenesis. These findings provide new insight into molecular mechanisms underlying spermatogenesis and how important factors for sperm morphogenesis and function are regulated.
The strengths of the study include the use of mouse and human samples, which demonstrate a likely relevance of the mouse model to humans; the use of multiple biochemical techniques to address the molecular mechanisms involved; the development of a new CRISPR mouse model; ample controls; and clearly displayed results. Assays are done rigorously and in a quantitative manner. Overall, the claims made by the authors in this manuscript are well-supported by the data provided.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
The work proposes a model of neural information processing based on a 'synergistic global workspace,' which processes information in three principal steps: a gatekeeping step (information gathering), an information integration step, and finally, a broadcasting step. They provided an interpretation of the reduced human consciousness states in terms of the proposed model of brain information processing, which could be helpful to be implemented in other states of consciousness. The manuscript is well-organized, and the results are important and could be interesting for a broad range of literature, suggesting interesting new ideas for the field to explore.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The main conclusion of the manuscript is that the presence of linker Histone H1 protects Arabidopsis pericentromeric heterochromatic regions and longer transposable elements from encroachment by other repressive pathways. The manuscript focuses on the RNA-dependent DNA-methylation (RdDM) pathway but indirectly finds that other pathways must also be ectopically enriched.
Strengths:
The authors present diverse sets of genomic data comparing Arabidopsis wild-type and h1 mutant background allowing an analysis of differential recruitment of RdDM component NPRE1, which is related to changes in DNA methylation and H1 coverage. The manuscript also contains recruitment data for SUVH1 in wild-type and h1 mutant backgrounds.<br /> Furthermore, the authors make use of a line that recruits NRPE1 ectopically to show that H1 occupancy is not altered because of this recruitment. These data clearly show that there is a hierarchy in which DNA-methylation is impacted by presence of H1 while H1 distribution is independent of DNA-methylation.
Weaknesses:
The manuscript is driven by a strong and reasonable hypothesis that absence of H1 results increased access of chromatin binding factors and that this explains how the RdDM machinery is restricted from encroaching heterochromatic regions, which are particularly enriched in H1. Indeed, increased binding of NPRE1 at pericentromeric sites is observed; however, the major DNA-methylation changes at these sites are symmetric and not related to the RdDM pathway. Thus, the authors propose that many factors redistribute, which is again reasonable. The authors show redistribution of SUVH1 and relate their data to a previous report showing redistribution of the PcG machinery in H1 depletion mutants (Teano et al. in Cell reports (Volume 42, Issue 8, 29 August 2023), but the manuscript provides limited mechanistic insight as to why there is a strong increase in heterochromatin symmetric DNA-methylation.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This paper uses a novel maze design to explore mouse navigation behaviour in an automated analogue of the Barnes maze. A major strength is the novel and clever experimental design which rotates the floor and intramaze cues before the start of each new trial, allowing the previous goal location to become the next starting position. The modelling sampling a Markov chain of navigation strategies is elegant, appropriate and solid, appearing to capture the behavioural data well. This work provides a valuable contribution and I'm excited to see further developments, such as neural correlates of the different strategies and switches between them.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, Jiang et al., explore the role of neurexins at glycinergic MNTB-LSO synapses. The authors utilize elegant and compelling ex vivo slice electrophysiology to assess how the genetic conditional deletion of Nrxns1-3 impacts inhibitory glycinergic synaptic transmission and found that TKO of neurexins reduced electrically and optically evoked IPSC amplitudes, slowed optically evoked IPSC kinetics and reduced presynaptic release probability. The authors use classic approaches including reduced [Ca2+] in ACSF and EGTA chelation to propose that changes in these evoked properties are likely driven by the loss of calcium channel coupling. Intriguingly, while evoked transmission was impaired, the authors reported that spontaneous IPSC frequency was increased, due to an increase in the number of synapses in LSO. Overall, this manuscript provides important insight into the role of neurexins at the glycinergic MNTB-LSO synapse and further emphasizes the need for continued study of both the non-redundant and redundant roles of neurexins.
The authors have addressed all of my previous concerns.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Weng et al. perform a comprehensive study of gene expression changes in young and old animals, in wild-type and daf-2 insulin receptor mutants, in the whole animal and specifically in the nervous system. Using this data, they identify gene families that are correlated with neuronal ageing, as well as a distinct set of genes that are upregulated in neurons of aged daf-2 mutants. This is particularly interesting as daf-2 mutants show both extended lifespan and healthier neurons in aged animals, reflected by better learning/memory in older animals compared with wild-type controls. Indeed, knockdown of several of these upregulated genes resulted in poorer learning and memory. In addition, the authors showed that several genes upregulated during ageing in wild-type neurons also contribute to learning and memory; specifically, knockdown of these genes in young animals resulted in improved memory. This indicates that (at least in this small number of cases), genes that show increased transcript levels with age in the nervous system somehow suppress memory, potentially by having damaging effects on neuronal health.
Finally, from a resource perspective, the neuronal transcriptome provided here will be very useful for C. elegans researchers as it adds to other existing datasets by providing the transcriptome of older animals (animals at day 8 of adulthood) and demonstrating the benefits of performing tissue-specific RNAseq instead of whole-animal sequencing.
The work presented here is of high quality and the authors present convincing evidence supporting their conclusions. I only have a few comments/suggestions:
(1) Do the genes identified to decrease learning/memory capacity in daf-2 animals (Figure 4d/e) also impact neuronal health? daf-2 mutant worms show delayed onset of age-related changes to neuron structure (Tank et al., 2011, J Neurosci). Does knockdown of the genes shown to affect learning also affect neuron structure during ageing, potentially one mechanism through which they modulate learning/memory?
(2) The learning and memory assay data presented in this study uses the butanone olfactory learning paradigm, which is well established by the same group. Have the authors tried other learning assays when testing for learning/memory changes after knockdown of candidate genes? Depending on the expression pattern of these genes, they may have more or less of an effect on olfactory learning versus for e.g. gustatory or mechanosensory-based learning.
(3) A comment on the 'compensatory vs dysregulatory' model as stated by the authors on page 7 - I understand that this model presents the two main options, but perhaps this is slightly too simplistic: gene expression that rises during ageing may be detrimental for memory (= dysregulatory), but at the same time may also be beneficial other physiological roles in other tissues (=compensatory).
Comments on revised version:
I am satisfied with how the authors have addressed all my comments/suggestions.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary
The manuscript "Uncovering Protein Ensembles: Automated Multiconformer Model building for X-ray Crystallography and Cryo-EM" by Wankowicz et al. describes updates to qFit, an algorithm for the characterization of conformational heterogeneity of protein molecules based on X-ray diffraction of Cryo-EM data. The work provides a clear description of the algorithm used by qFit. The authors then proceed to validate the performance of qFit by comparing to deposited X-ray entries in the PDB in the 1.2-1.5 Å resolution range as quantified by Rfree, Rwork-Rfree, detailed examination of the conformations introduced by qFit, and performance on stereochemical measures (MolProbity scores). To examine the effect of experimental resolution of X-ray diffraction data, they start from an ultra high-resolution structure (SARS-CoV2 Nsp3 macrodomain) to determine how the loss of resolution (introduced artificially) degrades the ability of qFit to correctly infer the nature and presence of alternate conformations. The authors observe a gradual loss of ability to correctly infer alternate conformations as resolution degrades past 2 Å. The authors repeat this analysis for a larger set of entries in a more automated fashion and again observe that qFit works well for structures with resolutions better than 2 Å, with a rapid loss of accuracy at lower resolution. Finally, the authors examine the performance of qFit on cryo-EM data. Despite a few prominent examples, the authors find only a handful (8) of datasets for which they can confirm a resolution better than 2.0 Å. The performance of qFit on these maps is encouraging and will be of much interest because cryo-EM maps will, presumably, continue to improve and because of the rapid increase in the availability of such data for many supramolecular biological assemblies. As the authors note, practices in cryo-EM analysis are far from uniform, hampering the development and assessment of tools like qFit.
Strengths
qFit improves the quality of refined structures at resolutions better than 2.0 A, in terms of reflecting true conformational heterogeneity and geometry. The algorithm is well-designed and does not introduce spurious or unnecessary conformational heterogeneity. I was able to install and run the program without a problem within a computing cluster environment. The paper is well-written and the validation thorough.<br /> I found the section on cryo-EM particularly enlightening, both because it demonstrates the potential for discovery of conformational heterogeneity from such data by qFit, and because it clearly explains the hurdles towards this becoming common practice, including lack of uniformity in reporting resolution, and differences in map and solvent treatment.
Weaknesses
Due to limitations of past software engineering, the paper lacks a careful comparison to past versions of qFit. In light of the extensive assessment of the current version of qFit, this is a minor concern.
Although qFit can handle supramolecular assemblies and bound organic molecules, analysis in the manuscript is limited to single-chain X-ray structures. I look forward to demonstration of its utility in such cases in future work.
Appraisal & Discussion
Overall, the authors convincingly demonstrate that qFit provides a reliable means to detect and model conformational heterogeneity within high-resolution X-ray diffraction datasets and (based on a smaller sample) in cryo-EM density maps. This represents the state of the art in the field and will be of interest to any structural biologist or biochemist seeking to attain an understanding of the structural basis of the function of their system of interest, including potential allosteric mechanisms-an area where there are still few good solutions. That is, I expect qFit to find widespread use.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Severe leptospirosis in humans and some mammals often meet death in the endpoint. In this article, authors explored the role of the gut microbiota in severe leptospirosis. They found that Leptospira infection promoted a dysbiotic gut microbiota with an expansion of Proteobacteria and LPS neutralization therapy synergized with antileptospiral therapy significantly improved the survival rates in severe leptospirosis. This study is well-organized and has potentially important clinical implications not only for severe leptospirosis but also for other gut-damaged infections.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
This study uses a range of methods to characterize heterogeneous neural populations within the nucleus incertus (NI). The authors focus on two major populations, expressing gsc2 and rln3a, and present convincing evidence that these cells have different patterns of connectivity, calcium activity and effects on behavior. Although the study does not go as far as clarifying the role of NI in any specific neural computation or aspect of behavioral control, the findings will be valuable in support of future endeavors to do so. In particular, the authors have made two beautiful knock-in lines that recapitulate endogenous expression pattern of gsc2 and rln3a which will be a powerful tool to study the roles of the relevant NI cells. Experiments are well done, data are high quality and most claims are well supported. In this revised version, the authors have added additional analysis that has clarified their results and strengthened some of the claims.
Two points of note:
• The data very clearly show different patterns of neurites for gsc2 and rln3a neurons in the IPN and the authors interpret these are being axonal arbors. However, they do not rule out the possibility that some of the processes might be dendritic in nature. Of relevance to this point, they cite a recent study (Petrucco et al. 2023) that confirmed that, as in other species, tegmental neurons in zebrafish extend spatially segregated dendritic as well as axonal arbors into IPN, and the authors speculate that these GABAergic tegmental cells might in fact be part of NI.
• Although the gsc2 and rln3a populations show differences in calcium activity, there is not as clear a dichotomy as stated in the abstract. For example, both populations clearly respond to electric shocks, albeit with different response time courses.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Summary:
This study proposes visual homogeneity as a novel visual property that enables observers perform to several seemingly disparate visual tasks, such as finding an odd item, deciding if two items are same, or judging if an object is symmetric. In Exp 1, the reaction times on several objects were measured in human subjects. In Exp 2, visual homogeneity of each object was calculated based on the reaction time data. The visual homogeneity scores predicted reaction times. This value was also correlated with the BOLD signals in a specific region anterior to LO. Similar methods were used to analyze reaction time and fMRI data in a symmetry detection task. It is concluded that visual homogeneity is an important feature that enables observers to solve these two tasks.
Strengths:
(1) The writing is very clear. The presentation of the study is informative.<br /> (2) This study includes several behavioral and fMRI experiments. I appreciate the scientific rigor of the authors.
Weaknesses:
(1) My main concern with this paper is the way visual homogeneity is computed. On page 10, lines 188-192, it says: "we then asked if there is any point in this multidimensional representation such that distances from this point to the target-present and target-absent response vectors can accurately predict the target-present and target-absent response times with a positive and negative correlation respectively (see Methods)". This is also true for the symmetry detection task. If I understand correctly, the reference point in this perceptual space was found by deliberating satisfying the negative and positive correlations in response times. And then on page 10, lines 200-205, it shows that the positive and negative correlations actually exist. This logic is confusing. The positive and negative correlations emerge only because this method is optimized to do so. It seems more reasonable to identify the reference point of this perceptual space independently, without using the reaction time data. Otherwise, the inference process sounds circular. A simple way is to just use the mean point of all objects in Exp 1, without any optimization towards reaction time data.
(2) Visual homogeneity (at least given the current from) is an unnecessary term. It is similar to distractor heterogeneity/distractor variability/distractor statics in literature. However, the authors attempt to claim it as a novel concept. The title is "visual homogeneity computations in the brain enable solving generic visual tasks". The last sentence of the abstract is "a NOVEL IMAGE PROPERTY, visual homogeneity, is encoded in a localized brain region, to solve generic visual tasks". In the significance, it is mentioned that "we show that these tasks can be solved using a simple property WE DEFINE as visual homogeneity". If the authors agree that visual homogeneity is not new, I suggest a complete rewrite of the title, abstract, significance, and introduction.
(3) Also, "solving generic tasks" is another overstatement. The oddball search tasks, same-different tasks, and symmetric tasks are only a small subset of many visual tasks. Can this "quantitative model" solve motion direction judgment tasks, visual working memory tasks? Perhaps so, but at least this manuscript provides no such evidence. On line 291, it says "we have proposed that visual homogeneity can be used to solve any task that requires discriminating between homogeneous and heterogeneous displays". I think this is a good statement. A title that says "XXXX enable solving discrimination tasks with multi-component displays" is more acceptable. The phrase "generic tasks" is certainly an exaggeration.
(4) If I understand it correctly, one of the key findings of this paper is "the response times for target-present searches were positively correlated with visual homogeneity. By contrast, the response times for target-absent searches were negatively correlated with visual homogeneity" (lines 204-207). I think the authors have already acknowledged that the positive correlation is not surprising at all because it reflects the classic target-distractor similarity effect. But the authors claim that the negative correlations in target-absent searches is the true novel finding.
(5) I would like to make it clear that this negative correlation is not new either. The seminal paper by Duncan and Humphreys (1989) has clearly stated that "difficulty increases with increased similarity of targets to nontargets and decreased similarity between nontargets" (the sentence in their abstract). Here, "similarity between nontargets" is the same as the visual homogeneity defined here. Similar effects have been shown in Duncan (1989) and Nagy, Neriani, and Young (2005). See also the inconsistent results in Nagy& Thomas, 2003, Vicent, Baddeley, Troscianko&Gilchrist, 2009.<br /> More recently, Wei Ji Ma has systematically investigated the effects of heterogeneous distractors in visual search. I think the introduction part of Wei Ji Ma's paper (2020) provides a nice summary of this line of research.
I am surprised that these references are not mentioned at all in this manuscript (except Duncan and Humphreys, 1989).
(6) If the key contribution is the quantitative model, the study should be organized in a different way. Although the findings of positive and negative correlations are not novel, it is still good to propose new models to explain classic phenomena. I would like to mention the three studies by Wei Ji Ma (see below). In these studies, Bayesian observer models were established to account for trial-by-trial behavioral responses. These computational models can also account for the set-size effect, behavior in both localization and detection tasks. I see much more scientific rigor in their studies. Going back to the quantitative model in this paper, I am wondering whether the model can provide any qualitative prediction beyond the positive and negative correlations? Can the model make qualitative predictions that differ from those of Wei Ji's model? If not, can the authors show that the model can quantitatively better account for the data than existing Bayesian models? We should evaluate a model either qualitatively or quantitatively.
(7) In my opinion, one of the advantages of this study is the fMRI dataset, which is valuable because previous studies did not collect fMRI data. The key contribution may be the novel brain region associated with display heterogeneity. If this is the case, I would suggest using a more parametric way to measure this region. For example, one can use Gabor stimuli and systematically manipulate the variations of multiple Gabor stimuli, the same logic also applies to motion direction. If this study uses static Gabor, random dot motion, object images that span from low-level to high-level visual stimuli, and consistently shows that the stimulus heterogeneity is encoded in one brain region, I would say this finding is valuable. But this sounds like another experiment. In other words, it is insufficient to claim a new brain region given the current form of the manuscript.
REFERENCES<br /> - Duncan, J., & Humphreys, G. W. (1989). Visual search and stimulus similarity. Psychological Review, 96(3), 433-458. doi: 10.1037/0033-295x.96.3.433<br /> - Duncan, J. (1989). Boundary conditions on parallel processing in human vision. Perception, 18(4), 457-469. doi: 10.1068/p180457<br /> - Nagy, A. L., Neriani, K. E., & Young, T. L. (2005). Effects of target and distractor heterogeneity on search for a color target. Vision Research, 45(14), 1885-1899. doi: 10.1016/j.visres.2005.01.007<br /> - Nagy, A. L., & Thomas, G. (2003). Distractor heterogeneity, attention, and color in visual search. Vision Research, 43(14), 1541-1552. doi: 10.1016/s0042-6989(03)00234-7<br /> - Vincent, B., Baddeley, R., Troscianko, T., & Gilchrist, I. (2009). Optimal feature integration in visual search. Journal of Vision, 9(5), 15-15. doi: 10.1167/9.5.15<br /> - Singh, A., Mihali, A., Chou, W. C., & Ma, W. J. (2023). A Computational Approach to Search in Visual Working Memory.<br /> - Mihali, A., & Ma, W. J. (2020). The psychophysics of visual search with heterogeneous distractors. BioRxiv, 2020-08.<br /> - Calder-Travis, J., & Ma, W. J. (2020). Explaining the effects of distractor statistics in visual search. Journal of Vision, 20(13), 11-11.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Regalado et al. studied how an extended motivational state, necessary for maintaining behavioural drive despite unrewarding experiences, could be encoded in the ACC and its potential causal implications for learning discriminatory behaviour and avoiding unrewarding stimuli. They designed a self-initiated learning task and identified bulk neural responses tuned specifically to reward delivery as well as trial initiation. Interestingly, in both cases, neural activity precedes behavioural onset, indicating the encoding of a motivational signal. To investigate the neural encoding of motivational signals during unrewarded, distracting stimuli presentation, they created a discrimination task by introducing 'no reward' cues, during which animals need to learn not to reduce running speed and not engage in licking. Interestingly, with mice learning to increase running speed and reduce licking rates after 'no reward' cues, the preceding ACC activity also gradually increased. Importantly, only the increase in running speed after 'no reward' cues was impaired upon optogenetic inhibition of ACC activity during early training, linking the extended motivational signal in ACC and learning to maximise rewards by actively avoiding distracting and unrewarded stimuli. Such motivational signals could also be observed in OFC-ACC projecting neurons. Especially the continuous ramping of activity upon repeated 'non-reward' cues, which could be exclusively observed in the 'fast learner' subgroup, provides an interesting concept of how an extended motivational signal necessary for learning avoidance of unrewarded stimuli could be implemented in ACC. The shift in the temporal activity of initially reward-responsive neurons towards the preceding 'no reward' cue, provides a potential mechanism linking extended motivation to reward maximisation. This mechanism seems to be particularly important in periods of persistent 'non-reward' cues, as demonstrated in the impairment of running speed increase after two consecutive 'non-reward' cues.
Appraisal:
The authors provide convincing experimental evidence to support their claims of an extended motivational signal encoded in the ACC that is implemented by OFC-ACC signalling and critically involved in learning avoidance of unrewarded stimuli. The newly designed task seems appropriate to identify correlates of relevant cognitive and behavioural variables (e.g. sustained motivation). The combination of recording Ca2+ transients (bulk as well as longitudinal single neuron recordings) to identify potential neural responses and subsequent evaluation of their causal role in establishing and maintaining this persistent motivational state using opto- and pharmacogenetic manipulations is generally accepted.
Impact:
The findings will be valuable for further research on the impact of motivational states on behaviour and cognition. The authors provided a promising concept of how persistent motivational states could be maintained, as well as established a novel, reproducible task assay. While experimental methods used are currently state-of-the-art, theoretical analysis seems to be incomplete/not extensive.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors present a new model for animal pose estimation. The core feature they highlight is the model's stability compared to existing models in terms of keypoint drift. The authors test this model across a range of new and existing datasets. The authors also test the model with two mice in the same arena. For the single animal datasets the authors show a decrease in sudden jumps in keypoint detection and the number of undetected keypoints compared with DeepLabCut and SLEAP. Overall average accuracy, as measured by root mean squared error, generally shows similar but sometimes superior performance to DeepLabCut and better performance compared to SLEAP. The authors confusingly don't quantify the performance of pose estimation in the multi (two) animal case instead focusing on detecting individual identity. This multi-animal model is not compared with the model performance of the multi-animal mode of DeepLabCut or SLEAP.
Strengths:
The major strength of the paper is successfully demonstrating a model that is less likely to have incorrect large keypoint jumps compared to existing methods. As noted in the paper, this should lead to easier-to-interpret descriptions of pose and behavior to use in the context of a range of biological experimental workflows.
Weaknesses:
There are two main types of weaknesses in this paper. The first is a tendency to make unsubstantiated claims that suggest either model performance that is untested or misrepresents the presented data, or suggest excessively large gaps in current SOTA capabilities. One obvious example is in the abstract when the authors state ADPT "significantly outperforms the existing deep-learning methods, such as DeepLabCut, SLEAP, and DeepPoseKit." All tests in the rest of the paper, however, only discuss performance with DeepLabCut and SLEAP, not DeepPoseKit. At this point, there are many animal pose estimation models so it's fine they didn't compare against DeepPoseKit, but they shouldn't act like they did. Similar odd presentation of results are statements like "Our method exhibited an impressive prediction speed of 90{plus minus}4 frames per second (fps), faster than DeepLabCut (44{plus minus}2 fps) and equivalent to SLEAP (106{plus minus}4 fps)." Why is 90{plus minus}4 fps considered "equivalent to SLEAP (106{plus minus}4 fps)" and not slower? I agree they are similar but they are not the same. The paper's point of view of what is "equivalent" changes when describing how "On the single-fly dataset, ADPT excelled with an average mAP of 92.83%, surpassing both DeepLabCut and SLEAP (Figure 5B)" When one looks at Figure 5B, however, ADPT and DeepLabCut look identical. Beyond this, oddly only ADPT has uncertainty bars (no mention of what uncertainty is being quantified) and in fact, the bars overlap with the values corresponding to SLEAP and DeepPoseKit. In terms of making claims that seem to stretch the gaps in the current state of the field, the paper makes some seemingly odd and uncited statements like "Concerns about the safety of deep learning have largely limited the application of deep learning-based tools in behavioral analysis and slowed down the development of ethology" and "So far, deep learning pose estimation has not achieved the reliability of classical kinematic gait analysis" without specifying which classical gait analysis is being referred to. Certainly, existing tools like DeepLabCut and SLEAP are already widely cited and used for research.
The other main weakness in the paper is the validation of the multi-animal pose estimation. The core point of the paper is pose estimation and anti-drift performance and yet there is no validation of either of these things relating to multi-animal video. All that is quantified is the ability to track individual identity with a relatively limited dataset of 10 mice IDs with only two in the same arena (and see note about train and validation splits below). While individual tracking is an important task, that literature is not engaged with (i.e. papers like Walter and Couzin, eLife, 2021: https://doi.org/10.7554/eLife.64000) and the results in this paper aren't novel compared to that field's state of the art. On the other hand, while multi-animal pose estimation is also an important problem the paper doesn't engage with those results either. The two methods already used for comparison in the paper, SLEAP and DeepPoseKit, already have multi-animal modes and multi-animal annotated datasets but none of that is tested or engaged with in the paper. The paper notes many existing approaches are two-step methods, but, for practitioners, the difference is not enough to warrant a lack of comparison. The authors state that "The evaluation of our social tracking capability was performed by visualizing the predicted video data (see supplement Videos 3 and 4)." While the authors report success maintaining mouse ID, when one actually watches the key points in the video of the two mice (only a single minute was used for validation) the pose estimation is relatively poor with tails rarely being detected and many pose issues when the mice get close to each other.
Finally, particularly in the methods section, there were a number of places where what was actually done wasn't clear. For example in describing the network architecture, the authors say "Subsequently, network separately process these features in three branches, compute features at scale of one-fourth, one-eight and one-sixteenth, and generate one-eight scale features using convolution layer or deconvolution layer." Does only the one-eight branch have deconvolution or do the other branches also? Similarly, for the speed test, the authors say "Here we evaluate the inference speed of ADPT. We compared it with DeepLabCut and SLEAP on mouse videos at 1288 x 964 resolution", but in the methods section they say "The image inputs of ADPT were resized to a size that can be trained on the computer. For mouse images, it was reduced to half of the original size." Were different image sizes used for training and validation? Or Did ADPT not use 1288 x 964 resolution images as input which would obviously have major implications for the speed comparison? Similarly, for the individual ID experiments, the authors say "In this experiment, we used videos featuring different identified mice, allocating 80% of the data for model training and the remaining 20% for accuracy validation." Were frames from each video randomly assigned to the training or validation sets? Frames from the same video are very correlated (two frames could be just 1/30th of a second different from each other), and so if training and validation frames are interspersed with each other validation performance doesn't indicate much about performance on more realistic use cases (i.e. using models trained during the first part of an experiment to maintain ids throughout the rest of it.)
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The paper from Li et al shows a mechanism by which axons can change direction during development. They use the sLNv neurons as a model. They find that the appearance of a new group of neurons (DNs) during post-embryonic proliferation secretes netrins and repels horizontally towards the midline, the axonal tip of the LNvs.
Strengths:
The experiments are well done and the results are conclusive.
Weaknesses:
The novelty of the study is overstated, and the background is understated. Both things need to be revised.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Rashid and colleagues demonstrate a novel hippocampal lateral septal circuit that is important for social recognition and drives the exploration of novel conspecifics. Their study spans from neural tracing to close-loop optogenetic experiments with clever controls and conditions to provide compelling evidence for their conclusion. They demonstrate that downstream of the hippocampal septal circuit, septal projections to the ventral tegmental area are necessary for general novelty discrimination. The study opens an avenue to study these circuits further to uncover the plasticity and synaptic mechanisms regulating social novelty preference.
Strengths:
Chemogenetic and optogenetic experiments have excellent behavioral controls. The synaptic tracing provides important information that informs the narrative of experiments presented and invites future studies to investigate the effects of septal input on dopaminergic activity.
Weaknesses:
There are unclear methodological important details for circuit manipulation experiments and analyses where multiple measures are needed but missing. Based on the legends, the chemogenetic experiment is done in a within-animal design. That is the same mouse receives SAL and CNO. However, the data is not presented in a within-animal manner such that we can distinguish if the behavior of the same animal changes with drug treatment. Similarly, the methods specify that the optogenetic manipulations were done in three different conditions, but the analyses do not report within-animal changes across conditions nor account for multiple measures within subjects. Finally, it is unclear if the order of drug treatment and conditions were counterbalanced across subjects.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this work, the authors show that GABAergic neurons play a role in sensing mitochondrial stress and regulating organismal aging. Thus, disrupting the mitochondrial mitochondria function in GABAergic neurons induces resistance to thermal and paraquat stresses, promotes longevity, and affects reproduction. This mechanism is regulated by the iron-sulfur subunit of complex III of the mitochondrial electron transport chain, ISP-1, and a mitochondrial quality control m-AAA protease, SPG-7, which in turn requires DAF-16/FoxO activity in GABAergic neurons.
Strengths:
A strength of this work is that the authors identify the specific site where mitochondrial stress promotes health and longevity, i.e., GABAergic neurons. In addition, the paper corroborates the findings with the appropriate experiments. How neuronal regulation of mitochondrial function impacts systemic health and aging is of interest to cell biology and neuroscience fields.
Weaknesses:
The entire paper is based on tissue-specific RNAi in GABAergic neurons, which was achieved using two different conditions of RNAi (although not for all experiments). However, multiple studies have shown deficiencies in the tissue-specific RNAi in C. elegans, especially for the rde-1(ne219) mutant used in this study. Therefore, it is necessary to repeat critical experiments by rescuing the isp-1 or spg-7 mutants in GABAergic neurons. Additionally, it is clear in the paper that perturbing mitochondrial function requires DAF-16/FoxO activity in GABAergic neurons to promote longevity, yet the downstream cellular pathways are not described.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In the manuscript, Xiang Mou and Daoyun JI investigate how ACC neurons activated by observational learning communicate with the hippocampus. They assess this line of communication through a complex behavioral technique, in vivo electrophysiology, pharmacological approaches, and data analytical techniques. Firstly, the authors find that observational performance is dependent on the ACC, and that the ACC possesses neurons that show side selectivity (trajectory-related) in both the observation box when shuttling to reward, and during subsequent maze running, shuttling to the corresponding same side for reward. The side-selective activation appears stronger for correct trials compared to error trials specifically during observation of Demo rats. They compare how the CA1 of the hippocampus encodes these two environments and find that ACC side-selective neurons show a correlation with side-selective CA1 ensembles during maze behavior, water consumption, and sharp-wave ripples.
Strengths:
Overall, the paper provides strong evidence that ACC neurons are activated by observational learning and that this activation seems to be correlated with CA1 activity.
Weaknesses:
Concerns, however, surround the strength of evidence that links ACC and CA1 activity during observational learning. Only weak correlations between the two regions are shown, and it is unclear if the ACC may lead to CA1 activity or vice versa. It is possible that these processes reflect two parallel pathways. Without manipulation of ACC, it is difficult to assess whether ACC activity influences hippocampal replay.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this study, the authors aim to identify the nuclear genome-encoded transcription factors that regulate mtDNA maintenance and mitochondrial biogenesis. They started with an RNAi screening in developing Drosophila eyes with reduced mtDNA content and identified a number of putative candidate genes. Subsequently, using ChIP-seq data, they built a potential regulatory network that could govern mitochondrial biogenesis. Next, they focused on a candidate gene, CG1603, for further characterization. Based on the expression of different markers, such as TFAM and SDHA, in the RNAi and OE clones in the midgut cells, they argue that CG1603 promotes mitochondrial biogenesis and the expression of ETC complex genes. Then, they used a mutant of CG1603 and showed that both mtDNA levels and mitochondrial protein levels were reduced. Using clonal analyses, they further show a reduction in mitochondrial biogenesis and membrane potential upon loss of CG1603. They made a reporter line of CG1603, showed that the protein is localized to the mitochondria, and binds to polytene chromosomes in the salivary gland. Based on the RNA-seq results from the mutants and the ChIP data, the authors argue that the nucleus-encoded mitochondrial genes that are downregulated >2 folds in the CG1603 mutants and that are bound by CG1603 are related to ETC biogenesis. Finally, they show that YL-1, another candidate in the network, is an upstream regulator of CG1603.
Strengths:
This is a valuable study, which identifies a potential regulator and a network of nucleus-encoded transcription factors that regulate mitochondrial biogenesis. Through in-vivo and in-vitro experimental evidence, the authors identify the role of CG1603 in this process. The screening strategy was smart, and the follow-up experiments were nicely executed.
Weaknesses:
Some additional experiments showing the effects of CG1603 loss on ETC integrity and functionality would strengthen the work.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The study by Deganutti and co-workers is a methodological report on an adaptive sampling approach, multiple walker supervised molecular dynamics (mwSuMD), which represents an improved version of the previous SuMD.
Case-studies concern complex conformational transitions in a number of G protein Coupled Receptors (GPCRs) involving long time-scale motions such as binding-unbinding and collective motions of domains or portions. GPCRs are specialized GEFs (guanine nucleotide exchange factors) of heterotrimeric Gα proteins of the Ras GTPase superfamily. They constitute the largest superfamily of membrane proteins and are of central biomedical relevance as privileged targets of currently marketed drugs.
MwSuMD was exploited to address:<br /> (1) Binding and unbinding of the arginine-vasopressin (AVP) cyclic peptide agonist to the V2 vasopressin receptor (V2R);<br /> (2) Molecular recognition of the β2-adrenergic receptor (β2-AR) and heterotrimeric GDP-bound Gs protein;<br /> (3) Molecular recognition of the A1-adenosine receptor (A1R) and palmitoylated and geranylgeranylated membrane-anchored heterotrimeric GDP-bound Gi protein;<br /> (4) The whole process of GDP release from membrane-anchored heterotrimeric Gs following interaction with the glucagon-like peptide 1 receptor (GLP1R), converted to the active state following interaction with the orthosteric non-peptide agonist danuglipron;<br /> (5) The heterodimerization of D2 dopamine and A2A adenosine receptors (D2R and A2AR, respectively) and binding to a bi-valent ligand.
The mwSuMD method is solid and valuable, has wide applicability, and is compatible with the most world-widely used MD engines. It may be of interest to the computational structural biology community.
The huge amount of high-resolution data on GPCRs makes those systems suitable, although challenging, for method validation and development.
While the approach is less energy-biased than other enhanced sampling methods, knowledge, at the atomic detail, of binding sites/interfaces and conformational states is needed to define the supervised metrics, the higher the resolution of such metrics is the more accurate the outcome is expected to be. The definition of the metrics is a user- and system-dependent process.
The too many and ambitious case-studies undermine the accuracy of the output and reduce the important details needed for a methodological report. In some cases, the available CryoEM structures could have been exploited better.
The most consistent example concerns AVP binding/unbinding to V2R. The consistency with CryoEM data decreases with an increase in the complexity of the simulated process and involved molecular systems (e.g. receptor recognition by membrane-anchored G protein and the process of nucleotide exchange starting from agonist recognition by an inactive-state receptor). The last example, GPCR hetero-dimerization, and binding to a bi-valent ligand, is the most speculative one as it does not rely on high-resolution structural data for metrics supervision.
-
-
-
Reviewer #2 (Public Review):
Summary:
The current article presents a new type of analytical approach to the sequential organisation of whale coda units.
Strengths:
The detailed description of the internal temporal structure of whale codas is something that has been thus far lacking.
Weaknesses:
It is unclear how the insight gained from these analyses differs or adds to the voluminous available literature on how codas varies between whale groups and populations. It provides new details, but what new aspects have been learned, or what features of variation seem to be only revealed by this new approach?<br /> The theoretical basis and concepts of the paper are problematical and indeed, hamper potentially the insights into whale communication that the methods could offer. Some aspects of the results are also overstated.
-
- May 2024
-
52.53.155.43 52.53.155.43
-
Reviewer #2 (Public Review):
Summary:
The manuscript by Djebar et al investigated the role and the underlying mechanism of the ciliary transition zone protein Rpgrip1l in zebrafish spinal alignment. They showed that rpgrip1l mutant zebrafish develop a nearly full penetrance of body curvature at juvenile stages. The mutant fish have cilia defects associated with ventricular dilations and loss of the Reissner fibers. Scoliosis onset and progression are also strongly associated with astrogliosis and neuroinflammation, and anti-inflammatory drug treatment prevents scoliosis in mutant zebrafish, suggesting a novel pathogenic mechanism for human idiopathic scoliosis. This study is quite comprehensive with high-quality data, and the manuscript is well written, providing important information on how the ciliary transition zone protein functions in maintaining the zebrafish body axis straightness.
Strengths:
Very clear and comprehensive analysis of the mutant zebrafish.
Weaknesses:
(1) In Figures 1D-G, magnified high-resolution pictures are required to show there are indeed no vertebral malformations.
(2) Are the transcriptome data and proteomic data consistent? Consistent targets in both analyses should be highlighted.
(3) What is the role of Anxa2 in neuroinflammation? Is increased Anxa2 expression in rpgrip1l mutant zebrafish reduced after anti-inflammatory drug treatment? What is the expression level of anxa2 in cep290 mutant zebrafish?
(4) More background about Rpgrip1l should be provided in the introduction, particularly the past studies of the mammalian homolog of Rpgrip11, if there are any.
(5) Is there any human disease associated with Rpgrip1l? Do these patients have scoliosis phenotype?
(6) A summary diagram at the end would be helpful for understanding the main findings.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors provide compelling evidence that stimulation of epidermal cells in Drosophila larvae results in the stimulation of sensory neurons that evoke a variety of behavioral responses. Further, the authors demonstrate that epidermal cells are inherently mechanoresponsive and implicate a role for store-operated calcium entry (mediated by Stim and Orai) in the communication to sensory neurons.
Strengths:
The study represents a significant advance in our understanding of mechanosensation. Multiple strengths are noted. First, the genetic analyses presented in the paper are thorough with appropriate consideration to potential confounds. Second, behavioral studies are complemented by sophisticated optogenetics and imaging studies. Third, identification of roles for store-operated calcium entry is intriguing. Lastly, conservation of these pathways in vertebrates raise the possibility that the described axis is also functional in vertebrates.
Weaknesses:
The study has a few conceptual weaknesses that are arguably minor. The involvement of store-operated calcium entry implicates ER calcium store release. Whether mechanical stimulation evokes ER calcium release in epidermal cells and how this might come about (e.g., which ER calcium channels, roles for calcium-induced calcium release etc.) remains unaddressed. On a related note, the kinetics of store-operated calcium entry is very distinct from that required for SV release. The link between SOC and epidermal cells-neuron transmission is not reconciled. Finally, it is not clear how optogenetic stimulation of epidermal cells results in the activation of SOC.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The manuscript titled, "Identification of a Musashi2 translocation as a novel oncogene in myeloid leukemia" by Spinler et al. studies the functional role of the translocation t(7;17)(p15;q23), resulting in MSI2/HOXA9 fusion gene, as a secondary driver in bcCML. MSI2-HOXA9 forced expression along with BCR-ABL enhances colony formation and leads to a more aggressive disease in vivo. Depletion of the RNA binding domain RRM1 or RRM2 of MSI2 led to a significant reduction in colony formation, with RRM1 depletion specifically impacting differentiation and blast cell counts. Mechanistically, the authors find that MSI2-HOXA9 aberrantly localizes to the nucleus, elevating the expression of mitochondrial polymerase Polrmt, thereby leading to upregulation of mitochondrial components and enhancing mitochondrial function and basal respiration. Overall, this study examines how the rare MSI2-HOXA9 fusion gene can act as a novel cooperating oncogene and could serve as a secondary hit in the progression of CML to blast crisis.
Strengths:
(1) Demonstration that MSI2-HOXA9 contributes to oncogenesis in the BCR-ABL context.
(2) Development of a novel cooperativity model for BCR-ABL and provides additional supporting data for the role of MSI2 in leukemogenesis.
(3) Evidence that MSI2-HOXA9 acts uniquely compared to MSI2 alone through nuclear vs. cytoplasmic localization and activation of mitochondrial polymerase Polrmt.
Weaknesses:
(1) MSI2-HOXA9 fusion is extremely rare as it has been only found in a handful of patients and it is not clear whether other MSI2 fusions function in a similar manner.
(2) The mechanism needs to be strengthened since MSI2 alone or the HOXA9 mutant may not be linked to the mitochondrial mechanism.
(3) It is not clear that the mitochondrial pathway is sufficient for the MSI2-HOXA9 oncogenic mechanism.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Through RNA analysis, Xie et al found LncRNA Snhg3 was one of the most down-regulated Snhgs by a high-fat diet (HFD) in mouse liver. Consequently, the authors sought to examine the mechanism through which Snhg3 is involved in the progression of metabolic dysfunction-associated fatty liver diseases (MASLD) in HFD-induced obese (DIO) mice. Interestingly, liver-specific Sngh3 knockout was reduced, while Sngh3 over-expression potentiated fatty liver in mice on an HFD. Using the RNA pull-down approach, the authors identified SND1 as a potential Sngh3 interacting protein. SND1 is a component of the RNA-induced silencing complex (RISC). The authors found that Sngh3 increased SND1 ubiquitination to enhance SND1 protein stability, which then reduced the level of repressive chromatin H3K27me3 on PPARg promoter. The upregulation of PPARg, a lipogenic transcription factor, thus contributed to hepatic fat accumulation.
The authors propose a signaling cascade that explains how LncRNA sngh3 may promote hepatic steatosis. Multiple molecular approaches have been employed to identify molecular targets of the proposed mechanism, which is a strength of the study. There are, however, several potential issues to consider before jumping to a conclusion.
(1) First of all, it's important to ensure the robustness and rigor of each study. The manuscript was not carefully put together. The image qualities for several figures were poor, making it difficult for the readers to evaluate the results with confidence. The biological replicates and numbers of experimental repeats for cell-based assays were not described. When possible, the entire immunoblot imaging used for quantification should be presented (rather than showing n=1 representative). There were multiple mislabels in figure panels or figure legends (e.g., Figure 2I, Figure 2K, and Figure 3K). The b-actin immunoblot image was reused in Figure 4J, Figure 5G, and Figure 7B with different exposure times. These might be from the same cohort of mice. If the immunoblots were run at different times, the loading control should be included on the same blot as well.
(2) The authors can do a better job in explaining the logic for how they came up with the potential function of each component of the signaling cascade. Sngh3 is down-regulated by HFD. However, the evidence presented indicates its involvement in promoting steatosis. In Figure 1C, one would expect PPARg expression to be up-regulated (when Sngh3 was down-regulated). If so, the physiological observation conflicts with the proposed mechanism. In addition, SND1 is known to regulate RNA/miRNA processing. How do the authors rule out this potential mechanism? How about the hosting snoRNA, Snord17? Does it involve the progression of NASLD?
(3) The role of PPARg in fatty liver diseases might be a rodent-specific phenomenon. PPARg agonist treatment in humans may actually reduce ectopic fat deposition by increasing fat storage in adipose tissues. The relevance of the findings to human diseases should be discussed.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors provide an interesting observation that ER-targeted excess misfolded proteins localize to the nucleus within membrane-entrapped vesicles for further quality control during cell division. This is useful information indicating transient nuclear compartmentalization as a quality control strategy for misfolded ER proteins in mitotic cells, although endogenous substrates of this pathway are yet to be identified.
Strengths:
This microscopy-based study reports unique membrane-based compartments of ER-targeted misfolded proteins within the nucleus. Quarantining aggregating proteins in membrane-less compartments is a widely accepted protein quality control mechanism. This work highlights the importance of membrane-bound quarantining strategies for aggregating proteins. These observations open up multiple questions on proteostasis biology. How do these membrane-bound bodies enter the nucleus? How are the single-layer membranes formed? How exactly are these membrane-bound aggregates degraded? Are similar membrane-bound nuclear deposits present in post-mitotic cells that are relevant in age-related proteostasis diseases? Etc. Thus, the observations reported here are potentially interesting.
Weaknesses:
This study, like many other studies, used a set of model misfolding-prone proteins to uncover the interesting nuclear-compartment-based quality control of ER proteins. The endogenous ER-proteins that reach a similar stage of overdose of misfolding during ER stress remain unknown.
The mechanism of disaggregation of membrane-trapped misfolded proteins is unclear. Do these come out of the membrane traps? The authors report a few vesicles in living cells. This may suggest that membrane-untrapped proteins are disaggregated while trapped proteins remain aggregates within membranes.
The authors figure out the involvement of proteasome and Hsp70 during the disaggregation process. However, the detailed mechanisms including the ubiquitin ligases are not identified. Also, is the protein ubiquitinated at this stage?
This paper suffers from a lack of cellular biochemistry. Western blots confirming the solubility and insolubility of the misfolded proteins are required. This will also help to calculate the specific activity of luciferase more accurately than estimating the fluorescence intensities of soluble and aggregated/compartmentalized proteins. Microscopy suggested the dissolution of the membrane-based compartments and probably disaggregation of the protein. This data should be substantiated using Western blots. Degradation can only be confirmed by Western blots. The authors should try time course experiments to correlate with microscopy data. Cycloheximide chase experiments will be useful.
The cell models express the ER-targeted misfolded proteins constitutively that may already reprogram the proteostasis. The authors may try one experiment with inducible overexpression.
It is clear that a saturating dose of ER-targeted misfolded proteins activates the pathway. The authors performed a few RT-PCR experiments to indicate the proteostasis-sensitivity. Proteome-based experiments will be better to substantiate proteostasis saturation.
The authors should immunostain the nuclear compartments for other ER-membrane resident proteins that span either the bilayer or a single layer. The data may be discussed.
All microscopy figures should include control cells with similarly aggregating proteins or without aggregates as appropriate. For example, is the nuclear-targeted FlucDM-EGFP similarly entrapped? A control experiment will be interesting. Expression of control proteins should be estimated by western blots.
There are few more points that may be out of the scope of the manuscript. For example, how do these compartments enter the nucleus? Whether similar entry mechanisms/events are ever reported? What do the authors speculate? Also, the bilayer membrane becomes a single layer. This is potentially interesting and should be discussed with probable mechanisms. Also, do these nuclear compartments interfere with transcription and thereby deregulate cell division? What about post-mitotic cells? Similar deposits may be potentially toxic in the absence of cell division. All these may be discussed.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This important work will be of interest to centriole and cilia cell biologists. It describes in detail how microtubules control multiple aspects of centriole amplification in brain multiciliated cells. This study provides a greater time-resolved and molecular proteomic mapping of the different steps involved, with or without microtubule disruption. Boudjema et al. show that microtubules are important throughout the centriole amplification process, from the early stages, where the procentrioles emerge from a pericentriolar "nest", through the growth stage where microtubules maintain the perinuclear localisation, to the detachment stage, where microtubules assist in perinuclear disengagement and apical migration. The results are generally well supported by the evidence, but the manuscript would benefit significantly from some heavy editing to introduce more niche terms, standardize abbreviations in text, and labels on figures to help bring the readers, especially non-specialists, along with them - increasing the accessibility of their work.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this work, the authors investigate the role of the Superoxide disumutase 1 (Sod1) enzyme, which acts to reduce the reactive oxygen species load, in the Drosophila testis. They show that the knockdown of Sod1 in somatic cells impacts stem cell numbers both autonomously in the soma and non-autonomously in the germline. Somatic stem cell numbers are increased, while germline stem cells are decreased and differentiate prematurely. The authors then show that in somatic Sod1 knockdowns, several signalling pathways are disrupted and that these may be responsible at least in part for the phenotypes observed. Finally, over-expression of Sod1 in the soma results in opposite phenotypes, suggesting that ROS levels do play a role in maintaining the balance between both stem cell populations in the testis.
Strengths:
The main strength of this work is to show a previously unappreciated role for Sod1 (and presumably by extension of ROS) in the Drosophila testis and in the regulation of stem cell self-renewal and differentiation. The authors use multiple readouts to show that the knockdown of Sod1 in the soma increases the number of somatic cells and also drives a non-autonomous, premature differentiation of germ cells. They also quantify the early differentiation of the germline using two different methods. Importantly, overexpression of Sod1 produces opposite phenotypes to knockdown, strengthening the conclusions.
Weaknesses:
Although the data presented are interesting, an important weakness of the manuscript as it currently stands is that many statements are not fully supported by the data. In particular, the authors do not provide any evidence of "cell redox-pairs" as indicated in the manuscript title, nor of intercellular redox gradients, as stated in several places throughout. While the data are consistent with non-autonomous regulation of ROS levels, this would not constitute a gradient. However, and crucially, there is no evidence provided to show that Sod1 manipulation in the soma is affecting ROS levels in the germline and that any of the phenotypes observed are a consequence of elevated ROS in the germline, rather than indirect effects caused by dysregulation of somatic self-renewal and differentiation, which is known to impact the germline. Indeed, there are many published reports of autonomous manipulations in the soma that influence either germline stem cell number (eg PMID: 19797664 among others) or differentiation (eg PMID: 17629483). The latter example is particularly relevant as the authors show altered somatic ERK levels, and the role of somatic ERK in promoting germ cell development is well established (PMID: 11048722, 11048723,...). Thus, whether Sod1 plays any non-autonomous role in controlling germ cell fate through ROS in the germline directly, or whether the phenotypes observed can all be explained by autonomous effects on somatic cell behaviour is debatable, but the experiments presented here do not distinguish between these two hypotheses. The only evidence presented by the authors for a non-autonomous role of Sod1 is the expression of a GFP reporter for gstD1. The quantifications and images are not clear and do not show unambiguously that this reporter is expressed in germ cells. Indeed, the quantifications show overlap between somatic and germline markers, suggesting that either the images themselves or the way they are quantified does not allow the authors to distinguish between the two cell types. Similarly, the claim that somatic mitochondria are enriched at the CySC-GSC interface and that this distribution maintains the redox balance in the niche is not supported by any experimental data. CySCs are extremely thin cells and much of the space is occupied by the nucleus (PMID: 114676), therefore it is likely that mitochondria would be enriched at the periphery. A careful analysis would be necessary to show that this enrichment is specific to the interface with GSCs. Moreover, no experiments are conducted to test whether mitochondrial distribution in CySCs has any impact on GSCs. Finally, no experiments are conducted to show definitively that the phenotypes observed upon Sod1 knockdown are indeed due to increased ROS, while this claim is made several times in the text. At present, the data presented here can support a role for Sod1 in somatic CySCs, but much more caution is required in attributing this to either ROS or intercellular ROS signaling. Therefore, several claims made in the title and throughout the text are not supported by evidence.
Besides this central point, there are other areas that should be improved. In particular, the data using the Fucci reporter to show accelerated proliferation do not appear convincing. It would seem that the proportions of cells in each phase are roughly similar, just that there are more cycling cells. A careful analysis of these results would distinguish between these two and determine whether Sod1 knockdown simply impairs differentiation (and therefore results in more somatic cells proliferating) or whether it speeds up the cell cycle (resulting in an increased mitotic index as suggested, but this requires a ratio to be shown). Similarly, several quantifications are not clearly explained, making it hard to understand what is being measured. As an example, while the decrease in pERK in CySCs is clear from the image and matched in the quantification, the increase in cyst cells is not apparent from the fire LUT used. The change in fluorescence intensity therefore may be that more cells have active ERK, rather than an increase per cell (similar arguments apply to the quantifications for p4E-BP or Ptc). Therefore, it is hard to know whether Sod1 knockdown results in increased or decreased signaling in individual cells.
Impact of study:
Demonstrating intercellular communication through ROS and its importance in maintaining the balance between two stem cell populations would be a finding of interest to a broad field. However, it remains to be demonstrated that this is the case, and given this, this study will have a limited impact.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The mechanism of microtubule formation, stabilization, and organization in neurites is important for neuronal function. In this manuscript, the authors examine the phenotype of neurons following alteration in the level of the protein HMMR, a microtubule-associated protein with established roles in mitosis. Neurite morphology is measured as well as microtubule stability and dynamic parameters using standard assays. A binding partner of HMMR, TPX2, is localized. The results support a role for HMMR in microtubule stabilization in neurons.
The results show that HMMR is distributed as puncta on neurons using standard immunofluorescence and PLA. Depletion of HMMR reduced neurite length and extent of branching; reduced post-translational acetylation of neurite microtubules. Conversely, overexpression of HMMR increased resistance to nocodazole. The parameters of microtubule dynamics were also impacted by reduction or overexpression of HMMR. The authors discuss the possibility HMMR regulates neurite morphological changes via regulation of microtubule nucleation and dynamics.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Summary:
The authors aim to demonstrate the effectiveness of their developed methodology, which utilizes super-resolution microscopy and single-molecule tracking in live cells on a high-throughput scale. Their study focuses on measuring the diffusion state of a molecule target, the estrogen receptor, in both ligand-bound and unbound forms in live cells. By showcasing the ability to screen 5067 compounds and measure the diffusive state of the estrogen receptor for each compound in live cells, they illustrate the capability and power of their methodology.
Readers are well introduced to the principles in the initial stages of the manuscript with highly convincing video examples. The methods and metrics used (fbound) are robust. The authors demonstrate high reproducibility of their screening method (R2=0.92). They also showcase the great sensitivity of their method in predicting the proliferation/viability state of cells (R2=0.84). The outcome of the screen is sound, with multiple compounds clustering identified in line with known estrogen receptor biology.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The manuscript submitted by Sujeethkumar et al. describes an alternative approach to skeletal tissue repair using extracellular matrix (ECM) deposited by genetically modified mesenchymal stromal/stem cells. Here, they generate a loss of function mutations in VEGF or RUNX2 in a BMP2-overexpressing MSC line and define the differences in the resulting tissue-engineered constructs following seeding onto a type I collagen matrix in vitro, and following lyophilization and subcutaneous and orthotopic implantation into mice and rats. Some strengths of this manuscript are the establishment of a platform by which modifications in cell-derived ECM can be evaluated both in vitro and in vivo, the demonstration that genetic modification of cells results in complexity of in vitro cell-derived ECM that elicits quantifiable results, and the admirable goal to improve endogenous cartilage repair. However, I recommend the authors clarify their conclusions and add more information regarding reproducibility, which was one limitation of primary-cell-derived ECMs.
Overcoming the limitations of native/autologous/allogeneic ECMs such as complete decellularization and reduction of batch-to-batch variability was not specifically addressed in the data provided herein. For the maintenance of ECM organization and complexity following lyophilization, evidence of complete decellularization was not addressed, but could be easily evaluated using polarized light microscopy and quantification of human DNA for example in constructs pre and post-lyophilization. It would be ideal to see minimization of batch-to-batch variability using this approach, as mitigation of using a sole cell line is likely not sufficient (considering that the sole cell line-derived Matrigel does exhibit batch-to-batch and manufacturer-to-manufacturer variability).
I recommend adding details regarding experimental design and outcomes not initially considered. Inter- and intra-experimental reproducibility was not adequately addressed. The size of in vitro-derived cartilage pellets was not quantified, and it is not clear that more than one independent 'differentiation' was performed from each gene-edited MSC line to generate in vitro replicates and constructs that were implanted in vivo.
The use of descriptive language in describing conclusions may mislead the reader and should be modified accordingly throughout the manuscript. For example, although this reviewer agrees with the comparative statements made by the authors regarding parental and gene-edited MSC lines, non-quantifiable terms such as 'frank' 'superior' (example, line 242) are inappropriate and should rather be discussed in terms of significance. Another example is 'rich-collagenous matrix,' which was not substantiated by uniform immunostaining for type II collagen (line 189).
I have similar recommendations regarding conclusive statements from the rat implantation model, which was appropriately used for the purpose of evaluating the response of native skeletal cells to the different cell-derived ECMs. Interpretations of these results should be described with more accuracy. For example, increased TRAP staining does not indicate reduced active bone formation (line 237). Many would not conclude that GAGs were retained in the RUNX2-KO line graft subchondral region based on the histology. Quantification of % chondral regeneration using histology is not accurate as it is greatly influenced by the location in the defect from which the section was taken. Chondral regeneration is usually semi-quantified from gross observations of the cartilage surface immediately following excision. The statements regarding integration (example line 290) are not founded by histological evidence, which should show high magnification of the periphery of the graft adjacent to the native tissue.
-
-
www.medrxiv.org www.medrxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors investigated the effects of the timing of dietary occasions on weight loss and well-being with the aim of explaining if a consistent, timely alignment of dietary occasions throughout the days of the week could improve weight management and overall well-being. The authors attributed these outcomes to a timely alignment of dietary occasions with the body's own circadian rhythms. However, the only evidence the authors provided for this hypothesis is the assumption that the individual timing of dietary occasions of the study participants identified before the intervention reflects the body's own circadian rhythms. This concept is rooted in understanding of dietary cues as a zeitgeber for the circadian system, potentially leading to more efficient energy use and weight management. Furthermore, the primary outcome, body weight loss, was self-reported by the study participants.
Strengths:
The innovative focus of the study on the timing of dietary occasions rather than daily energy intake or diet composition presents a fresh perspective in dietary intervention research. The feasibility of the diet plan, developed based on individual profiles of the timing of dietary occasions identified before the intervention, marks a significant step towards personalised nutrition.
Weaknesses:
Several methodological issues detract from the study's credibility, including unclear definitions not widely recognized in nutrition or dietetics (e.g., "caloric event"), lack of comprehensive data on body composition, and potential confounders not accounted for (e.g., age range, menstrual cycle, shift work, unmatched cohorts, inclusion of individuals with normal weight, overweight, and obesity). The primary outcome's reliance on self-reported body weight and subsequent measurement biases further undermines the reliability of the findings. Additionally, the absence of registration in clinical trial registries, such as the EU Clinical Trials Register or clinicaltrials.gov, and the multiple testing of hypotheses which were not listed a priori in the research protocol published on the German Register of Clinical Trials impede the study's transparency and reproducibility.
Achievement of Objectives and Support for Conclusions:
The study's objectives were partially met; however, the interpretation of the effects of meal timing on weight loss is compromised by the weaknesses mentioned above. The evidence only partially supports some of the claims due to methodological flaws and unstructured data analysis.
Impact and Utility:
Despite its innovative approach, significant methodological and analytical shortcomings limit the study's utility. If these issues were addressed, the research could have meaningful implications for dietary interventions and metabolic research. The concept of timing of dietary occasions in sync with circadian rhythms holds promise but requires further rigorous investigation.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript the authors have generated a single-cell atlas of the miracidium, the first free-living stage of an important human parasite, Schistosoma mansoni. Miracidia develop from eggs produced in the mammalian (human) host and are released into freshwater, where they can infect the parasite's intermediate snail host to continue the life cycle. This study adds to the growing single-cell resources that have already been generated for other life-cycle stages and, thus, provides a useful resource for the field.
Strengths:
Beyond generating lists of genes that are differentially expressed in different cell types, the authors validated many of the cluster-defining genes using in situ hybridization chain reaction. In addition to providing the field with markers for many of the cell types in the parasite at this stage, the authors use these markers to count the total number of various cell types in the organism. Because the authors realized that their cell isolation protocols were biasing the cell types they were sequencing, they applied a second method to help them recover additional cell types.
Schistosomes have ZW sex chromosomes and the authors make the interesting observation that the stem cells at this stage are already expressing sex (i.e. W)-specific genes.
Weaknesses:
The sample sizes upon which the in situ hybridization results and cell counts are based are either not stated (in most cases) or are very small (n=3). This lack of clarity about biological replicates and sample sizes makes it difficult for the reader to assess the robustness of the results and the extremely small sample sizes (when provided) are a missed opportunity to explore the variability of the system, or lack thereof.
Although assigning transcripts to a given cell type is usually straightforward via in situ experiments, the authors fail to consider the potential difficulty of assigning the appropriate nuclei to cells with long cytoplasmic extensions, like neurons. In the absence of multiple markers and a better understanding of the nervous system, it seems likely that the authors have overestimated the number of neurons and misassigned other cell types based on their proximity to neural projections.
The conclusion that germline genes are expressed in the miracidia stem cells seems greatly overstated in the absence of any follow-up validation. The expression scales for genes like eled and boule are more than 3 orders of magnitude smaller than those used for any of the robustly expressed genes presented throughout the paper. These scales are undefined, so it isn't entirely clear what they represent, but neither of these genes is detected at levels remotely high (or statistically significant) enough to survive filters for cluster-defining genes. Given that germ cells often develop early in embryogenesis and arrest the cell cycle until later in development, and that these transcripts reveal no unspliced forms, it seems plausible that the authors are detecting some maternally supplied transcripts that have yet to be completely degraded.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Immunogenic cell death (ICD) can lead to the release of factors such as DAMPs which promote an adaptive immune response. In the context of cancer, there is clear evidence of anti-tumour benefits as a result of ICD, perhaps induced by chemotherapy.
Lilong et al used TCGA data to explore whether a previously published 34 gene 'ICD-related' signature could stratify bladder cancer patients by prognosis and ultimately predict patient survival. The gene signature contains many genes involved in inflammation and immunity (IFNg, IL6, TNF, IL17A, TLR4, CD8B, etc) and those related to ICD (such as CALR, HMGB1, HSP, NLRP3, etc). The authors divide patients into 'ICD-high' and '-low' based on the expression of this gene set and find that 'ICD-high' is associated with longer survival in TCGA bladder cancer data. The authors further argue that ICD-high group responds better to PD1 therapies. From this 34-gene signature, it appears that LASSO regularisation and Cox analysis identifies a four-gene 'risk' signature (CALR, IL1R1, IFNB1, IFNG) which is associated with shorter patient survival and lower immunotherapy response rates. This is the primary finding. Their methodology is very similar to a publication in 2021 in Frontiers in Immunology instead in the context of head and neck squamous cell carcinoma. This paper is not referenced.
In terms of the strengths of the work, it is certainly plausible that the author's four gene signature has an association with survival in bladder cancer, at least based on the two datasets studied. However, the relatedness of their findings to ICD is unconvincing, and glaring omissions from the manuscript in terms of methods limit confidence in the work. The authors show a potential association with bladder cancer patient survival and their four gene signatures, but substantial revisions are required for this to be appropriately evidenced.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, Ghazi et reported that inhibition of KRASG12C signaling increases autophagy in KRASG12C-expressing lung cancer cells. Moreover, the combination of DCC 3116, a selective ULK1/2 inhibitor, plus sotorasib displays cooperative/synergistic suppression of human KRASG12C-driven lung cancer cell proliferation in vitro and tumor growth in vivo. Additionally, in genetically engineered mouse models of KRASG12C-driven NSCLC, inhibition of either KRASG12C or ULK1/2 decreases tumor burden and increases mouse survival. Additionally, this study found that LKB1 deficiency diminishes the sensitivity of KRASG12C/LKB1Null-driven lung cancer to the combination treatment, perhaps through the emergence of mixed adeno/squamous cell carcinomas and mucinous adenocarcinomas.
Strengths:
Both human cancer cells and mouse models were employed in this study to illustrate that inhibiting ULK1/2 could enhance the responsiveness of KRASG12C lung cancer to sotorasib. This research holds translational importance.
Weaknesses:
Additional validation of certain data is necessary.
(1) mCherry-EGFP-LC3 reporter was used to assess autophagy flux in Figure 1A. Please explain how autophagy status (high, medium, and low) was defined. It's also suggested to show WB of LC3 processing in different treatments as in Figure 1A at 48 hours.
(2) For Figures 1J, K, and L, please provide immunohistochemistry (IHC) images demonstrating RAS downstream signaling blockade by sotorasib and autophagy blockade by DCC 3116 in tumors.
(3) Given that both DCC 3116 and ULK1K46N exhibit the ability to inhibit autophagy and synergize with sotorasib in inhibiting cell proliferation, in addition to demonstrating decreased levels of pATG13 via ELISA assay, please include Western blot analyses of LC3 or p62 to confirm the blockade of autophagy by DCC 3116 and ULK1K46N in Figure 1 & Figure 2.
(4) Since adenocarcinomas, adenosquamous carcinomas (ASC), and mucinous adenocarcinomas were detected in KL lung tumors, please conduct immunohistochemistry (IHC) to detect these tumors, including markers such as p63, SOX2, Katrine 5.
(5) Please provide the sample size (n) for each treatment group in the survival study (Figure 4E). It appears that all mice were sacrificed for tumor burden analysis in Figure 4F. However, there doesn't seem to be a significant difference among the treatment groups in Figure 4F, which contrasts with the survival analysis in Figure 4E. It is suggested to increase the sample size in each treatment group to reduce variation.
(6) In KP mice (Figure 5), it seems that a single treatment alone is sufficient to inhibit established KP lung tumor growth. Combination treatment does not further enhance anti-tumor efficacy. Therefore, this result doesn't support the conclusion generated from human cancer cell lines. Please discuss.
-
-
-
documentLinks do: [:link | thisSnippet database importDocumentFrom: link ].
smalltalk myPages := documentLinks collect: [:link | thisSnippet database importDocumentFrom: link ].En lugar dedo:es uncollect:para que la nueva colección quede asignada a la variablemyPages. Una vez esto funcione, el resto de las intrucciones permite exportar sólo las nuevas páginas, en lugar de todas.
-
-
-
documentLinks
~~documentLinks~~ ~> myPages
Con este nuevo iterador, es posible trabajar con la colección deseada:
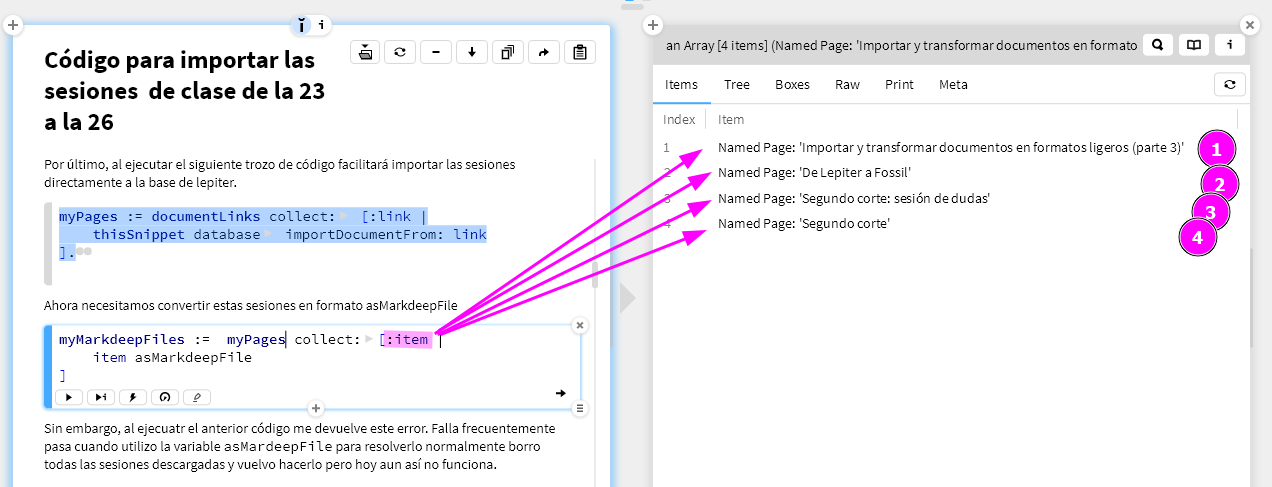
Una vez se trabaja con la colección correcta, el resto del código funciona y las páginas se pueden exportar.
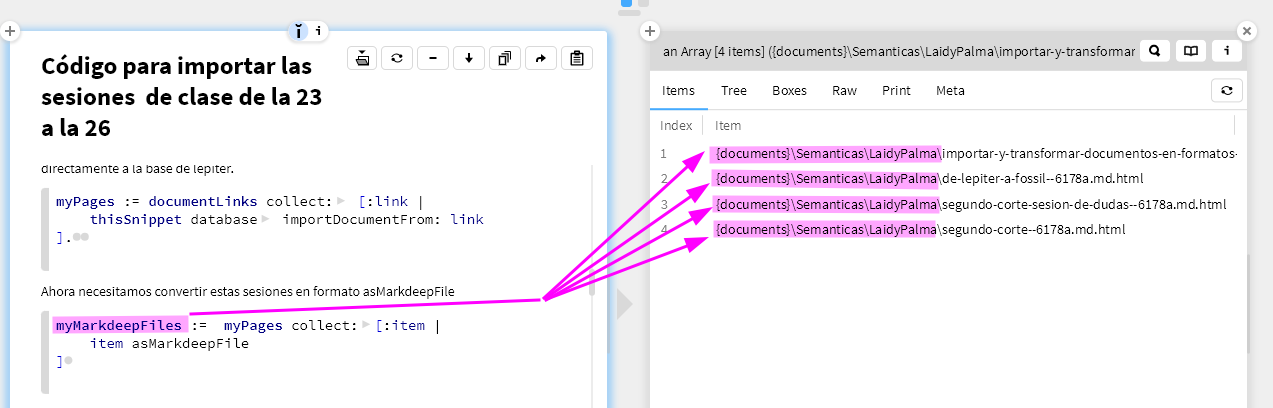
-
documentLinks collect: [:link | thisSnippet database importDocumentFrom: link ].
Este resultado hay que reusarlo, asignándolo a una variable.
smalltalk myPages := documentLinks collect: [:link | thisSnippet database importDocumentFrom: link ]. -
Falla frecuentemente pasa cuando utilizo la variable asMardeepFile para resolverlo normalmente borro todas las sesiones descargadas y vuelvo hacerlo pero hoy aun así no funciona.
Enviaste el mensaje
asMarkdeepFilea unString(cadena de texto en lugar de a una página de Lepiter. Tendrías que haber enviando el mensaje a unaNamed Page.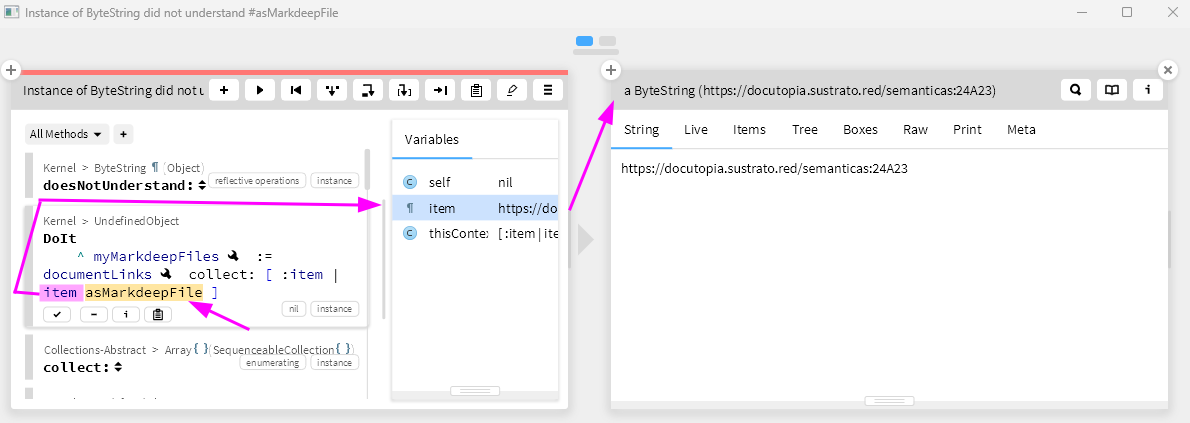
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Bates TA. et al. studied the biochemical characteristics of ESAT-6, a major virulence factor of Mycobacterium tuberculosis (Mtb), as part of the heterodimer with CFP10, a molecular chaperon of ESAT-6, as in homodimer and in homotetramer using recombinant ESAT-6 and CFP10 expressed in E. coli by applying several biochemical assays including Biolayer Interferometry (BLI) assay. The main findings show that ESAT-6 forms a tight interaction with CFP10 as a heterodimer at neutral pH, and ESAT-6 forms homodimer and even tetramer based larger molecular aggregates at acidic pH. Although the discussion of the potential problems associated with the contamination of ESAT-6 preparations with ASB-14 during the LPS removal step is interesting, but this research does not test the potential impact of residual ASB-14 contaminant on the biochemical behavior ESAT-6-CFP10 heterodimer and ESAT-6 homodimer or tetramer and their hemolytic activity in comparison with the ones without ASB-14. The main strength of this study is the generation of ESAT-6 specific nanobodies and demonstration of its anti-tuberculosis efficiency in THP-1 cell line infected with Mtb strains with reporter genes.
Strengths:
Generation and demonstration of the anti-ESAT-6 nanobodies against tuberculosis infection in cell line based Mtb infection model. Probably identifying potential anti-ESAT-6 nanobody interacting amino acid residues of ESAT-6 is critical in understanding their effects on ESAT-6 mediated membrane lytic activity.
Weaknesses:
Although the biochemistry studies provide quantitative data about the interactions of ESAT-6 with its molecular chaperon CFP10 and the interaction of ESAT-6 homodimer and tetramers, the novel information from these studies are minimal.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
In this work, the authors uncovered the effects of DNA dilution on E. coli, including a decrease in growth rate and a significant change in proteome composition. The authors demonstrated that the decline in growth rate is due to the reduction of active ribosomes and active RNA polymerases because of the limited DNA copy numbers. They further showed that the change in the DNA-to-volume ratio leads to concentration changes in almost 60% of proteins, and these changes mainly stem from the change in the mRNA levels.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors try to achieve a method of protection against pathogenic strains using saprophytic species. It is undeniable that the saprophytic species, despite not causing the disease, activates an immune response. However, based on these results, using the saprophytic species does not significantly impact the animal's infection by a virulent species.
Strengths:
Exposure to the saprophytic strain before the virulent strain reduces animal weight loss, reduces tissue kidney damage, and increases cellular response in mice.
Weaknesses:
Even after the challenge with the saprophyte strain, kidney colonization and the release of bacteria through urine continue. Moreover, the authors need to determine the impact on survival if the experiment ends on the 15th.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript by Alexander et al describes a careful and rigorous application of multiomics to mouse primordial germ cells (PGCs) and their surrounding gonadal cells during the period of sex differentiation.
Strengths:
In thoughtfully designed figures, the authors identify both known and new candidate gene regulatory networks in differentiating XX and XY PGCs and sex-specific interactions of PGCs with supporting cells. In XY germ cells, novel findings include the predicted set of TFs regulating Bnc2, which is known to promote mitotic arrest, as well as the TFs POU6F1/2 and FOXK2 and their predicted targets that function in mitosis and signal transduction. In XX germ cells, the authors deconstruct the regulation of the premeiotic replication regulator Stra8, which reveals TFs involved in meiosis, retinoic acid signaling, pluripotency, and epigenetics among predictions; this finding, along with evidence supporting the regulatory potential of retinoic acid receptors in meiotic gene expression is an important addition to the debate over the necessity of retinoic acid in XX meiotic initiation. In addition, a self-regulatory network of other TFs is hypothesized in XX differentiating PGCs, including TFAP2c, TCF5, ZFX, MGA, and NR6A1, which is predicted to turn on meiotic and Wnt signaling targets. Finally, analysis of PGC-support cell interactions during sex differentiation reveals more interactions in XX, via WNTs and BMPs, as well as some new signaling pathways that predominate in XY PGCs including ephrins, CADM1, Desert Hedgehog, and matrix metalloproteases. This dataset will be an excellent resource for the community, motivating functional studies and serving as a discovery platform.
Weaknesses:
My one major concern is that the conclusion that PGC sex differentiation (as read out by transcription) involves chromatin priming is overstated. The evidence presented in the figures includes a select handful of genes including Porcn, Rimbp1, Stra8, and Bnc2 for which chromatin accessibility precedes expression. Given that the authors performed all of their comparisons between XX versus XY datasets at each timepoint, have they missed an important comparison that would be a more direct test of chromatin priming: between timepoints for each sex? Furthermore, it remains possible that common mechanisms of differentiation to XX and XY could be missing from this analysis that focused on sex-specific differences.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Mistri et al explore the role of SLAM-SAP signaling in the developmental programming of innate-like gd T cell subsets. Using proteo-genomics, they determined that abrogation of SLAM-SAP signaling altered that programming, reducing some IL-17-producing subsets, including a novel Vγ4 γδT1 subset, and diverting gdTCR-expressing precursors to the ab fate. Altogether, this is a very thorough, thoughtfully interpreted study that adds significantly to our understanding of the contribution of the SLAM-SAP pathway to lineage specification. A particularly interesting element is the role of SLAM-SAP in preventing gd17 progenitors from switching fates and adopting the ab fate.
One thing to keep in mind in assessing the ultimate fate of the "ab wannabe cells" is that mechanisms exist to silence the gd TCR as cells differentiate to the DP stage and so their presence as diverted DP cells may not be evident by staining for gdTCR expression - and will only be evident transcriptomically.
Strengths:
This is an exceedingly well-designed and thorough study that significantly enriches our understanding of gd T cell development.
Weaknesses:
There are no major weaknesses identified by this reviewer.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Nguyen, Zhao et al. used bioinformatic analysis of mutational variants of SARS-CoV-2 Nucleocapsid (N) protein from the large genomic database of SARS-CoV-2 sequences to identify domains and regions of N where mutations are more highly represented, and computationally determined the effects of these mutations on the physicochemical properties of the protein. They found that the intrinsically disordered regions (IDRs) of N protein are more highly mutated than structured regions, and that these mutations can lead to higher variability in the physical properties of these domains. These computational predictions are compared to in vitro biophysical experiments to assess the effects of identified mutations on the thermodynamic stability, oligomeric state, particle formation, and liquid-liquid phase separation of a few exemplary mutants.
The paper is well written, easy to follow and the conclusions drawn are supported by the evidence presented. The analyses and conclusions are interesting and will be of value to virologists, cell biologists, and biophysicists studying SARS-CoV-2 function and assembly.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The origin of plasmatoid dendritic cells and their subclasses continues to be a debated field, akin to any immune cell field that is determined through the expression of surface markers (relative to clear subclass separation based on functional biology and experimentation). In this context, in this manuscript by Araujo et al, the authors attempt to demonstrate that a subtype of pDCs comes from lymphoid origin due to the presence of some B cell gene expression markers. They nomenclature these cells as B-pDCs. Strikingly, pDCs function via expression of IFNa where as B-pDCs do not express IFNa - thereby raising the question of what are their physiological or pathophysiological properties. B-pDCs also express AXL, a marker not seen in mouse pDCs but observed in human pDCs. Overall, using a combination of gene expression profiling of immune cells isolated from mice via RNA-seq and single-cell profiling the authors propose that B-pDCs are a novel subtype of pDCs in mice that were not previously identified and characterized.
Weaknesses:
My two points of discussion about this manuscript are as follows.
(1) How new are these observations that pDCs could also originate from common lymphoid progenitors. This fact has been previously outlined by many laboratories including Shigematsu et al, Immunity 2004. These studies in the manuscript can be considered new based on the single-cell profiling presented, only if the further characterization of the isolated B-pDCs is performed at the functional biology level. Overlapping gene expression profiles are often seen in developing immune cell types- especially when only evaluated at the RNA expression level- and can lead to cell type complexity (and identification of new cell types) that are not biologically and functionally relevant.
(2) The authors hardly perform any experiments to interrogate the function of these B-pDCs. The discussion on this topic can be enhanced. Ideally, some biological experiments would confirm that B-pDCs are important.
Tags
Annotators
URL
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, the authors provide structural analysis of the NHL domain for C. elegans NHL-2 and provide functional analysis of the NHL RNA binding domain. Their data support a model in which NHL-2 binding to mRNA targets through U rich motifs to promote miRISC regulation of translation and mRNA stability.
Strengths:
The authors present convincing data to describe the structure of the NHL-2 NHL domain along with functional analysis that supports an important role for two amino acids that are required for RNA binding activity. The function of these two amino acids were further studied through phenotypic assays to analyze their contribution to miRNA mediated regulation through the let-7 pathway. These data support an important role for RNA binding activity of NHL-2 in the regulation of miRNA dependent pathways. Genetic interactions support a role for the eIF4E binding protein IFET-1 in the miRISC activity.
Weaknesses:
The use of phenotypic assays to monitor let-7 pathway activity could be better explained so that the reader can more easily follow the significance of changes in alae formation or col-19::gfp expression.
The challenges of comparing expression levels using extrachromosomal arrays should be acknowledged.
The figure legends need to be revised to more clearly and accurately explain what is shown in the figures.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript by Cannarsa et. al., the authors describe the engineering of a light-entrainable synthetic biological oscillator in bacteria. It is based on an upgraded version of one of the first synthetic circuits to be constructed, the repressilator. The authors sought to make this oscillator entrainable by an external forcing signal, analogous to the way natural biological oscillators (like the circadian clock) are synchronized. They reasoned that an optogenetic system would provide a convenient and flexible means of manipulation. To this end, the authors exploited the CcaS-CcaA light-switchable system, which allows activation and deactivation of transcription by green and red light, respectively. They used this system to make the expression of one of the repressilator's transcription factors (lacI) light-controlled, from a construct separated from the main repressilator plasmid. This way, under red light the oscillator runs freely, but exposure to green light causes overexpression of the lacI, pushing the system into a specific state. Consequently, returning to red light will restore the oscillations from the same phase in all cells, effectively synchronizing the cell population.
After demonstrating the functionality of the basic concept, the authors combined modeling and experiments to show how periodic exposure to green light enables efficient entrainment, and how the frequency of the forcing signal affects the oscillatory behavior (detuning).
This work provides an important demonstration of engineering tunability into a foundational genetic circuit, expands the synthetic biology toolbox, and provides a platform to address critical questions about synchronization in biological oscillators. Due to the flexibility of the experimental system, it is also expected to provide a fertile ground for future testing of theoretical predictions regarding non-linear oscillators.
Strengths:
* The study provides a simple and elegant mechanism for the entertainment of a synthetic oscillator. The design relies on optogenetic proteins, which enable efficient experimentation compared to alternative approaches (like using chemical inducers). This way, a static culture (without microfluidics or change of growth media) can be easily exposed to flexible temporal sequences of the zeitgeber, and continuously measured through time.
* The study makes use of both plate-reader-based population-level readout and mother-machine single-cell measurements. Synchronization through entrainment is a single cell level phenomenon, but with a clear population-level manifestation. Thus, this experimental approach combination provides a strong validation to their system. At the same time, differences between the readout from the two systems have emerged, and provided a further opportunity for model refinement and testing.
* The authors correctly identified the main optimization goal, namely the effective leakiness of their construct even under red light. Then, they successfully overcame this issue using synthetic biology approaches.
* The work is supported by a simplified model of the repressilator, which provides a convenient analytical and numerical means to draw testable predictions. The model predictions are well aligned with the experimental evidence.
Weaknesses:
* Even after optimizing the expression level of the light-sensitive gene, the system is very sensitive, i.e., a very short exposure is sufficient to elicit the strongest entertainment. This limited dynamic range might hamper some model testing and future usage.
* As a result of the previous point, the system is entrained by transiently "breaking" the oscillator: each pulse of green light represents a Hopf bifurcation into a single attractor. it means that the system cannot oscillate in constant green light. In comparison, this is generally not the case for natural zeitgebers like light and temperature for the circadian rhythms. Extreme values might prevent oscillations (not necessarily due to breaking the core oscillator), but usually, free running is possible in a wide range of constant conditions. In some cases, the free-running period length will vary as a function of the constant value.
While the approach presented in this manuscript is valid, a comprehensive analysis of more subtle modes of repressilator entrainment could also be of value.
* The entire work makes use of a single intensity and single duration of the green pulse to force entrainment. While the model has clear predictions for how those modalities should affect entrainment, none of the experiments attempted to validate those predictions.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Studying Apteronotus leptorhynchus (the weakly electric brown ghost knifefish), the authors provide evidence that 'chirps' (brief modulations in the frequency and amplitude of the ongoing electric signal) function in active sensing (specifically homeoactive sensing) rather than communication. Chirping is a behavior that has been well studied, including numerous studies on the sensory coding of chirps and the neural mechanisms for chirp generation. Chirps are largely thought to function in communication behavior, so this alternative function is a very exciting possibility that could have a great impact on the field. The authors do provide convincing evidence that chirps may function in homeoactive sensing. However, their evidence arguing against a role for chirps in communication is not as strong, and fails to sufficiently consider the evidence from a large body of existing research. Ultimately, the manuscript presents very interesting data that is sure to stimulate discussion and follow-up studies, but it suffers from dismissing evidence in support of, or consistent with, a communicative function for chirps. The authors do acknowledge that chirps could function as both a communication and homeactive sensing signal, but it seems clear they wish to argue against the former and for the latter, and the evidence is not yet there to support this.
In the introduction, the authors state, "Since both chirps and positional parameters (such as size, orientation or motion) can only be detected as perturbations of the beat, and via the same electroreceptors, the inputs relaying both types of information are inevitably interfering." I disagree with this statement, which seems to be a key assumption. Both of these features certainly modulate the activity of electroreceptors, but that does not mean those modulations are ambiguous as to their source. You do not know whether the two types of modulations can be unambiguously decoded from electroreceptor afferent population activity.
My biggest issue with this manuscript is that it is much too strong in dismissing evidence that chirping correlates with context. In your behavioral observations, you found sex differences in chirping as well as differences between freely interacting and physically separated fish. Chirps tended to occur in close proximity to another fish. Your model of chirp variability found that environmental experience, social experience, and beat frequency (DF) are the most important factors explaining chirp variability. Are these not all considered behavioral or social context? Beat frequency (DF) in particular is heavily downplayed as being a part of "context" but it is a crucial part of the context, as it provides information about the identity of the fish you're interacting with. The authors show quite convincingly that the types of chirps produced do not vary with these contexts, but chirp rates do.
Further, in your playback experiments, fish responded differently to small vs. large DFs, males chirped more than females, type 2 chirps became more frequent throughout a playback, and rises tended to occur at the end of a playback. These are all examples of context-dependent behavior.
In the results, the authors state, "Overall, the majority of chirps were produced by male subjects, in comparable amounts regardless of environmental experience (resident, intruder or equal; Figure S1A,C), social status (dominant or subordinate; Figure S1B) or social experience (novel or experienced; Figure S1D)." This is not what is shown in Figure S1. S1A shows clear differences between resident vs. intruder males, S1B shows clear differences between dominant vs. subordinate males, and S1D shows clear differences between naïve and experienced males. The analysis shown in Figure 2 would seem to support this. Indeed, the authors state, "Overall, this analysis indicated that environmental and social experience, together with beat frequency (DF) are the most important factors explaining chirp variability."
The choice of chirp type varied widely between individuals but was relatively consistent within individuals across trials of the same experiment. The authors interpret this to mean that chirping does not vary with internal state, but is it not likely that the internal states of individuals are stable under stable conditions, and that individuals may differ in these internal states across the same conditions? Stable differences in communication signals between individuals are frequently interpreted as reflecting differences between those individuals in certain characteristics, which are being communicated by these signals.
I am not convinced of the conclusion drawn by the analysis of chirp transitions. The transition matrices show plenty of 1-2 and 2-1 transitions occurring. Further, the cross-correlation analysis only shows that chirp timing between individuals is not phase-locked at these small timescales. It is entirely possible that chirp rates are correlated between interacting individuals, even if their precise timing is not. Further, it is not clear to me how "transitions" were defined. The methods do not make this clear, and it is not clear to me how you can have zero chirp transitions between two individuals when those two individuals are both generating chirps throughout an interaction.
In the results, "Although all chirp types were used during aggressive interactions, these seemed to be rather less frequent in the immediate surround of the chirps (Figure 6A)." A lack of precise temporal correlation on short timescales does not mean there is no association between the two behaviors. An increased rate of chirping during aggression is still a correlation between the two behaviors, even if chirps and specific aggressive behaviors are not tightly time-locked.
In summary, it is simply too strong to say that chirping does not correlate with context, or to claim that there is convincing evidence arguing against a communication function of chirps. Importantly, however, this does not detract from your exciting and well-supported hypothesis that chirping functions in homeoactive sensing. A given EOD behavior could serve both communication and homeoactive sensing. I actually suspect this is quite common in electric fish (both gymnotiforms and mormyrids), and perhaps in other actively sensing species such as echolocating animals. The two are not mutually exclusive.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary: In Deconstructing Complexity: A Computational Topology Approach to Trajectory Inference in the Human Thymus with tviblindi, Stuchly et al. propose a new trajectory inference algorithm called tviblindi and a visualization algorithm called vaevictis for single-cell data. The paper utilizes novel and exciting ideas from computational topology coupled with random walk simulations to align single cells onto a continuum. The authors validate the utility of their approach largely using simulated data and establish known protein expression dynamics along CD4/CD8 T cell development in thymus using mass cytometry data. The authors also apply their method to track Treg development in single-cell RNA-sequencing data of human thymus.
The technical crux of the method is as follows: The authors provide an interactive tool to align single cells along a continuum axis. The method uses expected hitting time (given a user input start cell) to obtain a pseudotime alignment of cells. The pseudotime gives an orientation/direction for each cell, which is then used to simulate random walks. The random walks are then arranged/clustered based on the sparse region in the data they navigate using persistent homology.
Strengths:<br /> The notion of using persistent homology to group random walks to identify trajectories in the data is novel.<br /> The strength of the method lies in the implementation details that make computationally demanding ideas such as persistent homology more tractable for large scale single-cell data. This enables the authors to make the method more user friendly and interactive allowing real-time user query with the data.
Weaknesses:<br /> The interactive nature of the tool is also a weakness, by allowing for user bias leading to possible overfitting for a specific data.
The main weakness of the method is lack of benchmarking the method on real data and comparison to other methods. Trajectory inference is a very crowded field with many highly successful and widely used algorithms, the two most relevant ones (closest to this manuscript) are not only not benchmarked against, but also not sited. Including those that specifically use persistent homology to discover trajectories (Rizvi et.al. published Nat Biotech 2017). Including those that specifically implement the idea of simulating random walks to identify stable states in single-cell data (e.g. CellRank published in Lange et.al Nat Meth 2022), as well as many trajectory algorithms that take alternative approaches. The paper has much less benchmarking, demonstration on real data and comparison to the very many other previous trajectory algorithms published before it. Generally speaking, in a crowded field of previously published trajectory methods, I do not think this one approach will compete well against prior work (especially due to its inability to handle the noise typical in real world data (as was even demonstrated in the little bit of application to real world data provided).
Beyond general lack of benchmarking there are two issues that give me particular concern. As previously mentioned, the algorithm is highly susceptible to user bias and overfitting. The paper gives the example (Figure 4) of a trajectory which mistakenly shows that cells may pass from an apoptotic phase to a different developmental stage. To circumvent this mistake, the authors propose the interactive version of tviblindi that allows users to zoom in (increase resolution) and identify that there are in fact two trajectories in one. In this case, the authors show how the author can fix a mistake when the answer is known. However, the point of trajectory inference is to discover the unknown. With so much interactive options for the user to guide the result, the method is more user/bias driven than data-driven. So a rigorous and quantitative discussion of robustness of the method, as well as how to ensure data-driven inference and avoid over-fitting would be useful.
Second, the paper discusses the benefit of tviblindi operating in the original high dimensions of the data. This is perhaps adequate for mass cytometry data where there is less of an issue of dropouts and the proteins may be chosen to be large independent. But in the context of single-cell RNA-sequencing data, the massive undersampling of mRNA, as well as high degree of noise (e.g. ambient RNA), introduces very large degree of noise so that modeling data in the original high dimensions leads to methods being fit to the noise. Therefore ALL other methods for trajectory inference work in a lower dimension, for very good reason, otherwise one is learning noise rather than signal. It would be great to have a discussion on the feasibility of the method as is for such noisy data and provide users with guidance. We note that the example scRNA-seq data included in the paper is denoised using imputation, which will likely result in the trajectory inference being oversmoothed as well.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors conducted a comparative analysis of four networks, varying in the presence of excitatory assemblies and the architecture of inhibitory cell assembly connectivity. They found that co-tuned E-I assemblies provide network stability and a continuous representation of input patterns (on locally constrained manifolds), contrasting with networks with global inhibition that result in attractor networks.
Strengths:
The findings presented in this paper are very interesting and cutting-edge. The manuscript effectively conveys the message and presents a creative way to represent high-dimensional inputs and network responses. Particularly, the result regarding the projection of input patterns onto local manifolds and continuous representation of input/memory is very Intriguing and novel. Both computational and experimental neuroscientists would find value in reading the paper.
Weaknesses:
Intuitively, classification (decodability) in discrete attractor networks is much better than in networks that have continuous representations. This could also be shown in Figure 5B, along with the performance of the random and tuned E-I networks. The latter networks have the advantage of providing network stability compared to the Scaled I network, but at the cost of reduced network salience and, therefore, reduced input decodability. The authors may consider designing a decoder to quantify and compare the classification performance of all four networks.
Networks featuring E/I assemblies could potentially represent multistable attractors by exploring the parameter space for their reciprocal connectivity and connectivity with the rest of the network. However, for co-tuned E-I networks, the scope for achieving multistability is relatively constrained compared to networks employing global or lateral inhibition between assemblies. It would be good if the authors mentioned this in the discussion. Also, the fact that reciprocal inhibition increases network stability has been shown before and should be cited in the statements addressing network stability (e.g., some of the citations in the manuscript, including Rost et al. 2018, Lagzi & Fairhall 2022, and Vogels et al. 2011 have shown this).
Providing raster plots of the pDp network for familiar and novel inputs would help with understanding the claims regarding continuous versus discrete representation of inputs, allowing readers to visualize the activity patterns of the four different networks. (similar to Figure 1B).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript reports interesting findings about the navigational behavior of mice. The authors have dissected this behavior in various components using a sophisticated behavioral maze and statistical analysis of the data.
Strengths:
The results are solid and they support the main conclusions, which will be of considerable value to many scientists.
Weaknesses:
Figure 1: In some trials the mice seem to be doing thigmotaxis, walking along the perimeter of the maze. This is perhaps due to the fear of the open arena. But, these paths along the perimeter would significantly influence all metrics of navigation, e.g. the distance or time to reward. Perhaps analysis can be done that treats such behavior separately and the factors it out from the paths that are away from the perimeter.
Figure 1c: the color axis seems unusual. Red colors indicate less frequently visited regions (less than 25%) and white corresponds to more frequently visited places (>25%)? Why use such a binary measure instead of a graded map as commonly done?
Some figures use linear scale and others use logarithmic scale. Is there a scientific justification? For example, average latency is on a log scale and average speed is on a linear scale, but both quantify the same behavior. The y-axis in panel 1-I is much wider than the data. Is there a reason for this? Or can the authors zoom into the y-axis so that the reader can discern any pattern?<br /> <br /> 1F shows no significant reduction in distance to reward. Does that mean there is no improvement with experience and all the improvement in the latency is due to increasing running speed with experience?
Figure 3: The distance traveled was reduced by nearly 10-fold and speed increased by by about 3fold. So, the time to reach the reward should decrease by only 3 fold (t=d/v) but that too reduced by 10fold. How does one reconcile the 3fold difference between the expected and observed values?
Figure 4: The reader is confused about the use of a binary color scheme here for the checking behavior: gray for a large amount of checking, and pink for small. But, there is a large ellipse that is gray and there are smaller circles that are also gray, but these two gray areas mean very different things as far as the reader can tell. Is that so? Why not show the entire graded colormap of checking probability instead of such a seemingly arbitrary binary depiction?
Figure 4C: What would explain the large amount of checking behavior at the perimeter? Does that occur predominantly during thigmotaxis?
Was there a correlation between the amount of time spent by the animals in a part of the maze and the amount of reward checking? Previous studies have shown that the two behaviors are often positively correlated, e.g. reference 20 in the manuscript. How does this fit with the path integration hypothesis?
"Scratches and odor trails were eliminated by washing and rotating the maze floor between trials." Can one eliminate scratches by just washing the maze floor? Rotation of the maze floor between trials can make these cues unreliable or variable but will not eliminate them. Ditto for odor cues.
"Possible odor gradient cues were eliminated by experiments where such gradients were prevented with vacuum fans (Fig. S6E)" What tests were done to ensure that these were *eliminated* versus just diminished?
"Probe trials of fully trained mice resulted in trajectories and initial hole checking identical to that of regular trials thereby demonstrating that local odor cues are not essential for spatial learning." As far as the reader can tell, probe trials only eliminated the food odor cues but did not eliminate all other odors. If so, this conclusion can be modified accordingly. <br /> The interpretation of direction selectivity is a bit tricky. At different places in this manuscript, this is interpreted as a path integration signal that encodes goal location, including the Consync cells. However, studies show that (e.g. Acharya et al. 2016) direction selectivity in virtual reality is comparable to that during natural mazes, despite large differences in vestibular cues and spatial selectivity. How would one reconcile these observations with path integration interpretation?
The manuscript would be improved if the speculations about place cells, grid cells, BTSP, etc. were pared down. I could easily imagine the outcome of these speculations to go the other way and some claims are not supported by data. "We note that the cited experiments were done with virtual movement constrained to 1D and in the presence of landmarks. It remains to be shown whether similar results are obtained in our unconstrained 2D maze and with only self-motion cues available." There are many studies that have measured the evolution of place cells in non-virtual mazes, look up papers from the 1990s. Reference 43 reports such results in a 2D virtual maze.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The manuscript investigates the function of basal forebrain cholinergic axons in mouse primary visual cortex (V1) during locomotion using two-photon calcium imaging in head-fixed mice. Cholinergic modulation has previously been proposed to mediate the effects of locomotion on V1 responses. The manuscript concludes that the activity of basal forebrain cholinergic axons in visual cortex provides a signal which is more correlated with binary locomotion state than locomotion velocity of the animal and finds no evidence for modulation of cholinergic axons by locomotion velocity. Cholinergic axons did not seem to respond to grating stimuli or visuomotor prediction error. Optogenetic stimulation of these axons increased the amplitude of responses to visual stimuli and decreased the response latency of layer 5 excitatory neurons, but not layer 2/3 neurons. Moreover, optogenetic or chemogenetic stimulation of cholinergic inputs reduced pairwise correlation of neuronal responses. These results provide insight into the role of cholinergic modulation to visual cortex and demonstrate that it affects different layers of visual cortex in a distinct manner. The experiments are well executed and the data appear to be of high quality. However, further analyses may be required to fully support some of the study's conclusions. Specifically, the analyses of the effects of locomotion and stimulation of cholinergic inputs present grand averages of responses across all neurons, and therefore may mask heterogeneity across layer 2/3 and layer 5 neurons.
-
-
-
Reviewer #2 (Public Review):
Summary:
The current work describes a set of behavioral tasks to explore individual differences in the preferred perceptual and motor rhythms. Results show a consistent individual preference for a given perceptual and motor frequency across tasks and, while these were correlated, the latter is slower than the former one. Additionally, the adaptation accuracy to rate changes is proportional to the amount of rate variation and, crucially, the amount of adaptation decreases with age.
Strengths:
Experiments are carefully designed to measure individual preferred motor and perceptual tempo. Furthermore, the experimental design is validated by testing the consistency across tasks and test-retest, what makes the introduced paradigm a useful tool for future research.<br /> The obtained data is rigorously analyzed using a diverse set of tools, each adapted to the specificities across the different research questions and tasks.<br /> This study identifies several relevant behavioral features: (i) each individual shows a preferred and reliable motor and perceptual tempo and, while both are related, the motor is consistently slower than the pure perceptual one; (ii) the presence of hysteresis in the adaptation to rate variations; and (iii) the decrement of this adaptation with age. All these observations are valuable for the auditory-motor integration field of research, and they could potentially inform existing biophysical models to increase their descriptive power.
Weaknesses:
To get a better understanding of the mechanisms underlying the behavioral observations, it would have been useful to compare the observed pattern of results with simulations done with existing biophysical models. However, this point is addressed if the current study is read along with this other publication of the same research group: Kaya, E., & Henry, M. J. (2024, February 5). Modeling rhythm perception and temporal adaptation: top-down influences on a gradually decaying oscillator. https://doi.org/10.31234/osf.io/q9uvr
Tags
Annotators
URL
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This manuscript asks how different forms of selection affect the patterns of genetic diversity in microbial populations. One popular metric used to infer signatures of selection is dN/dS, the ratio of nonsynonymous to synonymous distances between two genomes. Previous observations across many bacterial species have found dN/dS decreases with dS, which is a proxy for the divergence time. The most common interpretation of this pattern was proposed by Rocha et al. (2006), who suggested the excess in nonsynonymous mutations on short divergence times represent transient deleterious mutations that have not yet been purged by selection.
In this study, the authors propose an alternative model based on the population structure of human gut bacteria, in which dN is dominated by selective sweeps of SNPs that revert previous mutations within local populations. The authors argue that contrary to standard population genetics models, which are based on the population dynamics of large eukaryotes, the large populations in the human gut mean that reversions may be quite common and may have a large impact on evolutionary dynamics. They show that such a model can fit the decrease of dN/dS in time at least as well as the purifying selection model.
Strengths
The main strength of the manuscript is to show that adaptive sweeps in gut microbial populations can lead to small dN/dS. While previous work has shown that using dN/dS to infer the strength of selection within a population is problematic (see Kryazhimskiy and Plotkin, 2008, cited in the paper) the particular mechanism proposed by the authors is new to my knowledge. In addition, despite the known caveats, dN/dS values are still routinely reported in studies of microbial evolution, and so their interpretation should be of considerable interest to the community.
The authors provide compelling justification for the importance of adaptive reversions and make a good case that these need to be carefully considered by future studies of microbial evolution. The authors show that their model can fit the data as well as the standard model based on purifying selection and the parameters they infer appear to be plausible given known data. More generally, I found the discussion on the implications of traditional population genetics models in the context of human gut bacteria to be a valuable contribution of the paper.
Weaknesses
The authors argue that the purifying selection model would predict a gradual loss in fitness via Muller's ratchet. This is true if recombination is ignored, but this assumption is inconsistent with the data from Garud, et al. (2019) cited in the manuscript, who showed a significant linkage decrease in the bacteria also used in this study.
I also found that the data analysis part of the paper added little new to what was previously known. Most of the data comes directly from the Garud et al. study and the analysis is very similar as well. Even if other appropriate data may not currently be available, I feel that more could be done to test specific predictions of the model with more careful analysis.
Finally, I found the description of the underlying assumptions of the model and the theoretical results difficult to understand. I could not, for example, relate the fitting parameters nloci and Tadapt to the simulations after reading the main text and the supplement. In addition, it was not clear to me if simulations involved actual hosts or how the changes in selection coefficients for different sites was implemented. Note that these are not simply issues of exposition since the specific implementation of the model could conceivably lead to different results. For example, if the environmental change is due to the colonization of a different host, it would presumably affect the selection coefficients at many sites at once and lead to clonal interference. Related to this point, it was also not clear that the weak mutation strong selection assumption is consistent with the microscopic parameters of the model. The authors also mention that "superspreading" may somehow make a difference to the probability of maintaining the least loaded class in the purifying selection model, but what they mean by this was not adequately explained.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors investigate replay (defined as sequential reactivation) and clustered reactivation during retrieval of an abstract cognitive map. Replay and clustered reactivation were analysed based on MEG recordings combined with a decoding approach. While the authors state to find evidence for both, replay and clustered reactivation during retrieval, replay was exclusively present in low performers. Further, the authors show that reactivation strength declined with an increasing graph distance.
Strengths:
The paper raises interesting research questions, i.e., replay vs. clustered reactivation and how that supports retrieval of cognitive maps. The paper is well written, well structured and easy to follow. The methodological approach is convincing and definitely suited to address the proposed research questions.
The paper is a great combination between replicating previous findings (Wimmer et al. 2020) with a new experimental approach but at the same time presenting novel evidence (reactivation strength declines as a function of graph distance).
What I also want to positively highlight is their general transparency. For example, they pre-registered this study but with a focus on a different part of the data and outlined this explicitly in the paper.
The paper has very interesting findings. However, there are some shortcomings, especially in the experimental design. These are shortly outlined below but are also openly and in detail discussed by the authors.
Weaknesses:
The individual findings are interesting. However, due to some shortcomings in the experimental design they cannot be profoundly related to each other. For example, the authors show that replay is present in low but not in high performers with the assumption that high performers tend to simultaneously reactivate items. But then, the authors do not investigate clustered reactivation (= simultaneous reactivation) as a function of performance due to a low number of retrieval trials and ceiling performance in most participants.<br /> As a consequence of the experimental design, some analyses are underpowered (very low number of trials, n = ~10, and for some analyses, very low number of participants, n = 14).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The authors characterized the recombinase-based cumulative fate maps for vesicular glutamate transporters (Vglut1, Vglut2 and Vglut3) expression and compared those maps to their real-time expression profiles in central NA neurons by RNA in situ hybridization in adult mice. Authors have revealed a new and intriguing expression pattern for Vglut2, along with an entirely uncharted co-expression domain for Vglut3 within central noradrenergic neurons. Interestingly, and in contrast to previous studies, the authors demonstrated that glutamatergic signaling in central noradrenergic neurons does not exert any influence on breathing and metabolic control either under normoxic/normocapnic conditions or after chemoreflex stimulation. Also, they showed for the first-time the Vglut3-expressing NA population in C2/A2 nuclei. In addition, they were also able to demonstrate Vglut2 expression in anterior NA populations, such as LC neurons, by using more refined techniques, unlike previous studies.
A major strength of the study is the use of a set of techniques to investigate the participation of NA-based glutamatergic signaling in breathing and metabolic control. The authors provided a full characterization of the recombinase-based cumulative fate maps for Vglut transporters. They performed real-time mRNA expression of Vglut transporters in central NA neurons of adult mice. Further, they evaluated the effect of knocking down Vglut2 expression in NA neurons using a DBH-Cre; Vglut2cKO mice on breathing and control in unanesthetized mice. Finally, they injected the AAV virus containing Cre-dependent Td tomato into LC of v-Glut2 Cre mice to verify the VGlut2 expression in LC-NA neurons. A very positive aspect of the article is that the authors combined ventilation with metabolic measurements. This integration holds particular significance, especially when delving into the exploration of respiratory chemosensitivity. Furthermore, the sample size of the experiments is excellent.<br /> Despite the clear strengths of the paper, some weaknesses exist. It is not clear in the manuscript if the experiments were performed in males and females and if the data were combined. I believe that the study would have benefited from a more comprehensive analysis exploring the sex specific differences. The reason I think this is particularly relevant is the developmental disorders mentioned by the authors, such as SIDS and Rett syndrome, which could potentially arise from disruptions in central noradrenergic (NA) function, exhibit varying degrees of sex predominance. Moreover, some of the noradrenergic cell groups are sexually dimorphic. For instance, female Wistar rats exhibit a larger LC size and more LC-NA neurons than male subjects (Pinos et al., 2001; Garcia-Falgueras et al., 2005). More recently, a detailed transcriptional profiling investigation has unveiled the identities of over 3,000 genes in the LC. This revelation has highlighted significant sexual dimorphisms, with more than 100 genes exhibiting differential expression within LC-NA neurons at the transcript level. Furthermore, this investigation has convincingly showcased that these distinct gene expression patterns have the capacity to elicit disparate behavioral responses between sexes (Mulvey et al., 2018). Therefore, the authors should compare the fate maps, Vglut transporters in males and females, at least considering LC-NA neurons. Even in the absence of identified sex differences, this information retains significant importance.<br /> An important point well raised by the authors is that although suggestive, these experiments do not definitively rule out that NA-Vglut2 based glutamatergic signaling has a role in breathing control. Subsequent experiments will be necessary to validate this hypothesis.
An improvement could be made in terms of measuring body temperature. Opting for implanted sensors over rectal probes would circumvent the need to open the chamber, thereby preventing alterations in gas composition during respiratory measurements. Further, what happens to body temperature phenotype in these animals under different gas exposures? These data should be included in the Tables.
Is it plausible that another neurotransmitter within NA neurons might be released in higher amounts in DBH-Cre; Vglut2 cKO mice to compensate for the deficiency in glutamate and prevent changes in ventilation?
Continuing along the same line of inquiry is there a possibility that Vglut2 cKO from NA neurons not only eliminates glutamate release but also reduces NA release? A similar mechanism was previously found in VGLUT2 cKO from DA neurons in previous studies (Alsio et al., 2011; Fortin et al., 2012; Hnasko et al., 2010). Additionally, does glutamate play a role in the vesicular loading of NA? Therefore, could the lack of effect on breathing be explained by the lack of noradrenaline and not glutamate?
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This is an interesting and well-performed study that develops a new modeling approach (MoA-HMM) to simultaneously characterize reinforcement learning parameters of different RL agents, as well as latent behavioral states that differ in the relative contributions of those agents to the animal's choices. They performed this study in rats trained to perform the two-step task. While the major advance of the paper is developing and rigorously validating this novel technical approach, there are also a number of interesting conceptual advances. For instance, humans performing the two-step task are thought to exhibit a trade-off between model-free and model-based strategies. However, the MoA-HMM did not reveal such a trade-off in the rats, but instead suggested a trade-off between model-based exploratory vs. exploitative strategies. Additionally, the firing rates of neurons in the orbitofrontal cortex (OFC) reflected latent behavioral states estimated from the HMM, suggesting that (1) characterizing dynamic behavioral strategies might help elucidate neural dynamics supporting behavior, and (2) OFC might reflect the contributions of one or a subset of RL agents that are preferentially active or engaged in particular states identified by the HMM.
Strengths:
The claims were generally well-supported by the data. The model was validated rigorously and was used to generate and test novel predictions about behavior and neural activity in OFC. The approach is likely to be generally useful for characterizing dynamic behavioral strategies.
Weaknesses:
There were a lot of typos and some figures were mis-referenced in the text and figure legends.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This research shows compelling and detailed evidence showing that aging influences intrinsic membrane properties of peripheral sympathetic motor neurons, which become hyperexcitable. The authors found that sympathetic motor neurons from old mice exhibit increased firing rates (spontaneous and evoked), more depolarized membrane resting potential, and increased rheobase. Furthermore, the study investigates cellular mechanisms underlying age-associated hyperexcitability and shows solid evidence supporting that a decreased activity of KCNQ channels during aging is a major contributor to the increased excitability of sympathetic old neurons. All conclusions of this paper are well supported by the data.
Strengths:
Detailed and rigorous analysis of electrical responses of peripheral sympathetic motor neurons using electrophysiology (perforated patch and whole-cell recordings). The study identifies a decrease in KCNQ current as a cellular mechanism behind age-induced hyperexcitability in sympathetic motor neurons.
Weaknesses:
None, the revised version of the manuscript has addressed all my concerns.
-
-
www.researchsquare.com www.researchsquare.com
-
Reviewer #2 (Public Review):
Summary:
The paper presents PPI-hotspot a method to predict PPI-hotspots. Overall, it could be useful but serious concerns about the validation and benchmarking of the methodology make it difficult to predict its reliability.
Strengths:
Develops an extended benchmark of hot-spots.
Weaknesses:
(1) Novelty seems to be just in the extended training set. Features and approaches have been used before.
(2) As far as I can tell the training and testing sets are the same. If I am correct, it is a fatal flaw.
(3) Comparisons should state that: SPOTONE is a sequence (only) based ML method that uses similar features but is trained on a smaller dataset. FTmap I think predicts binding sites, I don't understand how it can be compared with hot spots. Suggesting superiority by comparing with these methods is an overreach.
(4) Training in the same dataset as SPOTONE, and then comparing results in targets without structure could be valuable.
(5) The paper presents as validation of the prediction and experimental validation of hotspots in human eEF2. Several predictions were made but only one was confirmed, what was the overall success rate of this exercise?
Tags
Annotators
URL
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This article has characterized the mouse Schlemm's canal expression profile using a comprehensive approach based on sorted SEC, LEC, and BEC total RNA-Seq, scRNA-Seq, and snRNA-Seq to enrich the selection of SECs. The study has successfully profiled genome-wide gene expression using sorted SECs, demonstrating that SECs have a closer similarity to LECs than BECs. The combined scRNA- and snRNA-Seq data with deep coverage of gene expression led to the successful identification of many novel biomarkers for inner wall SECs, outer wall SECs, collector channel ECs, and pericytes. In addition, the study also identified two novel states of inner wall SECs separated by new markers. The study provides significant novel information about the biology and expression profile of SECs in the inner and outer walls. It is of great significance to have this novel, convincing, and comprehensive study led by leading researchers published in this journal.
Strengths:
This is a comprehensive study using various data to support the expression characterization of mouse SECs. First, the study profiled genome-wide expression using sorted SECs, LECs, and BECs from the same tissue/organ to identify the similarities and differences among the three types of cells. Second, snRNA-Seq was applied to enrich the number of SECs from mouse ocular tissues significantly. Increased sampling of SECs and other cells led to more comprehensive coverage and characterization of cells, including pericytes. Third, the combined scRNA- and snRNA-Seq data analyses increase the power to further characterize the subtle differences within SECs, leading to identifying the expression markers of Inner and Outer wall SECs, collector channel ECs, and distal region cells. Fourth, the identified unique markers were validated for RNA and protein expression in mouse ocular tissues. Fifth, the study explored how the IOP- and glaucoma-associated genes are expressed in the ScRNA- and snRNA-Seq data, providing potential connections of these GWAS genes with IOP and glaucoma. Sixth, the initial pathway and network analyses generated exciting hypotheses that could be tested in other independent studies.
Weaknesses:
A few minor weaknesses have been noted. First, since snRNA-Seq and scRNA-Seq generated different coverage of expressed genes in the cells, how did the combined analyses balance the un-equal sequencing coverage and missing data points in the snRNA-Seq data? Second, the RNA/protein validation of selected SEC molecular markers was done using mouse anterior segment tissues. It would be more helpful to examine whether these molecular markers for SECs could work well in human SECs. Third, the effort to characterize the GWAS-identified IOP- and glaucoma-associated genes is exciting but with limited new information. Additional work could be performed to prioritize these genes.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The authors carried out the current studies with the justification that the biochemical mechanisms that lead to alcohol addiction are incompletely understood. The topic and question addressed here are impactful and indeed deserve further research. To this end, a metabolomics approach toward investigating the metabolic effects of alcohol use disorder and the effect of alcohol withdrawal in AUD subjects is valuable. However, it is primarily descriptive in nature, and these data alone do not meet the stated goal of investigating biochemical mechanisms of alcohol addiction. The current work's most significant limitation is the cross-sectional study design, though inadequate description and citation of the underlying methodological approaches also hampers interest.
Most of the data are cross-sectional in the study design, i.e., alcohol use disorder vs controls. However, it is well established that there is a high degree of interpersonal variation with metabolism, and further, there is somewhat high intra-personal variation in metabolism over time. This means that the relatively small cohort of subjects is unlikely to reflect the broader condition of interest (AUD/withdrawal). The authors report a comparison of a later time-point after alcohol withdrawal (T2) vs. the AUD condition. However, without replicative time points from the control subjects it is difficult to assess how much of these changes are due to withdrawal vs the intra-personal variation described above. Overall, there is not enough experimental context to interpret these findings into a biological understanding. For example, while several metabolites are linked with AUD and associated with microbiome or host metabolism based on existing literature, it's unclear from the current study what function these changes have concerning AUD, if any. The authors also argue that alcohol withdrawal shifts the AUD plasma metabolic fingerprint towards healthy controls (line 153). However, this is hard to assess based on the plots provided since the change in the direction of the orange data subset is considers AUD T2 vs T1. In contrast, AUD T2 vs Control would represent the claimed shift. To support these claims, the authors would better support their argument by showing this comparison as well as showing all experimental groups (including control subjects) in their multi-dimensional model (e.g., PCA). The authors attempt to extend the significance of their findings by assessing post-mortem brain tissues from AUD subjects; however, the finding that many of the metabolites changed in T2/T1 are also present in AUD brain tissues is interesting; however, not strongly supporting of the authors' claims that these metabolites are markers of AUD (line 173). Concerning the plasma cohort itself, it is unclear how the authors assessed for compliance with alcohol withdrawal or whether the subjects' blood-alcohol levels were independently verified.
The second area of concern is the need for more description of the analytical methodology, the lack of metabolite identification validation evidence, and related statistical questions. The authors cite reference #59 regarding the general methodology. However, this reference from their group is a tutorial/review/protocol-focused resource paper, and it is needs to be clarified how specific critical steps were actually applied to the current plasma study samples given the range of descriptions provided in the citations. The authors report a variety of interesting metabolites, including their primary fragment intensities, which are appreciated (Supplementary Table 3), but no MS2 matching scores are provided for level 2 or 3 hits. Further, level 1 hits under their definition are validated by an in-house standard, but no supporting data are provided besides this categorization. Finally, a common risk in such descriptive studies is finding spurious associations, especially considering many factors described in the current work. These include AUD, depression, anxiety, craving, withdrawal, etc. The authors describe the use of BH correction for multiple-hypothesis testing. However, this approach only accounts for the many possible metabolite association tests within each comparison (such as metabolites vs depression). It does not account for the multi-variate comparisons to the many behavior/clinical factors described above. The authors should employ one of several common strategies, such as linear mixed effects models, for these types of multi-variate assessments.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors Fu et al., developed polymer models that combine loop extrusion with attractive interactions to best describe Hi-C population average data. They analyzed Hi-C data of the MYC locus as an example and developed an optimization strategy to extract the parameters that best fit this average Hi-C data.
Strengths:
The model has an intuitive nature and the authors masterfully fitted the model to predict relevant biology/Hi-C methodology parameters. This includes loop extrusion parameters, the need for self-interaction with specific energies, and the time and distance parameters expected for Hi-C capture.
Weaknesses:
(1) We are no longer in the age in which the community only has access to population average Hi-C. Why was only the population average Hi-C used in this study?
Can single-cell data: i.e. single-cell Hi-C/Dip-C data or chromatin tracing data (i.e. see Tan et al Science 2018 - for Dip-C, Bintu et al Science 2018, Su et al Cell 2020 for chromatin tracing, etc.) or even 2 color DNA FISH data (used here only as validation) better constrain these models? At the very least the simulations themselves could be used to answer this essential question.
I am expecting that the single-cell variance and overall distributions of distances between loci might better constrain the models, and the authors should at least comment on it.
(2) The authors claimed "Our parameter optimization can be adapted to build biophysical models of any locus of interest. Despite the model's simplicity, the best-fit simulations are sufficient to predict the contribution of loop extrusion and domain interactions, as well as single-cell variability from Hi-C data. Modeling dynamics enables testing mechanistic relationships between chromatin dynamics and transcription regulation. As more experimental results emerge to define simulation parameters, updates to the model should further increase its power." The focus on the Myc locus in this study is too narrow for this claim. I am expecting at least one more locus for testing the generality of this model.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The motivation for this study is to provide a comprehensive assessment of motor unit firing rate responses of entire pools during isometric contractions. The authors have used new quantitative methods to extract more unique motor units across contractions than prior studies. This was achieved by recording muscle fibre action potentials from four high-density surface electromyogram (HDsEMG) arrays (Caillet et al., 2023), quantifying residual EMG comparing the recorded and data-based simulation (Figure 1A-B), and developing a metric to compare the spatial identification for each motor unit (Figure 1D-E). From identified motor units, the authors have provided a detailed characterization of recruitment and firing rate responses during slow voluntary isometric contractions in the vastus lateralis and tibialis anterior muscles up to 80% of maximum intensity. In the lower limb, it is interesting how lower threshold motor units have firing rate responses that saturate, whereas higher threshold units that presumably produce higher muscle contractile forces continue to increase their firing rate. In many ways, these results agree with the rate coding of motor units in the extensor digitorum communis muscle (Monster and Chan, 1977). The paper is detailed, and the analyses are well explained. However, there are several points that I think should be addressed to strengthen the paper.
General comments:
(1) The authors claim they have measured the complete rate coding profiles of motor units in the vastus lateralis and tibialis anterior muscles. However, this study quantified rate coding during slow and prolonged voluntary isometric contractions whereas the function of rate coding during movements (Grimby and Hannerz, 1977) or more complex isometric contractions (Cutsem and Duchateau, 2005; Marshall et al., 2022) remains unexplored. For example, supraspinal inputs may not scale the same way across low and higher threshold motor units, or between muscles (Devanne et al., 1997), making the response of firing rates to increasing isometric contraction force less clear. Conceptually, the authors focus on the literature on intrinsic motoneurone properties, but in vivo, other possibilities are that descending supraspinal drive, spinal network dynamics, and afferent inputs have different effects across motor unit sizes, muscles, and types of contractions. Also, the influence from local muscles that act as synergists (e.g., vastii muscles for the vastus lateralis, and peroneal muscles that evert the foot for the tibialis anterior) or antagonists (coactivation during higher contraction intensities would stiffen the joint) may provide differential forms of proprioceptive feedback across motor pools.
(2) The evidence that the entire motor unit pool was recorded per muscle is not clear. There appears to be substantial residual EMG (Figure 1B), signal cancellation of smaller motor units (lines 172-176), some participants had fewer than 20 identified motor units, and contractions never went above 80% of MVC. Also, to my understanding, there remains no gold-standard in awake humans to estimate the total motor unit number in order to determine if the entire pool was decomposed. Furthermore, using four HDsEMG arrays also raises questions about how some channels were placed over non-target muscles, and if motor units were decomposed from surrounding synergists.
(3) The authors claim (Abstract L51; Discussion L376) that a commonly held view in the field is that rate coding is similar across motor units from the same pool. Perhaps this is in reference to some studies that have carefully assessed lower threshold motor units during lower force ramp contractions (e.g., Fuglevand et al., 2015; Revill and Fuglevand, 2017). However, a more complete integration of the literature exploring motor unit firing rate responses during rapid isometric contractions, comparing different muscles and contraction intensities would be helpful. From Figure 3, the range of rate coding in the tibialis anterior (~7-40 Hz) is greater than the vastus lateralis (~5-22 Hz) muscle across contraction levels. In agreement with other studies, the range of rate coding within some muscles is different than others (Kirk et al., 2021) and during maximal intensity (Bellemare et al., 1983) or rapid contractions (Desmedt and Godaux, 1978). Likewise, within a motor pool, there is a diversity of firing rate responses across motor units of different sizes as a function of isometric force (Monster and Chan, 1977; Desmedt and Godaux, 1977; Kukula and Clamann, 1981; Del Vecchio et al., 2019; Marshall et al., 2022). A strength of this paper is how firing rate responses are quantified across a wide range of motor unit recruitment thresholds and between two muscles. I suggest improving clarity for the general reader, especially in the motivation for testing two lower limb muscles, and elaborating on some of the functional implications.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Tóth and Bazeli et al. find rapamycin activates heterologously-expressed TRPM8 and dissociated sensory neurons in a TRPM8-dependent way with Ca2+-imaging. With electrophysiology and STTD-NMR, they confirmed the activation is through direct interaction with TRPM8. Using mutants and computational modeling, the authored localized the binding site to the groove between S4 and S5, different than the binding pocket of cooling agents such as menthol. The hydroxyl group on carbon 40 within the cyclohexane ring in rapamycin is indispensable for activation, while other rapalogs with its replacement, such as everolimus, still bind but cannot activate TRPM8. Overall, the findings provide new insights into TRPM8 functions and may indicate previously unknown physiological effects or therapeutic mechanisms of rapamycin.
Strengths:
The authors spent extensive effort on demonstrating that the interaction between TRPM8 and rapamycin is direct. The evidence is solid. In probing the binding site and the structural-function relationship, the authors combined computational simulation and functional experiments. It is very impressive to see that "within" a rapamycin molecule, the portion shared with everolimus is for "binding", while the hydroxyl group in the cyclohexane ring is for activation. Such detailed dissection represents a successful trial in the computational biology-facilitated, functional experiment-validated study of TRP channel structural-activity relationship. The research draws the attention of scientists, including those outside the TRP channel field, to previously neglected effects of rapamycin, and therefore the manuscript deserves broad readership.
Weaknesses:
The significance of the research could be improved by showing or discussing whether a similar binding pocket is present in other TRP channels, and hence rapalogs might bind to or activate these TRP channels. Additionally, while the finding on TRPM8 is novel, it is worthwhile to perform more comprehensive pharmacological characterization, including single-channel recording and a few more mutant studies to offer further insight into the mechanism of rapamycin binding to S4~S5 pocket driving channel opening. It is also necessary to know if rapalogs have independent or synergistic effects on top of other activators, including cooling agents and lower temperature, and their dependence on regulators such as PIP2.
Additional discussion that might be helpful:
The authors did confirm that rapamycin does not activate TRPV1, TRPA1 and TRPM3. But other TRP channels, particularly other structurally similar TRPM channels, should be discussed or tested. Alignment of the amino acid sequences or structures at the predicted binding pocket might predict some possible outcomes. In particular, rapamycin is known to activate TRPML1 in a PI(3,5)P2-dependent manner, which should be highlighted in comparison among TRP channels (PMID: 35131932, 31112550).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this article, Wen et. al. describe the development of a 'proof-of-concept' bi-functional vector based on HSV-deltaICP-34.5's ability to purge latent HIV-1 and SIV genomes from cells. They show that co-infection of latent J-lat T-cell lines with an HSV-deltaICP-34.5 vector can reactivate HIV-1 from a latent state. Over- or stable expression of ICP 34.5 ORF in these cells can arrest latent HIV-1 genomes from transcription, even in the presence of latency reversal agents. ICP34.5 can co-IP with- and de-phosphorylate IKKa/b to block its interaction with NF-k/B transcription factor. Additionally, ICP34.5 can interact with HSF1 which was identified by mass-spec. Thus, the authors propose that the latency reversal effect of HSV-deltaICP-34.5 in co-infected JLat cells is due to modulatory effects on the IKKa/b-NF-kB and PP1-HSF-1 pathway.
Next, the authors cleverly construct a bifunctional HSV-based vector with deleted ICP34.5 and 47 ORFs to purge latency and avoid immunological refluxes, and additionally, expand the application of this construct as a vaccine by introducing SIV genes. They use this 'vaccine' in mouse models and show the expected SIV-immune responses. Experiments in rhesus macaques (RM), further elicit the potential for their approach to reactivate SIV genomes and at the same time block their replication by antibodies. What was interesting in the SIV experiments is that the dual-functional vector vaccine containing sPD1- and SIV Gag/Env ORFs effectively delayed SIV rebound in RMs and in some cases almost neutralized viral DNA copy detection in serum. Very promising indeed, however, there are some questions I wish the authors had explored to get answers to, detailed below.
Overall, this is an elegant and timely work demonstrating the feasibility of reducing virus rebound in animals, with the potential to expand to clinical studies. The work was well-written, and sections were clearly discussed.
Strengths:
The work is well designed, rationale explained, and written very clearly for lay readers.
Claims are adequately supported by evidence and well-designed experiments including controls.
Weaknesses:
(1) While the mechanism of ICP34.5 interaction and modulation of the NF-kB and HSF1 pathways are shown, this only proves ICP34.5 interactions but does not give away the mechanism of how the HSV-deltaICP-34.5 vector purges HIV-1 latency. What other components of the vector are required for latency reversal? Perhaps serial deletion experiments of the other ORFs in the HSV-deltaICP-34.5 vector might be revealing.
(2) The efficacy of the HSV vaccine vectors was evaluated in Rhesus Macaque model animals. Animals were chronically infected with SIV (a parent of HIV), treated with ART, challenged with bi-functional HSV vaccine or controls, and discontinued treatment, and the resulting virus burden and immune responses were monitored. The animals showed SIV Gag and Env-specific immune responses, and delayed virus rebound (however rebound is still there), and below-detection viral DNA copies. What would make a more convincing argument to this reviewer will be data to demonstrate that after the bi-functional vaccine, the animals show overall reduction in the number of circulating latent cells. The feasibility of obtaining such a result is not clearly demonstrated.
(3) The authors state that the reduced virus rebound detected following bi-functional vaccine delivery is due to latent genomes becoming activated and steady-state neutralization of these viruses by antibody response. This needs to be demonstrated. Perhaps cell-culture experiments from specimens taken from animals might help address this issue. In lab cultures one could create environments without antibody responses, under these conditions one would expect a higher level of viral loads to be released in response to the vaccine in question.
(4) How do the authors imagine neutralizing HIV-1 envelope epitopes by a similar strategy? A discussion of this point may also help.
(5) I thought the empty HSV-vector control also elicited somewhat delayed kinetics in virus rebound and neutralization, can the authors comment on why this is the case?
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This paper derives the first three functional gradients in the left and right hippocampus across two datasets. These gradient maps are then compared to dopamine receptor maps obtained with PET, associated with age, and linked to memory. Results reveal links between dopamine maps and gradient 2, age with gradients 1 and 2, and memory performance.
Strengths:
This paper investigates how hippocampal gradients relate to aging, memory, and dopamine receptors, which are interesting and important questions. A strength of the paper is that some of the findings were replicated in a separate sample.
Weaknesses
The paper would benefit from added clarification on the number of models/comparisons for each test. Furthermore, it would be helpful to clarify whether or not multiple comparison correction was performed and - if so - what type or - if not - to provide a justification. The manuscript would furthermore benefit from code sharing and clarifying which results did/did not replicate.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Summary:
This paper describes transcriptomes from three tardigrade species with or without treatment with ionizing radiation (IR). The authors show that IR produces numerous single strand and double strand breaks as expected and that these are substantially repaired within 4-8 hours. Treatment with IR induces strong upregulation of transcripts from numerous DNA repair proteins, and from the newly described protein TDR1 with homologs in both Hypsibioidea and Macrobiotoidea supefamilies. The authors show that TDR1 transcription produces newly translated TDR1 protein, which can bind DNA and co-localizes with DNA in the nucleus. At higher concentrations TDR appears to form aggregates with DNA, which might be relevant to a possible function in DNA damage repair. When introduced into human U2OS cells treated with the radiomimetic drug bleomycin, TDR1 reduces the number of double-strand breaks as detected by gamma H2AX spots. This paper will be of interest to the DNA repair field and to radiobiologists.
Strengths:
The paper is well-written and provides solid evidence of the upregulation of DNA repair enzymes after irradiation of tardigrades, as well as upregulation of the TRD1 protein. The reduction of gamma-H2A.X spots in U2OS cells after expression of TRD1 supports a role in a DNA damage.
Weaknesses:<br /> Genetic tools are still being developed in tardigrades, so there is no mutant phenotype to support a DNA repair function for TRD1, but this may be available soon.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
In this study, Huang et al. performed a scRNA-seq analysis of lung adenocarcinoma (LUAD) specimens from 9 human patients, including 5 who received neoadjuvant chemotherapy (NCT), and 4 without treatment (control). The new data was produced using 10 × Genomics technology and comprises 83622 cells, of which 50055 and 33567 cells were derived from the NCT and control groups, respectively. Data was processed via R Seurat package, and various downstream analyses were conducted, including CNV, GSVA, functional enrichment, cell-cell interaction, and pseudotime trajectory analyses. Additionally, the authors performed several experiments for in vitro and in vivo validation of their findings, such as immunohistochemistry, immunofluorescence, flow cytometry, and animal experiments.
The study extensively discusses the heterogeneity of cell populations in LUAD, comparing the samples with and without chemotherapy. However, there are several shortcomings that diminish the quality of this paper:
• The number of cells included in the dataset is limited, and the number of patients from different groups is low, which may reduce the attractiveness of the dataset for other researchers to reuse. Additionally, there is no metadata on patients' clinical characteristics, such as age, sex, history of smoking, etc., which would be valuable for future studies.<br /> • Several crucial details about the data analysis are missing: How many PCs were used for reduction? Which versions of Seurat/inferCNV/other packages were used? Why monocle2 was used and not monocle3 or other packages? Also, the authors use R version 3.6.1, and the current version is 4.3.2.<br /> • It seems that the authors may lack a fundamental understanding of scRNA-seq data processing and the functions of Seurat. For instance, they state, 'Next, we classified cell types through dimensional reduction and unsupervised clustering via the Seurat package.' However, dimensional reduction and unsupervised clustering are not methods for cell classification. Typically, cell types are classified using marker genes or other established methods.<br /> "Therefore, to identify subclusters within each of these nine major cell types, we performed principal component analysis" (Line 127). Principal component analysis is a method for dimensionality reduction, not cell clustering.<br /> The authors did not mention the normalization or scaling of the data, which are crucial steps in scRNA-seq data preprocessing.<br /> • Numerous style and grammar mistakes are present in the main text. For instance, certain sections of the methods are written in the present tense, suggesting that parts of a protocol were copied without text editing. Furthermore, some sections of the introduction are written in the past tense when the present tense would be more suitable. Clusters are inconsistently referred to by numbers or cell types, leading to confusion. Additionally, the authors frequently use the term "evolution" when describing trajectory analysis, which may not be appropriate. Overall, significant revisions to the main text are required.<br /> • Some figures are not mentioned in order or are not referenced in the text at all, such as Figure 5l (where it is also unclear how the authors selected the root cells). Additionally, many figures have text that is too small to be read without zooming in. Overall, the quality of the figures is inconsistent and sometimes very poor.<br /> • At times, the authors' statements are incomplete (ex. Lines 67-69, Line 177, Line 629, Lines 646-648 and 678).
The results section lacks clarity on several points:<br /> • The authors state that "myofibroblasts exclusively originated from the control group". However, pathways up-regulated in myofibroblasts (such as glycolysis) were enhanced after chemotherapy, as indicated by GSVA score. Similarly, why are some clusters of TAMs from the control group associated with pathways enriched in chemotherapy group?<br /> • Further explanation is necessary regarding the distinctions between malignant and non-malignant cells, as well as regarding the upregulation of metabolism-related pathways in fibroblasts from the NCT group. Additionally, clarification is needed regarding why certain TAMs from the control group are associated with pathways enriched in the chemotherapy group.<br /> • In the section titled 'Chemo-driven Pro-mac and Anti-mac Metabolic Reprogramming Exerted Diametrically Opposite Effects on Tumor Cells': The markers selected to characterize the anti- and pro-macrophages are commonly employed for describing M1 or M2 polarization. It is uncertain whether this new classification into anti- and pro-macrophages is necessary. Additionally, it should be noted that pro-macrophages are anti-inflammatory, while anti-macrophages are pro-inflammatory, which could lead to confusion. M2 macrophages are already recognized for their role in stimulating tumor relapse after chemotherapy.<br /> • The authors suggest that there is "reprogramming of CD8+ cytotoxic cells" following chemotherapy (Line 409). It remains unclear whether they imply the reprogramming of other CD8+ T cells into cytotoxic cells. While it is indicated that cytotoxic cells from the control group differ from those in the NCT group and that NCT cytotoxic T cells exhibit higher cytotoxicity, the authors did not assess the expression of NK and NK-like T cell markers (aside from NKG7), which may possess greater cytotoxic potential than CD8+ cytotoxic cells. This could also elucidate why cytotoxic cells from the NCT and control groups are positioned on separate branches in trajectory analysis. Overall, with 22.5k T cells in the dataset, only 3 subtypes were identified, suggesting a need for improved cell annotations by the authors.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The article uses a cell-based model to investigate how mutations and cells spread throughout a tumour. The paper uses published data and the proposed model to understand how growth and death mechanisms lead to the observed data. This work provides an insight into the early stages of tumour development. From the work provided here, the results are solid, showing a thorough analysis. However, the work has not fully specified the model, which can lead to some questions around the model's suitability. The article is well-written and presents a very suitable and rigorous analysis to describe the data. The authors did a particularly nice job of the discussion and decision of their "metrics of interest", though this is not the main aim of this work.
Strengths:
Due to the particularly nice and tractable cell-based model, the authors are able to perform a thorough analysis to compare the published data to that simulated with their model. They then used their computational model to investigate different growth mechanisms of volume growth and surface growth. With this approach, the authors are able to compare the metric of interest (here, the direction angle of a new mutant clone, the dispersion of mutants throughout the tumour) to quantify how the different growth models compare to the observed data. The authors have also used inference methods to identify model parameters based on the data observed. The authors performed a rigorous analysis and have chosen the metrics in an appropriate manner to compare the different growth mechanisms.
Weaknesses:
The work contained within this article considers a single cell-based model. While ideally, this is sufficient, results from simulated multi-cellular systems can often be sensitive to the model choice. Performing this work with various other standard models would strengthen the results significantly. This is, however, not an easy task.
Context:
Improved mechanistic understanding into the early developmental stages of tumours will further assist in disease treatment and quantification. Understanding how readily and quickly a tumour is evolving is key to understanding how it will develop and progress. This work provides a solid example as to how this can be achieved with data alongside simulated models.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
In this study, the authors assess whether selective pressure from drug chemotherapy influences the emergence of drug resistance through the acquisition of genetic mutations or phenotypic tolerance. I commend the authors on their approach of utilizing the mutation accumulation (MA) assay as a means to answer this and whole genome sequencing of clones from the assay convincingly demonstrates low mutation rates in Mycobacteria when exposed to sub-inhibitory concentrations of antibiotics. Also, quantitative PCR highlighted the upregulation of DNA repair genes in Mycobacteria following drug treatment, implying the preservation of genomic integrity via specific repair pathways.
Even though the findings stem from M. smegmatis exposure to antibiotics under in vitro conditions, this is still relevant in the context of the development of drug resistance so I can see where the authors' train of thought was heading in exploring this. However, I think important experiments to perform to more fully support the conclusion that resistance is largely associated with phenotypic rather than genetic factors would have been to either sequence clones from the ciprofloxacin tolerance assay (to show absence/ minimal genetic mutations) or to have tested the MIC of clones from the MA assay (to show an increase in MIC). There seems to be a disconnect between making these conclusions from experiments conducted under different conditions, or perhaps the authors can clarify why this was done. With regards to the sub-inhibitory drug concentration applied, there is significant variation in the viability as calculated by CFUs following the different treatments and there is evidence that cell death greatly affects the calculation of mutation rate (PMCID: PMC5966242). For instance, the COMBO treatment led to 6% viability whilst the INH treatment led to 80% cell viability. Are there any adjustments made to take this into account? It would also be useful to the reader to include a supplementary table of the SNPs detected from the lineages of each treatment - to determine if at any point rifampicin treatment led to mutations in rpoB, isoniazid to katG mutations, etc. Overall, while this study is tantalizingly suggestive of phenotypic tolerance playing a leading role in drug resistance (and perhaps genetic mutations a sub-ordinate role) a more substantial link is needed to clarify this.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
SNX4 is thought to mediate recycling from endosomes back to the plasma membrane in cells. In this study, the authors demonstrate the increases in the amounts of transmitter release and the number of docked vesicles by combining genetics, electrophysiology, and EM. They failed to find evidence for its role in synaptic vesicle cycling and endocytosis, which may be intuitively closer to the endosome function.
Strengths:
The electrophysiological data and EM data are in principle, convincing, though there are several issues in the study.
Weaknesses:
It is unclear why the increase in the amounts of transmitter release and docked vesicles happened in the SNX4 KO mice. In other words, it is unclear how the endosomal sorting proteins in the end regulate or are connected to presynaptic, particularly the active zone function.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The paper entitled "Goal-directed motor actions drive acetylcholine dynamics in sensory cortex" aims to characterize the dynamics of cholinergic signaling in sensory cortex during perceptual behavior. The authors showed that acetylcholine release in S1 was linked to goal-directed motor actions rather than sensory input or reward delivery, a pattern also observed in the auditory cortex (A1). This release was specifically associated with whisking and licking and was potentiated by training. The results contribute to a better understanding of neuromodulator actions. That said, several aspects of the manuscript could benefit from improved writing, data presentation, and statistical analysis.
Strengths:
The evidence provided is clear to link ACh response to different task-related events. Implementing two different tasks to show generality is appreciated. Important control analysis is included.
Weaknesses:
The quantification of ACh signal differences across different trial types or between expert and early-training mice is lacking. Although statistical significance is occasionally mentioned, the indication of significance in figures seems rare. For example, in Figures 5A and E, it is difficult to tell when p is < 0.05. Based on the sentence "small, but significant increase on Hits over False Alarm trials (Figure 5A, S Figure 4A)" there is indeed a time point where the difference is significant, and more details should be added (when and the p-value).
For Figure 5D, it seems like there is no significant difference between Hit and False alarm trials, however, for the trials with 1 or 2 lick there appears to be a difference. Is it due to a lack of power? Moreover, in Figure 5 H the first licks also seem to differ.
Linear regression: the coefficient of determination (R²) is absent, in Figures 4E, F, and 6B, H, making it hard to evaluate the goodness of the fitting.
Similar comments apply to Figure 7: the lack of quantitative comparisons between the coefficients of first lick and other regressors, and between early and expert training, as well as the change in goodness of fit by removing a regressor.
The writing of the introduction and discussion could be improved to enhance readability, and the manuscript could improve its discussion on orofacial movement and acetylcholine release by citing relevant studies demonstrating the association between neuronal activity and orofacial/body movements.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
While many studies have explored the impacts of pathogens on hosts, the effect of hosts on pathogens has received less attention. In this manuscript, Wang et al. utilize Drosophila melanogaster and an opportunistic pathogen, Serratia marcescens, to explore how the host impacts pathogenicity. Beginning with an observation that larval presence and density impacted microbial growth in fly vials (which they assess qualitatively as the amount of 'slick' and quantitatively as microbial load/CFUs), the authors focus on the impact of axenic/germ-free larvae on an opportunistic pathogen S. marcescens. Similar to their observations with general microbial load, they find that larvae reduce the presence of a pinkish slick of Sm, indicative of its secondary metabolite prodigiosin. The presence of larvae alters prodigiosin production, pathogen load, pathogen cellular morphology, and virulence, and this effect is through transcriptional and metabolic changes in the pathogen. Overall, they observe a loss of virulence factors/pathways and an increase in pathways contributing to growth. Given the important role the host plays in this lifestyle shift, the authors then examined host features that might influence these effects, focusing on the role of antimicrobial peptides (Amps). The authors combine the use of synthetic Amps and an Amp-deficient fly line and conclude much of the larval inhibitory effect is due to their production of AMPs.
Strengths:
This is a very interesting question and the use of Drosophila-Serratia marcescens is a great model to explore these interactions and effects.
The authors have an interesting and compelling phenotype and are asking a unique question on the impact of the host on the pathogen. The use of microbial transcriptomics and metabolomics is a strength, especially in order to assess these impacts on the pathogen level and at single-cell level to capture heterogeneity.
Weaknesses:
Overall, the writing style in the manuscript makes it difficult to fully understand and appreciate the data and its interpretation.
The data on the role of AMPs would benefit from strengthening. Some of the arguments in the text of that section are also counterintuitive. The authors show that AMP larvae have a reduced impact on Sm as compared to wt larvae, but it seems less mild of an effect than that observed with wt excreta (assuming the same as secreta in Figures 7, should be corrected or harmonized). Higher doses of AMPs give a phenotype similar to wt larvae, but a lower dose (40 ng/ul) gives phenotypes more similar to controls. The authors argue that this data suggests AMPs are the factor responsible for much of the inhibition, but their data seems more to support that it's synergistic- you seem to still need larvae (or some not yet defined feature larvae make, although secreta/excreta was not sufficient) + AMPs to see similar effects as wt. Based on positioning and color scheme guessing that AMP 40ng/ul was used in Figures 7D-H, but could not find this detail in the text, methods, or figure legend and it should be indicated. This section does not seem to be well supported by the provided data, and this inconsistency greatly dampened this reviewer's enthusiasm for the paper.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The manuscript illustrates how spatial targeting (perisomatic vs distal, apical, and basal dendritic) and timing of inhibition are crucial to distinct effects on neuronal integration and show that beta and gamma oscillations differentially engage dendritic spiking mechanisms.
Strengths:
The strength of this study lies in the integrative biophysical modelling of a layer 5 pyramidal neuron by bringing together in vitro and in vivo observations.
Weaknesses:
The weaknesses are probably in some of the parameterizations of inhibitory synaptic dynamics. A unitary peak conductance of 1nS is very high for inhibitory synapses. This high value could invariably skew some of the network-level predictions. The authors could obtain specific parameters from the Neocortical Collaboration Portal (https://bbp.epfl.ch/nmc-portal/microcircuit.html), which is an incredible resource for cortical neurons and synapses.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, the authors examine the question of whether discrete action sequences and coarticulated continuous sequential actions can be produced from the same controller, without having to derive separate control policies for each sequential movement. Using modeling and behavioral experiments, the authors demonstrate that this is indeed possible if the constraints of the policy are appropriately specified. These results are of interest to those interested in motor sequences, but it is unclear whether these findings can be interpreted to apply to the control of sequences more broadly (see weaknesses below).
Strengths:
The authors provide an interesting and novel extension of the stochastic optimal control model to demonstrate how different temporal constraints can lead to either individual or coarticulated movements. The authors use this model to make predictions about patterns of behavior (e.g., in response to perturbations), which they then demonstrate in human participants both by measuring movement kinematics as well as EMG. Together this work supports the authors' primary claims regarding how changes in task instructions (i.e., task constraints) can result in coarticulated or separated movement sequences and the extent to which the subsequent movement goal affects the planning and control of the previous movement.
Weaknesses:
I reviewed a prior version of this manuscript, and appreciate the authors addressing many of my previous comments. However, there are some concerns, particularly with regard to how the authors interpret their findings.
(1) It would be helpful for the authors to discuss whether they think there is a fundamental distinction between a coarticulated sequence and a single movement passing through a via point (or equivalently, avoiding an obstacle). The notion of a coarticulated sequence brings with it the notion of sequential (sub)movements and temporal structure, whereas the latter can be treated as more of a constraint on the production of a single continuous movement. If I am interpreting the authors' findings correctly it seems they are suggesting that these are not truly different kinds of movements at the level of a control policy, but it would be helpful for the authors to clarify this claim.
(2) The authors' model clearly shows that each subsequent target only influences the movement of one target back, but not earlier ones (page 7 lines 199-204). This stands in contrast to the paper they cite from Kashefi 2023, in which those authors clearly show that people account for at least 2 targets in the future when planning/executing the current movement. It would be useful to know whether this distinction arises because of a difference in experimental methodology, or because the model is not capturing something about human behavior.
(3) In my prior review I raised a concern that the authors seem to be claiming that because they can use a single control policy for both coarticulated and separated movement sequences, there need not be any higher-level or explicit specification of whether the movements are sequential. While much of that language has been removed, it still appears in a few places (e.g., p. 13, lines 403-404). As previously noted, the authors' control policy can generate both types of movements as long as the proper constraints are provided to the model. However, these constraints must be specified somewhere (potentially explicitly, as the authors do by providing them as task instructions). Moreover, in typical sequence tasks, although some movements become coarticulated, people also tend to form chunks with distinct chunk boundaries, which presumably means that there is at least some specification of the sequential ordering of these chunks that must exist (otherwise the authors' model might suggest that people can coarticulate forever without needing to exhibit any chunk boundaries). Hence the authors should limit themselves to the narrow claim that a single control policy can lead to separated or coarticulated movements given an appropriate set of constraints, but acknowledge that their work cannot speak to where or how those constraints are specified in humans (i.e., that there could still be an explicit sequence representation guiding coarticulation).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript aims to tackle the antimicrobial resistance through the development of vaccines. Specifically, the authors test the potential of the RSP protein as a vaccine candidate. The RSP protein contains bacterial Ig-like domains that are typically carried in IncHl1 plasmids like R27. The extracellular location of the RSP protein and its role in the conjugation process makes it a good candidate for a vaccine. The authors then use Salmonella carrying an IncHl plasmid to test the efficacy of the RSP protein as a vaccine antigen in providing protection against infection of antibiotic-resistant bacteria carrying the IncHl plasmid. The authors found no differences in total IgG or IgA levels, nor in pro-inflammatory cytokines between immunized and non-immunized mice. They however found differences in specific IgG and IgA, attenuated disease symptoms, and restricted systemic infection.
The manuscript also evaluates the potential use of nanobodies specifically targeting the RSP protein by expressing it in E. coli and evaluating their interference in the conjugation of IncHl plasmids. The authors found that E. coli strains expressing RSP-specific nanobodies bind to Salmonella cells carrying the R27 plasmid thereby reducing the conjugation efficacy of Salmonella.
Strengths:
- The main strength of this manuscript is that it targets the mechanism of transmission of resistance genes carried by any bacterial species, thus making it broad.
- The experimental setup is sound and with proper replication.
Weaknesses:
- The two main experiments, evaluating the potential of the RSP protein and the effects of nanobodies on conjugation, seem as parts of two different and unrelated strategies.
- The survival rates shown in Figure 1A and Figure 3A for Salmonella pHCM1 and non-immunized mice challenged with Salmonella, respectively, are substantially different. In the same figures, the challenge of immunized mice and Salmonella pHCM1 and mice challenged with Salmonella pHCM1 with and without ampicillin are virtually the same. While this is not the only measure of the effect of immunization, the inconsistencies in the resulting survival curves should be addressed by the authors more thoroughly as they can confound the effects found in other parameters, including total and specific IgG and IgA, and pro-inflammatory cytokines.
- Overall the results are inconsistent and provide only partial evidence of the effectiveness of the RSP protein as a vaccine target.
- The conjugative experiments use very long conjugation times, making it harder to asses if the resulting transconjugants are the direct result of conjugation or just the growth of transconjugants obtained at earlier points in time. While this could be assessed from the obtained results, it is not a direct or precise measure.
- While the potential outcomes of these experiments could be applied to any bacterial species carrying this type of plasmids, it is unclear why the authors use Salmonella strains to evaluate it. The introduction does a great job of explaining the importance of these plasmids but falls short in introducing their relevance in Salmonella.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The study is devoted to the deep investigation of the spermatogonial stem cell (SSC) niche in trans women after gender-affirming hormone therapy (GAHT). Both cellular structure and functionality of the niche were studied. The authors evidently demonstrated that all cellular components of SSC niche were affected by hormone therapy. Interestingly, the signs of "rejuvenation" within the niche were also observed indicating the possible reverse to the immature condition.
Strengths:
The obtained findings are important for the better understanding of hormonal regulation of testis and SSC niche and provide some clues for using the biomaterials from these specific and even unique donors for biomedical research.
Weaknesses:
This study has some limitations. Many studies can't be done using the testes cells of trans women, since their cells are significantly different from adult man cells and less from prepubertal and pubertal cells. The authors themselves identify some of the limitations: this material is suitable only for studying prepubertal processes in the testis. However, the authors also report large variability in data due to different hormonal therapy regimens and, apparently, age. Accordingly, not all material obtained from trans women can also be used for studies of prepubertal processes.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Bowman and colleagues have compiled a large comparative genomic dataset to examine the molecular evolution of genes in mammals, with the primary goal of identifying how changes in the gorilla mating system have shaped the evolution of spermatogenesis. They report several patterns pointing to signal of relaxed purifying selection on genes involved in male fertility, a pattern that they interpret as a response to changes in the mating system of gorillas. Many previous studies have used comparisons among species of primates and other mammals to understand how changes in mating systems have shaped the evolution or reproductive traits and genes. These collective works have provided some of the best evidence that changes in the form and intensity of sexual selection has had a strong effect on the evolution of male reproduction. The current study builds on this rich history by exploring molecular evolution of over 13,310 genes across 261 mammals. This very large phylogenetic dataset allows affords considerable power to characterize patterns of molecular evolution along the gorilla lineage. This allows for some added power relative to a previous study that interrogated the same lineage-specific patterns (Scally et al. 2021). They report a subset of genes showing evidence for either positive directional selection (less than 1% of genes) or relaxed purifying selection (4% of genes) in gorillas. Relaxed purifying selection is more common than positive selection, and genes showing signatures of relaxed constraint are enriched for spermatogenesis functions using various tests based on functional annotation or gene expression and infertility associations in humans and mice. The authors also report new functional data - the only original data in this study - using a high throughput genetic screen showing that some of these genes are also expressed in spermatogenesis in flies, and when perturbed they affect male fertility.
These results are interpreted as strong evidence that changes in mating system, specifically that loss of sperm competition, has shaped the evolution of male reproduction in gorillas. The authors argue that these discoveries illustrate, for the first time, the genome-wide effect of striking changes in mating behavior in gorillas on the genetic underpinnings of male reproduction and provide new candidates relevant to male fertility in humans. Support for these central conclusions is eroded by a lack of appropriate comparative contrasts needed clarify the uniqueness of these patterns to gorillas and, critically, establish a direct phylogenetic association with mating system or correlated reproductive traits.
Strengths:
The presentation is engaging, clear, and easy to follow throughout. I enjoyed reading the overall narrative and I think that the authors did a good job of presenting the details of male reproductive biology in an informative and accessible manner. Given the general interest in gorilla evolution, and the clear relevance to humans, studies of this scope on male reproductive biology are likely to be of broad interest to both evolutionary and reproductive biologists.
The reported signatures of molecular evolution in gorillas appear robust, well-executed, and supported by several lines of evidence that establish some links with male reproduction. The authors have presented a series of molecular evolution analyses that demonstrate both rigor and attention to analytical details and quality control. Although all the primary sequence data has been previously published by others, the compilation of a high-quality curated comparative dataset of this scale is impressive and inspires confidence in the underlying molecular results. Likewise, the incorporation of diverse other data from mice and humans helps shape the overall narrative. To my knowledge, this represents the most focused and detailed analysis of protein-coding evolution specific to gorillas to date (although parallel results from the landmark gorilla genome study - Scally et al. 2012 - are downplayed somewhat).
Likewise, the inclusion of new functional data from Drosophila establishes a subset of genes showing recent changes in molecular evolution in gorillas that appear to be both deeply conserved in animals and related to male fertility.
Weaknesses:
This study lacks the necessary comparative framework needed to ascribe any of the reported patterns to changes in the reproductive system of gorillas, or to really understand the uniqueness of these patterns relative to other species. Although wording is careful at times, the authors repeatedly ascribe the patterns they are finding directly to the specific changes in mating system biology that has occurred in gorillas. The general framing and significance rests on the central finding that "these data provide compelling evidence that reduced sperm competition in gorillas is associated with relaxed purifying selection on genes related to male reproductive function (Abstract)". No such association between variation in mating system or at any correlated reproductive traits and molecular evolution is ever directly tested let alone established as a clear statistical correlation. The massive comparative dataset is used to localize patterns of molecular evolution to the gorilla lineage and then these patterns are interpreted in the context of changes in mating system, as an assumption of the study not a direct result. Although basic information of the reproductive system (or correlates thereof) likely exists for many of the 261 species included here, this information is never used to test for a relationship between changes in positive or purifying selection and reproduction.
The lack of any such comparisons is especially curious given that there are many previous studies that have sought and established such connections for traits and/or genes in mammals (dozens now?), and especially great apes, before. This comparative approach is the gold standard to making claims linking mating system to molecular evolution and yet this is not pursued here. The authors are correct in that they provide a rigorous genome-wide analysis (but not at all for the first time, see Scally et al. 2012), but they skip this critical central step to rigorous inference in comparative genomics. This is essentially a broad comparative study, but the central conclusion (a direct link between mating system and molecular evolution) is speculative and not actually tested.
Note that despite the framing here, there are of course several aspects of lineage specific biology that undoubtedly shape molecular evolution of male reproduction and fertility but could be unrelated to sperm competition per se. For example, shift in operational sex ratios can have profound effects on effective population sizes and the efficacy of selection, which of course would be expected to change the intensity and direction of molecular evolution. Likewise, shifts in population size, structure, and diet all can affect molecular evolution and reproduction.
In the absence of a broad phylogenetically independent contrast (which would be really interesting here), the authors need to at least establish that there is indeed something noteworthy about the specific findings they report relative to other systems that have a different mating system. Such comparisons would be readily available within the great apes, especially compared to chimpanzees and bonobos (Pan). Most of the patterns are presented in such a way to suggest a clear connection between the result and the unique features of gorilla reproduction, but are these clearly outliers? Relaxed purifying selection is much more common than positive selection, is this result qualitatively or quantitatively unique to gorillas as implied (I would honestly be surprised if it was as this is a common outcome of these dn/ds-based tests)? Similar questions and the need for more context apply to the various enrichment tests. That genes involved in male reproduction evolve rapidly and that this reflects both relaxed constraint and positive selection is an exceptionally well-established pattern, as is enrichment for reproductive functions/expression of such genes in unbiased genome-wide screens (as cited by the authors, including in gorillas by Scally et al. 2012 who performed a very similar analysis albeit with some model advances used in the current study). Do chimpanzees or humans lack these specific signatures of relaxed constraint at reproductive genes or is it a much stronger enrichment in gorillas? Establishing these baseline comparisons would help a lot with interpretation of the core findings. A little bit of this is explored with the human comparisons but not in a parallel genome-wide manner that places the signatures in gorillas in context.
I had similar questions related to the high-throughput Drosophila screen. This is a creative and novel component of the study. However, I am unclear on how to interpret the results or the conclusions drawn from them. It is very interesting that a subset of genes showing relaxed constraint are conserved to Drosophila and that perturbation of some of these cause fertility issues. However, the conclusion that these genes reflect novel candidates not implicated in sperm biology is a bit overstated. Here implicated means genes with an annotated sterility phenotype in humans, mice, flies, or gorillas - specific annotations which are pretty limited at least in the mammalian systems. The entire design was conditioned on analyzing genes that were reliably expressed during Drosophila spermatogenesis, and then focusing on those. But the comparative set for the enrichment test was a random set of genes. Shouldn't the background be a random set of testis-expressed genes? I would say that genes that are reliably expressed during spermatogenesis in both mammals and flies are implicated in sperm biology and genetic manipulation of such genes would be expected to produce fertility phenotypes at some appreciable rate. So the result here adds some interesting data but it does not seem unexpected or significant as framed.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The paper 'Sequence characteristic and an accurate model of abundant hyperactive loci in human genome' by Hydaiberdiev and Ovcharenko offers comprehensive analyses and insights about the 'high-occupancy target' (HOT) loci in the human genome. These are considered genomic regions that overlap with transcription factor binding sites. The authors provided very comprehensive analyses of the TF composition characteristics of these HOT loci. They showed that these HOT loci tend to overlap with annotated promoters and enhancers, GC-rich regions, open chromatin signals, and highly conserved regions, and that these loci are also enriched with potentially causal variants with different traits.
Strengths:
Overall, the HOT loci' definition is clear and the data of HOT regions across the genome can be a useful dataset for studies that use HepG2 or K562 as a model. I appreciate the authors' efforts in presenting many analyses and plots backing up each statement.
Weaknesses:
It is noteworthy that the HOT concept and their signature characteristics as being highly functional regions of the genome are not presented for the first time here. Additionally, I find the main manuscript, though very comprehensive, long-winded and can be put in a shorter, more digestible format without sacrificing scientific content.
The introduction's mention of the blacklisted region can be rather misleading because when I read it, I was anticipating that we are uncovering new regulatory regions within the blacklisted region. However, the paper does not seem to address the question of whether the HOT regions overlap, if any, with the ENCODE blacklisted regions afterward. This plays into the central assessment that this manuscript is long-winded.
The introduction also mentioned that HOT regions correspond to 'genomic regions that seemingly get bound by a large number of TFs with no apparent DNA sequence specificity' (this point of 'no sequence specificity' is reiterated in the discussion lines 485-486). However, later on in the paper, the authors also presented models such as convolutional neural networks that take in one-hot-encoded DNA sequence to predict HOT performed really well. It means that the sequence contexts with potential motifs can still play a role in forming the HOT loci. At the same time, lines 59-60 also cited studies that "detected putative drive motifs at the core segments of the HOT loci". The authors should edit the manuscript to clarify (or eradicate) contradictory statements.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this study, Styer et al. impose artificial selection on root-associated microbiomes to increase drought tolerance in rice plants using different soils as starting microbiomes. Using NDVI and biomass as a proxy for plant health, they find that iterative passaging of the microbiomes of the best-performing plants increased plant resilience to drought stress in a soil-dependent manner. The study makes use of numerous controls. The authors survey the microbiota of the plants across generations, using an array of interesting analyses to characterize their observations. Firstly, the authors find that the acquired microbiomes are divergent towards the beginning of the selection experiment, but nearly converge later suggesting that the selected communities become more similar over time. One reason is that the diversity of the microbiomes severely decreases after only one or two generations of selection AND that microbes from each inoculation source appear to easily disperse across the experiment, leading to microbiome homogeneity. The authors then present an analysis to correlate ASVs with the NDVI and Biomass over the course of the experiment (using the rice soil selection lines) to develop hypotheses about which ASVs may impact plant traits.
Strengths:
The authors set out to refine the understanding of microbiome artificial selection, a topic of recent interest to the plant microbiome field. The authors use an established approach (Mueller et al), expanding upon it by including multiple starting soil inocula to ask whether the strength of selection varies by input microbiome. This is an important and novel question. Using drought resilience as measured by NDVI and plant biomass to select upon was a wise choice for this type of study, given their relative ease and quickness to assess. The inclusion of several types of controls, multiple selection lines, and several starting soil inocula showed a thoughtful experimental design. The analyses were diverse, non-standard, and attempted to address microbiome dynamics on multiple fronts. I am not necessarily convinced by some of the conclusions (see below), however, I think this study examines an important and exciting topic in the area of plant microbiomes. I predict the findings of the experiments will inform a wide audience of researchers attempting similar studies and be helpful in their designs.
Weaknesses:
Although the controls were well designed, the dispersal of the microbiomes erased the utility of the sterile inoculated (SI) controls, at least from my reading of the manuscript. Perhaps the original intent of the SI plants was to contrast the selected microbiomes vs axenic plants to show that plant resilience to drought increased generation after generation. If the controls had worked properly under my presumed scenario, this would allow the authors to account for batch variation across the generations (due to slight differences in MS media prep, water quality, etc.). Instead, the SI lines acquired microbes from the experiment and never appeared to significantly deviate from the SL plants. The dispersal of the microbes amongst soils and selection lines also minimizes any conclusions that can be made about the different starting inocula and how prone to selection they may be.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This is a short and unpretentious paper. It is an interesting area and therefore, although much of this area of research was pioneered in flies, extending basic findings to butterflies would be worthwhile. Indeed, there is an intriguing observation but it is technically flawed and these flaws are serious.
The authors show that mirror is expressed at the back of the wing in butterflies (as in flies). They present some evidence that is required for the proper development of the back of the wing in butterflies (a region dubbed the vannus by the ancient guru Snodgrass). But there are problems with that evidence. First, concerning the method, using CRISP they treat embryos and the expectation is that the mirror gene will be damaged in groups of cell lineages, giving a mosaic animal in which some lines of cells are normal for mirror and others are not. We do not know where the clones or patches of cells that are defective for mirror are because they are not marked. Also, we do not know what part of the wing is wild type and what part is mutant for mirror. When the mirror mutant cells colonise the back of the wing and that butterfly survives (many butterflies fail to develop), the back of the wing is altered in some selected butterflies. This raises a second problem: we do not know whether the rear of the wing is missing or transformed. From the images, the appearance of the back of the wing is clearly different from the wild type, but is that due to transformation or not? And then I believe we need to know specifically what the difference is between the rear of the wing and the main part. What we see is a silvery look at the back that is not present in the main part, is it the structure of the scales? We are not told. There are other problems. Mirror is only part of a group of genes in flies and in flies both iroquois and mirror are needed to make the back of the wing, the alula (Kehl et al). What is known about iro expression in butterflies?
In flies, mirror regulates a late and local expression of dpp that seems to be responsible for making the alula. What happens in butterflies? Would a study of the expression of Dpp in wildtype and mirror compromised wings be useful?
Thus, I find the paper to be disappointing for a general journal as it does little more than claim what was discovered in Drosophila is at least partly true in butterflies. Also, it fails to explain what the authors mean by "wing domains" and "domain specification". They are not alone, butterfly workers, in general, appear vague about these concepts, their vagueness allowing too much loose thinking.
Since these matters are at the heart of the purpose and meaning of the work reported here, we readers need a paper containing more critical thought and information. I would like to have a better and more logical introduction and discussion.
The authors do define what they mean by the vannus of the wing. In flies the definition of compartments is clear and abundantly demonstrated, with gene expression and requirement being limited precisely to sets of cells that display lineage boundaries. It is true that domains of gene expression in flies, for example of the iroquois complex, which includes mirror, can only be related to patterns with difficulty. Some recap of what is known plus the opinion of the authors on how they interpret papers on possible lineage domains in butterflies might also be useful as the reader, is no wiser about what the authors might mean at the end of it!
The references are sometimes inappropriate. The discovery of the AP compartments should not be referred to Guillen et al 1995, but to Morata and Lawrence 1975. Proofreading is required.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This paper reports investigations of chromosome stiffness in oocytes and spermatocytes. The paper shows that prophase I spermatocytes and MI/MII oocytes yield high Young Modulus values in the assay the authors applied. Deficiency in each one of three meiosis-specific cohesins they claim did not affect this result and increased stiffness was seen in aged oocytes but not in oocytes treated with the DNA-damaging agent etoposide.
The paper reports some interesting observations which are in line with a report by the same authors of 2020 where increased stiffness of spermatocyte chromosomes was already shown. In that sense, the current manuscript is an extension of that previous paper, and thus novelty is somewhat limited. The paper is also largely descriptive as it does neither propose a mechanism nor report factors that determine the chromosomal stiffness.
There are several points that need to be considered.
(1) Limitations of the study and the conclusions are not discussed in the "Discussion" section and that is a significant gap. Even more so as the authors rely on just one experimental system for all their data - there is no independent verification - and that in vitro system may be prone to artefacts.
(2) It is somewhat unfortunate that they jump between oocytes and spermatocytes to address the cohesin question. Prophase I (pachytene) spermatocytes chromosomes are not directly comparable to MI or MII oocyte chromosomes. In fact, the authors report Young Modulus values of 3700 for MI oocytes and only 2700 for spermatocyte prophase chromosomes, illustrating this difference. Why not use oocyte-specific cohesin deficiencies?
(3) It remains unclear whether the treatment of oocytes with the detergent TritonX-100 affects the spindle and thus the chromosomes isolated directly from the Triton-lysed oocytes. In fact, it is rather likely that the detergent affects chromatin-associated proteins and thus structural features of the chromosomes.
(4) Why did the authors use mouse strains of different genetic backgrounds, CD-1, and C57BL/6? That makes comparison difficult. Breeding of heterozygous cohesin mutants will yield the ideal controls, i.e. littermates.
(5) How did the authors capture chromosome axes from STAG3-deficienct spermatocytes which feature very few if any axes? How representative are those chromosomes that could be captured?
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:<br /> The paper explores a mathematical model of subsecond time perception, engaging with established theories such as the linear psychophysical law, Weber's law, and dopaminergic modulation of subjective durations. While it ambitiously attempts to confirm specific mechanisms of time perception and presents a comprehensive description of these mechanisms, the work is presented as data-driven but its empirical backing and model generalization capabilities are questionable. The title's implication of a robust empirical foundation is misleading, as the main figures do not reflect empirical data directly but rather model outputs aligned with general trends in psychophysical studies. This disjunction raises concerns about the model's applicability and the strength of the claims made regarding time perception mechanisms.
Strengths:<br /> (1) The paper describes specific mechanisms of time perception, providing a theoretical examination of linear psychophysical law, Weber's law, and dopaminergic modulation. This aspect is valuable for readers seeking a theoretical understanding of temporal perception.
(2) The authors describe a range of psychophysical studies and theories, attempting to position their model within the broader scientific discourse on time perception.
Weaknesses:<br /> (1) Lack of Empirical Data: The absence of two things: 1) quantification of error between model and empirical data with interpretation of what this degree of error means, and 2) clear comparisons between model and empirical data in all figures and tables, to substantiate the model's predictions stands out. The reliance on general trends rather than specific empirical studies undermines the strength and reliability of the model's claims. The paper would benefit from quantitative and qualitative simulations of results from specific, large-sample studies to anchor the model's predictions in concrete empirical evidence.
(2) Methodological Ambiguities: The training and testing procedures lack robust checks for generalization, leading to potential overfitting issues. Clarifications are needed on whether and how the model reaches a steady state before stimulation and the implications of the chosen model time constants in the absence of stimulation. The overlap between training (50ms) and testing (25ms) steps and the implications for model generalization need validation with "traditional" parameter fitting protocols, such as formal model cross-validation across well-defined datasets and splits, as well as evaluations to understand and assess potential overfitting.
(3) Inadequate Visualization of Empirical Data: References to empirical data are vague and not directly visualized alongside model outputs. Future iterations should include empirical data, not general trends from psychophysics, in figures for a clear comparison.
(4) Limitations in Model Scope and Dynamics: The exploration of limitations is narrowly focused on interval length and noise. Expanding the model limitations to consider isochronous pulse processing and the emergence of limit-cycle behaviors after prolonged stimulation would provide a more comprehensive understanding of the model's capabilities and limitations. Additionally, the justification for using \(N_{Poisson}\) as a proxy for more connections is unclear and warrants a more direct approach. Adding more units to a truly data-driven model should be trivial.
(5) Omissions and Redundancies: Certain omissions, such as the lack of a condition in Figure 7A or missing references to relevant models and reviews, detract from the paper's thoroughness. Moreover, some statements and terms like "internal clock" are used without a clear mechanistic definition within the model.
Guidance for Readers<br /> Readers should approach this paper as a theoretical exploration into the mechanisms of subsecond-time perception. The model offers a detailed theoretical framework that engages with established laws and theories in time perception. However, it's crucial to note the model's reliance on general trends and its lack of direct empirical backing. The findings should be interpreted as a hypothesis-generating exercise rather than conclusive evidence.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Summary:
The authors are trying to find a vaccine solution for invasive candidiasis.
Strengths:
The testing of the antifungal activity of EDTA on Candida is not new as many other papers have examined this effect. The novelty here is on the use of this such EDTA treated strain as a vaccine to protect against a secondary challenge with wild-type Candida.
Weaknesses:
However, data presented in Fig. 5 and in Fig. 6 are not convincing and need further experimental controls and analysis as the authors do not show a time-dependent effect on the CFU of their vaccine formulation. Specific points are below.
Methodology used is also an issue. As it stands, the impact is minor, if any.
Comments on revised version:
The data provided in the revised paper are simply not satisfactory and do not give confidence that a rigorous design and methodologies were used to obtain the results illustrated in this paper.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Microbial degradation of synthetic organic compounds is the basis of bioremediation. Biodegradation of 1NA has not been previously reported. The report describes a complete study of 1NA biodegradation by a new isolate Pseudomonas sp. strain JS3066. The study includes the enrichment and isolation of the 1NA-degrading bacterium Pseudomonas sp. strain JS3066, the identification of the genes and enzymes involved in 1NA degradation, and the detailed characterization of γ-glutamylorganoamide synthetase by using biochemical and structural analysis. In the discussion, the potential evolution of 1NA degradation pathway, the similarity and difference between γ-glutamylorganoamide synthetase and glutamine synthetase, and the significance were explained. The conclusions were well supported by the results presented.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The paper by Gilbert et al. is well-written in a detailed format and the authors are candid in their data interpretation by acknowledging that the described ninein bone defects are mild, transient, and do not lead to major long-lasting defects in adulthood.
The main strength of the study is presenting a novel link between a centrosomal protein and osteoclasts in the mouse. However, the majority of the work is dedicated to describing the premature ossification phenotype and less attention is paid to how a centrosomal protein affects osteoclast proliferation, survival, and/or differentiation into mature osteoclasts.
Based on the decrease in the number of osteoclasts (Fig 5E, G, and also per coverslip after 2 days in culture), the authors suggest that the loss of ninein impacts osteoclast proliferation. First, proliferation can be directly quantified using Ki67 staining or EdU incorporation. Second, other interpretations are also plausible and can also be experimentally tested. These include less adhesion and attachment of the mutants to the coverslips, but perhaps more relevant in vivo is cell death of the ninein mutant osteoclasts. It has been established that the loss of centrosome function activates p53-dependent cell death and osteoclasts might be a vulnerable cell population. Quantifying p53 immunoreactivity and/or cell death in osteoclasts might help clarify the phenotype of osteoclast reduction.
-
-
martin.kleppmann.com martin.kleppmann.com
-
When updates are generatedconcurrently and then merged, the next update containsmultiple predecessors
Given many updates are merged at the same time pointers are = many. But given we merge right away having learned an update - we always merge two updates - hence two pointers.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The research uses a large collection of Arabidopsis thaliana accessions from various geographic scales to investigate the natural genetic variation underlying the response of ionome (elemental) composition to elevated CO2 (eCO2), a concern for future food security. While most accessions show a decrease in elemental accumulation, the authors demonstrate a wide variety of responses to eCO2 across the diversity of Arabidopsis, including lines that increase elemental content in eCO2. The demonstration of genetic diversity in eCO2 response is a significant contribution to our understanding of this important phenomenon.
Comments on revised version:
The authors made significant improvements in the manuscript from the original preprint, and the conclusions are now well supported by the evidence presented.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Summary:
Zai et al. test whether birds can modify their vocal behavior in a manner consistent with planning. They point out that while some animals are known to be capable of volitional control of vocalizations, it has been unclear if animals are capable of planning vocalizations-that is, modifying vocalizations towards a desired target without the need to learn this modification by practising and comparing sensory feedback of practised behavior to the behavioral target. They study zebra finches that have been trained to shift the pitch of song syllables away from their baseline values. It is known that once this training ends, zebra finches have a drive to modify pitch so that it is restored back to its baseline value. They take advantage of this drive to ask whether birds can implement this targeted pitch modification in a manner that looks like planning, by comparing the time course and magnitude of pitch modification in separate groups of birds who have undergone different manipulations of sensory and motor capabilities. A key finding is that birds who are deafened immediately before the onset of this pitch restoration paradigm, but after they have been shifted away from baseline, are able to shift pitch partially back towards their baseline target. In other words, this targeted pitch shift occurs even when birds don't have access to auditory feedback, which argues that this shift is not due to reinforcement-learning-guided practice, but is instead planned based on the difference between an internal representation of the target (baseline pitch) and current behavior (pitch the bird was singing immediately before deafening).
The authors present additional behavioral studies arguing that this pitch shift requires auditory experience of song in its state after it has been shifted away from baseline (birds deafened early on, before the initial pitch shift away from baseline, do not exhibit any shift back towards baseline), and that a full shift back to baseline requires auditory feedback. The authors synthesize these results to argue that different mechanisms operate for small shifts (planning, which does not need auditory feedback) and large shifts (through a mechanism that requires auditory feedback).
The authors also make a distinction between two kinds of planning: covert-not requiring any motor practice and overt-requiring motor practice but without access to auditory experience from which target mismatch could be computed. They argue that birds plan overtly, based on these deafening experiments as well as an analogous experiment involving temporary muting, which suggests that indeed motor practice is required for pitch shifts.
Strengths:
The primary finding (that partially restorative pitch shift occurs even after deafening) rests on strong behavioral evidence. It is less clear to what extent this shift requires practice, since their analysis of pitch after deafening takes the average over within the first two hours of singing. If this shift is already evident in the first few renditions then this would be evidence for covert planning. Technical hurdles, such as limited sample sizes and unstable song after surgical deafening, make this difficult to test. (Similarly, the authors could test whether the first few renditions after recovery from muting already exhibit a shift back towards baseline.)
This work will be a valuable addition to others studying birdsong learning and its neural mechanisms. It documents features of birdsong plasticity that are unexpected in standard models of birdsong learning based on reinforcement and are consistent with an additional, perhaps more cognitive, mechanism involving planning. As the authors point out, perhaps this framework offers a reinterpretation of the neural mechanisms underlying a prior finding of covert pitch learning in songbirds (Charlesworth et al., 2012).
A strength of this work is the variety and detail in its behavioral studies, combined with sensory and motor manipulations, which on their own form a rich set of observations that are useful behavioral constraints on future studies.
Weaknesses:
The argument that pitch modification in deafened birds requires some experience hearing their song in its shifted state prior to deafening (Fig. 4) is solid but has an important caveat. Their argument rests on comparing two experimental conditions: one with and one without auditory experience of shifted pitch. However, these conditions also differ in the pitch training paradigm: the "with experience" condition was performed using white noise training, while the "without experience" condition used "lights off" training (Fig. 4A). It is possible that the differences in ability for these two groups to restore pitch to baseline reflects the training paradigm, not whether subjects had auditory experience of the pitch shift. Ideally, a control study would use one of the training paradigms for both conditions, which would be "lights off" or electrical stimulation (McGregor et al. 2022), since WN training cannot be performed in deafened birds. In the Discussion, in response to this point, the authors point out that birds are known to recover their pitch shift if those shifts are driven using electrical stimulation as reinforcement (McGregor et al. 2022); however, it is arguably still relevant to know whether a similar recovery occurs for the "lights off" paradigm used here.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Several publications during the past years provided evidence that NMD protects tumor cells from being recognized by the immune system by suppressing the display of neoantigens, and hence NMD inhibition is emerging as a promising anti-cancer approach. However, the lack of an efficacious and specific small-molecule NMD inhibitor with suitable pharmacological properties is currently a major bottleneck in the development of therapies that rely on NMD inhibition. In this manuscript, the authors describe their screen for identifying NMD inhibitors, which is based on isogenic cell lines that either express wild-type or NMD-sensitive transcript isoforms of p53 and STAG2. Using this setup, they screened a library of 2658 FDA-approved or late-phase clinical trial drugs and had 8 hits. Among them they further characterized LY3023414, showing that it inhibits NMD in cultured cells and in a mouse xenograft model, where it, however, was very toxic. Because LY3023414 was originally developed as a PI3K inhibitor, the authors claim that it inhibits NMD by inhibiting SMG1. While this is most likely true, the authors do not provide experimental evidence for this claim. Instead, they use this statement to switch their attention to another previously developed SMG1 inhibitor (SMG1i-11), of which they design and test several derivatives. Of these derivatives, KVS0001 showed the best pharmacological behavior. It upregulated NMD-sensitive transcripts in cultured cells and the xenograft mouse model and two predicted neoantigens could indeed be detected by mass spectrometry when the respective cells were treated with KVS0001. A bispecific antibody targeting T cells to a specific antigen-HLA complex led to increased IFN-gamma release and killing of cancer cells expressing this antigen-HLA complex when they were treated with KVS0001. Finally, the authors show that renal (RENCA) or lung cancer cells (LLC) were significantly inhibited in tumor growth in immunocompetent mice treated with KVS0001. Overall, this establishes KVS0001 as a novel and promising ant-cancer drug that by inhibiting SMG1 (and therewith NMD) increases the neoantigen production in the cancer cells and reveals them to the body's immune system as "foreign".
Strengths:
The novelty and significance of this work consists in the development of a novel and - judging from the presented data - very promising NMD inhibiting drug that is suitable for applications in animals. This is an important advance for the field, as previous NMD inhibitors were not specific, lacked efficacy, or were very toxic and hence not suitable for animal application. It will be still a long way with many challenges ahead towards an efficacious NMD inhibitor that is safe for use in humans, but KVS0001 appears to be a molecule that bears promise for follow-up studies. In addition, while the idea of inhibiting NMD to trigger neoantigen production in cancer cells and so reveal them to the immune system has been around for quite some time, this work provides ample and compelling support for the feasibility of this approach, at least for tumors with a high mutational burden.
Main weaknesses:
There is a disconnect between the screen and the KVS0001 compound, that they describe and test in the second part of the manuscript since KVS0001 is a derivative of the SMG1 inhibitors developed by Gopalsamy et al. in 2012 and not of the lead compound identified in the screen (LY3023414). Because of high toxicity in the mouse xenograft experiments, the authors did not follow up LY3023414 but instead switched to the published SMG1i-11 drug of Gopalsamy and colleagues, a molecule that is widely used among NMD researchers for NMD inhibition in cultured cells. Therefore, in my view, the description of the screen is obsolete, and the paper could just start with the optimization of the pharmacological properties of SMG1i-11 and the characterization of KVS0001. Even though the screen is based on an elegant setup and was executed successfully, it was ultimately a failure as it didn't reveal a useful lead compound that could be further optimized.
Additional points:
- Compared to SMG1i-11, KVS0001 seems less potent in inhibiting SMG1 (higher IC50). It would therefore be important to also compare the specificity of both drugs for SMG1 over other kinases at the applied concentrations (1 uM for SMG1i-11, 5 uM for KVS0001). The Kinativ Assay (Fig. S13) was performed with 100 nM KVS0001, which is 50-fold less than the concentration used for functional assays and hence not really meaningful. In addition, more information on the pharmacokinetic properties and toxicology of KVS0001 would allow a better judgment of the potential of this molecule as a future therapeutic agent.
- On many figures, the concentrations of the used drugs are missing. Please ensure that for every experiment that includes drugs, the drug concentration is indicated.
- Do the authors have an explanation for why LY3023414 has a much stronger effect on the p53 than on the STAG2 nonsense allele (Figure 1B, S8), whereas emetine upregulates the STAG2 nonsense alleles more than the p53 nonsense allele (Figure S5). I find this curious, but the authors do not comment on it.
- While it is a strength of the study that the NMD inhibitors were validated on many different truncation mutations in different cell lines, it would help readers if a table or graphic illustration was included that gives an overview of all mutant alleles tested in this study (which gene, type of mutation, in which cell type). In the current version, this information is scattered throughout the manuscript.
- Lines 194 and 302: That SMG1i-11 was highly insoluble in the hands of the authors is surprising. It is unclear why they used variant 11j, since variant 11e of this inhibitor is widely used among NMD researchers and readily dissolves in DMSO.
- Line 296: The authors claim that they were able to show that LY3023414 inhibited the SMG1 kinase, which is not true. To show this, they would have for example to show that LY3023414 prevents SMG1-mediated UPF1 phosphorylation, as they did for KVS0001 and SMG1i-11 in Fig. 3F. Unless the authors provide this data, the statement should be deleted or modified.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
This study seeks to investigate one aspect of disparity in academia: how gender balance in a discipline is valued in terms of evaluated research quality score and funding success. This is important in understanding disparities within academia.<br /> This study uses publicly available data to investigate covariation between gender balance in an academic discipline and:<br /> i) Individual research quality scores of New Zealand academics as evaluated by one of 14 broader subject panels.<br /> ii) Funding success in Australia, Canada, Europe, UK.
The study would benefit from further discussion of it limitations, and from the clarification of some technical points (as described in the recommendations for the authors).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary
Recent evidence indicates that cells of the navigation system representing different directions and whole spatial routes fire in a rhythmic alternation during 5-10 Hz (theta) network oscillation (Brandon et al., 2013, Kay et al., 2020). This phenomenon of theta cycle skipping was also reported in broader circuitry connecting the navigation system with the cognitive control regions (Jankowski et al., 2014, Tang et al., 2021). Yet nothing was known about the translation of these temporally separate representations to midbrain regions involved in reward processing as well as the hypothalamic regions, which integrate metabolic, visceral, and sensory signals with the descending signals from the forebrain to ensure adaptive control of innate behaviors (Carus-Cadavieco et al., 2017). The present work aimed to investigate theta cycle skipping and alternating representations of trajectories in the lateral septum, neurons of which receive inputs from large number of CA1 and nearly all CA3 pyramidal cells (Risold and Swanson, 1995). While spatial firing has been reported in the lateral septum before (Leutgeb and Mizumori, 2002, Wirtshafter and Wilson, 2019), its dynamic aspects have remained elusive. The present study replicates the previous findings of theta-rhythmic neuronal activity in the lateral septum and reports a temporal alternation of spatial representations in this region, thus filling an important knowledge gap and significantly extending the understanding of the processing of spatial information in the brain. The lateral septum thus propagates the representations of alternative spatial behaviors to its efferent regions. The results can instruct further research of neural mechanisms supporting learning during goal-oriented navigation and decision-making in the behaviourally crucial circuits entailing the lateral septum.
Strengths
To this end, cutting-edge approaches for high-density monitoring of neuronal activity in freely behaving rodents and neural decoding were applied. Strengths of this work include comparisons of different anatomically and probably functionally distinct compartments of the lateral septum, innervated by different hippocampal domains and projecting to different parts of the hypothalamus; large neuronal datasets including many sessions with simultaneously recorded neurons; consequently, the rhythmic aspects of the spatial code could be directly revealed from the analysis of multiple spike trains, which were also used for decoding of spatial trajectories; and comparisons of the spatial coding between the two differently reinforced tasks.
Weaknesses
Without using perturbation techniques, the present approach could not identify the aspects of the spatial code actually influencing the generation of behaviors by downstream regions.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript builds on previous work suggesting that the CCK peptide is the releasing hormone for FSH in fishes, which is different than that observed in mammals where both LH and FSH release are under the control of GnRH. Based on data using calcium imaging as a readout for stimulation of the gonadotrophs, the researchers present data supporting the hypothesis that CCK stimulates FSH-containing cells in the pituitary. In contrast, LH-containing cells show a weak and variable response to CCK but are highly responsive to GnRH. Data are presented that support the role of CCK in the release of FSH. Researchers also state that functional overlap exists in the potency of GnRH to activate FSH cells, thus the two signalling pathways are not separate.
The results are of interest to the field because for many years the assumption has been that fishes use the same signalling mechanism. These data present an intriguing variation where a hormone involved in satiation acts in the control of reproduction.
Strengths:
The strengths of the manuscript are that researchers have shed light on different pathways controlling reproduction in fishes.
Weaknesses:
Weaknesses are that it is not clear if multiple ligand/receptors are involved (more than one CCK and more than one receptor?). The imaging of the CCK terminals and CCK receptors needs to be reinforced.
Reviewer consultation summary:
- The data presented establish sufficiency, but not necessity of CCK in FSH regulation. The paper did not show that CCK endogenously regulates FSH in fish. This has not been established yet.
- The paper presents the pharmacological effects of CCK on ex vivo preparations but does not establish the in vivo physiological function of the peptide. The current evidence for a novel physiological regulatory mechanism is incomplete and would require further physiological experiments. These could include the use of a CCK receptor antagonist in adult fish to see the effects on FSH and LH release, the generation of a CCK knockout, or cell-specific genetic manipulations.
- Zebrafish have two CCK ligands: ccka, cckb and also multiple receptors: cckar, cckbra and cckbrb. There is ambiguity about which CCK receptor and ligand are expressed and which gene was knocked out.
- Blocking CCK action in fish (with receptor KO) affects FSH and LH. Therefore, the work did not demonstrate a selective role for CCK in FSH regulation in vivo and any claims to have discovered FSHRH need to be more conservative.
- The labelling of the terminals with anti-CCK looks a lot like the background and the authors did not show a specificity control (e.g. anti-CCK antibody pre-absorbed with the peptide or anti-CCK in morphant/KO animals).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The physiology and behaviour of animals are regulated by a huge variety of neuropeptide signalling systems. In this paper, the authors focus on the neuropeptide ion transport peptide (ITP), which was first identified and named on account of its effects on the locust hindgut (Audsley et al. 1992). Using Drosophila as an experimental model, the authors have mapped the expression of three different isoforms of ITP (Figures 1, S1, and S2), all of which are encoded by the same gene.
The authors then investigated candidate receptors for isoforms of ITP. Firstly, Drosophila orthologs of G-protein coupled receptors (GPCRs) that have been reported to act as receptors for ITPa or ITPL in the insect Bombyx mori were investigated. Importantly, the authors report that ITPa does not act as a ligand for the GPCRs TkR99D and PK2-R1 (Figure S3). Therefore, the authors investigated other putative receptors for ITPs. Informed by a previously reported finding that ITP-type peptides cause an increase in cGMP levels in cells/tissues (Dircksen, 2009, Nagai et al., 2014), the authors investigated guanylyl cyclases as candidate receptors for ITPs. In particular, the authors suggest that Gyc76C may act as an ITP receptor in Drosophila.
Evidence that Gyc76C may be involved in mediating effects of ITP in Bombyx was first reported by Nagai et al. (2014) and here the authors present further evidence, based on a proposed concordance in the phylogenetic distribution ITP-type neuropeptides and Gyc76C (Figure 2). Having performed detailed mapping of the expression of Gyc76C in Drosophila (Figures 3, S4, S5, S6), the authors then investigated if Gyc76C knockdown affects the bioactivity of ITPa in Drosophila. The inhibitory effect of ITPa on leucokinin- and diuretic hormone-31-stimulated fluid secretion from Malpighian tubules was found to be abolished when expression of Gyc76C was knocked down in stellate cells and principal cells, respectively (Figure 4). However, as discussed below, this does not provide proof that Gyc76C directly mediates the effect of ITPa by acting as its receptor. The effect of Gyc76C knockdown on the action of ITPa could be an indirect consequence of an alteration in cGMP signalling.
Having investigated the proposed mechanism of ITPa in Drosophila, the authors then investigated its physiological roles at a systemic level. In Figure 5 the authors present evidence that ITPa is released during desiccation and accordingly, overexpression of ITPa increases survival when animals are subjected to desiccation. Furthermore, knockdown of Gyc76C in stellate or principal cells of Malphigian tubules decreases survival when animals are subject to desiccation. However, whilst this is correlative, it does not prove that Gyc76C mediates the effects of ITPa. The authors investigated the effects of knockdown of Gyc76C in stellate or principal cells of Malphigian tubules on i). survival when animals are subject to salt stress and ii). time taken to recover from of chill coma. It is not clear, however, why animals over-expressing ITPa were also not tested for its effect on i). survival when animals are subject to salt stress and ii). time taken to recover from of chill coma. In Figures 6 and S8, the authors show the effects of Gyc76C knockdown in the female fat body on metabolism, feeding-associated behaviours and locomotor activity, which are interesting. Furthermore, the relevance of the phenotypes observed to potential in vivo actions of ITPa is explored in Figure 7. The authors conclude that "increased ITPa signaling results in phenotypes that largely mirror those seen following Gyc76C knockdown in the fat body, providing further support that ITPa mediates its effects via Gyc76C." Use of the term "largely mirror" seems inappropriate here because there are opposing effects- e.g. decreased starvation resistance in Figure 6A versus increased starvation resistance in Figure 7A. Furthermore, as discussed above, the results of these experiments do not prove that the effects of ITPa are mediated by Gyc76C because the effects reported here could be correlative, rather than causative.
Lastly, in Figures 8, S9, and S10 the authors analyse publicly available connectomic data and single-cell transcriptomic data to identify putative inputs and outputs of ITPa-expressing neurons. These data are a valuable addition to our knowledge ITPa expressing neurons; but they do not address the core hypothesis of this paper - namely that Gyc76C acts as an ITPa receptor.
Strengths:
(1) The main strengths of this paper are i) the detailed analysis of the expression and actions of ITP and the phenotypic consequences of over-expression of ITPa in Drosophila. ii). the detailed analysis of the expression of Gyc76C and the phenotypic consequences of knockdown of Gyc76C expression in Drosophila.
(2) Furthermore, the paper is generally well-written and the figures are of good quality.
Weaknesses:
(1) The main weakness of this paper is that the data obtained do not prove that Gyc76C acts as a receptor for ITPa. Therefore, the following statement in the abstract is premature: "Using a phylogenetic-driven approach and the ex vivo secretion assay, we identified and functionally characterized Gyc76C, a membrane guanylate cyclase, as an elusive Drosophila ITPa receptor." Further experimental studies are needed to determine if Gyc76C acts as a receptor for ITPa. In the section of the paper headed "Limitations of the study", the authors recognise this weakness. They state "While our phylogenetic analysis, anatomical mapping, and ex vivo and in vivo functional studies all indicate that Gyc76C functions as an ITPa receptor in Drosophila, we were unable to verify that ITPa directly binds to Gyc76C. This was largely due to the lack of a robust and sensitive reporter system to monitor mGC activation." It is not clear what the authors mean by "the lack of a robust and sensitive reporter system to monitor mGC activation". The discovery of mGCs as receptors for ANP in mammals was dependent on the use of assays that measure GC activity in cells (e.g. by measuring cGMP levels in cells). Furthermore, more recently cGMP reporters have been developed. The use of such assays is needed here to investigate directly whether Gyc76C acts as a receptor for ITPa. In summary, insufficient evidence has been obtained to conclude that Gyc76C acts as a receptor for ITPa. Therefore, I think there are two ways forward, either:<br /> (a) The authors obtain additional biochemical evidence that ITPa is a ligand for Gyc76C.<br /> or<br /> (b) The authors substantially revise the conclusions of the paper (in the title, abstract, and throughout the paper) to state that Gyc76C MAY act as a receptor for ITPa, but that additional experiments are needed to prove this.
(2) The authors state in the abstract that a phylogenetic-driven approach led to their identification of Gyc76C as a candidate receptor for ITPa. However, there are weaknesses in this claim. Firstly, because the hypothesis that Gyc76C may be involved in mediating effects of ITPa was first proposed ten years ago by Nagai et al. 2014, so this surely was the primary basis for investigating this protein. Nevertheless, investigating if there is correspondence in the phylogenetic distribution of ITP-type and Gyc76C-type genes/proteins is a valuable approach to addressing this issue. Unfortunately, the evidence presented is rather limited in scope. Essentially, the authors report that they only found ITP-type and Gyc76C-type genes/proteins in protostomes, but not in deuterostomes. What is needed is a more fine-grained analysis at the species level within the protostomes. Thus, are there protostome species in which both ITP-type and Gyc76C-type genes/proteins have been lost? Furthermore, are there any protostome species in which an ITP-type gene is present but an Gyc76C-type gene is absent, or vice versa? If there are protostome species in which an ITP-type gene is present but a Gyc76C-type gene is absent or vice versa, this would argue against Gyc76C being a receptor for ITPa. In this regard, it is noteworthy that in Figure 2A there are two ITP-type precursors in C. elegans, but there are no Gyc76C-type proteins shown in the tree in Figure 2B. Thus, what is needed is a more detailed analysis of protostomes to investigate if there really is correspondence in the phylogenetic distribution of Gyc76C-type and ITP-type genes at the species level.
(3) The manuscript would benefit from a more comprehensive overview and discussion of published literature on Gyc76C in Drosophila, both as a basis for this study and for interpretation of the findings of this study.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Latham A.P. et al. apply simulations and FLIM to analyse several di-block elastin-like polypetides and connect their sequence to the micro-structure of coacervates resulting from their phase-separation.
Strengths:
Understanding the molecular grammar of phase separating proteins and the connection with mesoscale properties of the coacervates is highly relevant. This work provides insights into micro-structures of coacervates resulting from di-block polypetides.
Weaknesses:
The results apply to a very specific architecture (di-block polypetides) with specific sequences.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Thakare et al. present the DIETS assay for quantifying food consumption in adult Drosophila. DIETS measures food intake by weighing fly food before and after feeding. Technically, this is a well-designed, executed, and analyzed study. The interpretations are generally conservative and justified by the results. Although the results aren't always consistent with other published studies, which might reflect some of the unique conditions of the DIETS assay, the technique can clearly distinguish between some expected differences in food intake. Although lifespan is shortened in the DIETS chamber, the method seems robust for various time scales up to a week. DIETS adds another useful and versatile tool for fly researchers interested in studying feeding behavior.
Strengths:
The authors test various conditions, including food presentation, surface area, and humidity (by changing the food cup distance to an agar base) to demonstrate an optimized technique for quantifying consumption. Under these conditions, evaporation is generally limited to <10%.
The authors use DIETS to validate diverse feeding paradigms, including the published effects of temperature, food dilution, and intermittent fasting on food intake.
Weaknesses:
The studies to optimize and test the DIETS assay are technically rigorous and well-designed. However, the results reveal some weaknesses or potential caveats of the assay. As highlighted below, how much nutrition flies are actually obtaining may be misestimated due to vapor diffusion, and crowding/competition for food. This appears largely acceptable though, since the 'group' measurement can still clearly distinguish between expected feeding differences under different conditions, but it likely reduces accuracy, which may be important in some studies, and probably nullifies the effectiveness of using DIETS to restrict caloric intake.
It is my understanding that flies suck out nutrients from the medium, leaving behind the agar/cornmeal matrix. This seems consistent with the images in Figure S2B, where the spheroidal medium in the food cup maintains its shape as it shrinks, but there don't seem to be any pits or holes from fly consumption. Given that flies in DIETS consume a significant portion of the available food, it seems that the food concentration at the medium surface may be changing throughout the experiment. This may also make it challenging to use other common fly food ingredients, such as cornmeal, much of which is indigestible.
Similarly, vapor diffusion is expected between the agar bed and food cup (which the authors observed; in line 385), which may further affect assay accuracy, especially in comparisons between foods with different osmolarity.
In DIETS, increased feeding is observed with increased flies per chamber, but this is not observed in other techniques, such as EX-Q (Wu et al. 2020). It is unclear whether sensitivity to adult density is a DIETS-specific feature, or if adult density instead directly affects food intake estimates using DIETS (e.g., by affecting chamber humidity).
In another example, there is a ~300% difference in absolute feeding when the DIETS food cup is presented in different formats (Figure 3C). Again, it is unclear whether food presentation has an inherently greater effect in DIETS, or if the measurements themselves are highly sensitive to the environment.
Although the control of total food mass given to the animals is a novel feature of the assay, the likely differences in nutrient intake between individuals (and shortened lifespan) in a DIETS chamber makes this a challenging method to use to study caloric restriction. The shortened lifespan likely stems from the high adult density per vial, which has been explored in previous publications (e.g., Pearl in the 1920s; Mueller in the 1990s).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This manuscript explores mechanisms underlying heart contractility problems in metabolic disease using Drosophila as a model. They confirm, as others have demonstrated, that a high-fat diet (HFD) induces cardiac problems in flies. They showed that a high-fat diet increased Akh mRNA levels and calcium levels in the Akh-producing cells (APC), suggesting there is increased production and release of this hormone in a HFD context. When they knock down Akh production in the APCs using RNAi they see that cardiac contractility problems are abolished. They similarly show that levels of the Akh receptor (Akhr) are increased on a HFD and that loss of Akhr also rescues contractility problems on a HFD.
One highlight of the paper was the identification of a pair of neurons that express a receptor for the metabolic hormone Akh, and showing initial data that these neurons innervate the cardiac muscle. They then overexpress cell death gene reaper (rpr) in all Akhr-positive cells with Akhr-GAL4 and see that cardiac contractility becomes abnormal.
However, this paper contains several findings that have been reported elsewhere and it contains key flaws in both experimental design and data interpretation. There is some rationale for doing the experiments, and the data and images are of good quality. However, others have shown that HFD induces cardiac contractility problems (Birse 2010), that Akh mRNA levels are changed with HFD (Liao 2021) that Akh modulates cardiac rhythms (Noyes 1995), so Figures 1-4 are largely a confirmation of what is already known. This limits the overall magnitude of the advances presented in these figures. Overall, the stated concerns limit the impact of the manuscript in advancing our understanding of heart contractility.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This is an interesting method that addresses the important problem of assessing G protein localization at endogenous levels. The data are generally convincing.
Specific comments
Methods:<br /> The description of the gene editing method is unclear. There are two different CRISPR cell lines made in two different cell backgrounds. The methods should clearly state which CRISPR guides were used on which cell line. It is also not clear why HiBit is included in the mNG-β1 construct. Presumably, this is not critical but it would be helpful to explicitly note. In general, the Methods could be more complete.
Results:<br /> The explanation of validation experiments in Figures 1 C and D is incomplete and difficult to follow. The rationale and explanation of the experiments could be expanded. In addition, because this is an interesting method, it would be helpful to know if the endogenous editing affects normal GPCR signaling. For example, the authors could include data showing an Iso-induced cAMP response. This is not critical to the present interpretation but is relevant as a general point regarding the method. Also, it may be relevant to the interpretation of receptor effects on G protein localization.
Discussion:<br /> The conclusion that beta-gamma subunits do not redistribute after GPCR activation seems new and different from previous reports. Is this correct? Can the authors elaborate on how the results compare to previous literature?
Can the authors note that OpenCell has endogenously tagged Gβ1 and reports more obvious internal localization? Can the authors comment on this point?
Is this the first use of CRISPR / HiBit for BRET assay? It would be helpful to know this or cite previous work if not. Also, as this is submitted as a tools piece, the authors might say a little more about the potential application to other questions.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Kisspeptin neurons of the arcuate nucleus (ARC) are thought to be responsible for the pulsatile GnRH secretory pattern and to mediate feedback regulation of GnRH secretion by estradiol (E2). Evidence in the literature, including the work of the authors, indicates that ARC kisspeptin coordinate their activity through reciprocal synaptic interactions and the release of glutamate and of neuropeptide neurokinin B (NKB), which they co-express. The authors show here that E2 regulates the expression of genes encoding different voltage-dependent calcium channels, calcium-dependent potassium channels, and canonical transient receptor potential (TRPC5) channels and of the corresponding ionic currents in ARC kisspeptin neurons. Using computer simulations of the electrical activity of ARC kisspeptin neurons, the authors also provide evidence of what these changes translate into in terms of these cells' firing patterns. The experiments reveal that E2 upregulates various voltage-gated calcium currents as well as 2 subtypes of calcium-dependent potassium currents while decreasing TRPC5 expression (an ion channel downstream of NKB receptor activation), the slow excitatory synaptic potentials (slow EPSP) elicited in ARC kisspeptin neurons by NKB release and expression of the G protein-associated inward-rectifying potassium channel (GIRK). Based on these results, and on those of computer simulations, the authors propose that E2 promotes a functional transition of ARC kisspeptin neurons from neuropeptide-mediated sustained firing that supports coordinated activity for pulsatile GnRH secretion to a less intense firing in glutamatergic burst-like firing pattern that could favor glutamate release from ARC kisspeptin. The authors suggest that the latter might be important for the generation of the preovulatory surge in females.
Strengths:
The authors combined multiple approaches in vitro and in silico to gain insights into the impact of E2 on the electrical activity of ARC kisspeptin neurons. These include patch-clamp electrophysiology combined with selective optogenetic stimulation of ARC kisspeptin neurons, reverse transcriptase quantitative PCR, pharmacology, and CRIPR-Cas9-mediated knockdown of the Trpc5 gene. The addition of computer simulations for understanding the impact of E2 on the electrical activity of ARC kisspeptin cells is also a strength.
The authors add interesting information on the complement of ionic currents in ARC kisspeptin neurons and on their regulation by E2 to what was already known in the literature. Pharmacological and electrophysiological experiments appear of the highest standards. Robust statistical analyses are provided throughout, although some experiments (illustrated in Figures 7 and 8) do have rather low sample numbers.
The impact of E2 on calcium and potassium currents is compelling. Likewise, the results of Trpc5 gene knockdown do provide good evidence that the TRPC5 channel plays a key role in mediating the NKB-mediated slow EPSP. Surprisingly, this also revealed an unsuspected role for this channel in regulating the membrane potential and excitability of ARC kisspeptin neurons.
Weaknesses:
The manuscript also has weaknesses that obscure some of the conclusions drawn by the authors.
One has to do with the fact that "burst-like" firing that the authors postulate ARC kisspeptin neurons transition to after E2 replacement is only seen in computer simulations, and not in slice patch-clamp recordings. A more direct demonstration of the existence of this firing pattern, and of its prominence over neuropeptide-dependent sustained firing under conditions of high E2 would make a more convincing case for the authors' hypothesis.
In addition, and quite importantly, the authors compare here two conditions, OVX versus OVX replaced with high E2, that may not reflect the physiological conditions (the diestrous [low E2] and proestrous [high E2] stages of the estrous cycle) under which the proposed transition between neuropeptide-dependent sustained firing and less intense burst firing might take place. This is an important caveat to keep in mind when interpreting the authors' findings. Indeed, that E2 alters certain ionic currents when added back to OVX females, does not mean that the magnitude of these ionic currents will vary during the estrous cycle.
Lastly, the results of some of the pharmacological and genetic experiments may be difficult to interpret as presented. For example, in Figure 3, although it is possible that blockade of individual calcium channel subtypes suppresses the slow EPSP through decreased calcium entry at the somato-dendritic compartment to sustain TRPC5 activation and the slow depolarization (as the authors imply), a reasonable alternative interpretation would be that at least some of the effects on the amplitude of the slow EPSP result from suppression of presynaptic calcium influx and, thus, decreased neurotransmitter and neuropeptide secretion. Along the same lines, in Figure 12, one possible interpretation of the observed smaller slow EPSPs seen in mice with mutant TRPC5 could be that at least some of the effect is due to decreased neurotransmitter and neuropeptide release due to the decreased excitability associated with TRPC5 knockdown.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Wang, He et al. shed insight into the molecular mechanisms of deep-sea chemosymbiosis at the single-cell level. They do so by producing a comprehensive cell atlas of the gill of Gigantidas platifrons, a chemosymbiotic mussel that dominates the deep-sea ecosystem. They uncover novel cell types and find that the gene expression of bacteriocytes, the symbiont-hosting cells, supports two hypotheses of host-symbiont interactions: the "farming" pathway, where symbionts are directly digested, and the "milking" pathway, where nutrients released by the symbionts are used by the host. They perform an in situ transplantation experiment in the deep sea and reveal transitional changes in gene expression that support a model where starvation stress induces bacteriocytes to "farm" their symbionts, while recovery leads to the restoration of the "farming" and "milking" pathways.
A major strength of this study includes the successful application of advanced single nucleus techniques to a non-model, deep sea organism that remains challenging to sample. I also applaud the authors for performing an in situ transplantation experiment in a deep sea environment. From gene expression profiles, the authors deftly provide a rich functional description of G. platifrons cell types that is well-contextualized within the unique biology of chemosymbiosis. These findings offer significant insight into the molecular mechanisms of deep-sea host-symbiont ecology, and will serve as a valuable resource for future studies into the striking biology of G. platifrons.
The authors' conclusions are generally well-supported by their results. However, I recognize that the difficulty of obtaining deep-sea specimens may have impacted experimental design and no replicates were sampled.
It is notable that the Fanmao cells were much more sparsely sampled. It appears that fewer cells were sequenced, resulting in the Starvation and Reconstitution conditions having 2-3x more cells after doublet filtering. These discrepancies also are reflected in the proportion of cells that survived QC, suggesting a distinction in quality or approach. However, the authors provide clear and sufficient evidence via bootstrapping that batch effects between the three samples are negligible. While batch effect does not appear to have affected gene expression profiles, the proportion of cell types may remain sensitive to sampling techniques, and thus interpretation of Fig. S12 must be approached with caution.
-
-
www.researchsquare.com www.researchsquare.com
-
Reviewer #2 (Public Review):
This manuscript by Xue et al. describes the effects of a long noncoding RNA, lncDACH1, on the localization of Nav channel expression, the magnitude of INa, and arrhythmia susceptibility in the mouse heart. Because lncDACH1 was previously reported to bind and disrupt membrane expression of dystrophin, which in turn is required for proper Nav1.5 localization, much of the findings are inferred through the lens of dystrophin alterations.
The results report that cardiomyocyte-specific transgenic overexpression of lncDACH1 reduces INa in isolated cardiomyocytes; measurements in whole heart show a corresponding reduction in conduction velocity and enhanced susceptibility to arrhythmia. The effect on INa was confirmed in isolated WT mouse cardiomyocytes infected with a lncDACH1 adenoviral construct. Importantly, reducing lncDACH1 expression via either a cardiomyocyte-specific knockout or using shRNA had the opposite effect: INa was increased in isolated cells, as was conduction velocity in heart. Experiments were also conducted with a fragment of lnDACH1 identified by its conservation with other mammalian species. Overexpression of this fragment resulted in reduced INa and greater proarrhythmic behavior. Alteration of expression was confirmed by qPCR.
The mechanism by which lnDACH1 exerts its effects on INa was explored by measuring protein levels from cell fractions and immunofluorescence localization in cells. In general, overexpression was reported to reduce Nav1.5 and dystrophin levels and knockout or knockdown increased them.
The strengths of this manuscript include convincing evidence of a link between lncDACH1 and Na channel function. The identification of a lncDACH1 segment conserved among mammalian species is compelling. The observation that lncDACH1 is increased in a heart failure model and provides a plausible hypothesis for disease mechanism.
One limitation of the fractionation approach is the uncertain disposition of Na channel protein deemed "cytoplasmic." It seems likely that the membrane fraction includes ER membrane. The signal may reasonably be attributed to Na channel protein in stalled transport vesicles, or alternatively in stress granules, but this was not directly addressed.
Tags
Annotators
URL
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
In this study the authors aim to elucidate the role of RAPSYN in BCR-ABL-mediated leukemogenesis. RAPSYN is mainly known as a scaffolding protein for anchoring acetylcholine receptors (AChRs) to the cytoskeleton in muscle cells, facilitating AChR clustering through neddylation (Li et al., 2016). The authors demonstrate, through a broad and rigorous array of biochemical assays, that RAPSYN also plays a crucial role in the neddylation of BCR-ABL in leukemia cells. Their results indicate that this process shields BCR-ABL from ubiquitination and subsequent degradation, likely through a mechanism involving competition for binding with the BCR-ABL ubiquitin ligase c-CBL. In addition, the authors delve into the regulatory mechanisms underlying RAPSYN stability, demonstrating that it is enhanced through phosphorylation by SRC. This discovery further deepens our understanding of the complex dynamics of the molecular interactions that regulate BCR-ABL stability in leukemia.
To confirm the physiological significance of their findings, the authors effectively utilize cell viability assays and in vivo models. The integration of these approaches lends strength and validity to their conclusions.
The implications of the findings presented in this study are important, particularly in relation to our understanding of the pathogenesis and potential therapeutic strategies for Philadelphia chromosome-positive leukemias. By illuminating the role of RAPSYN in the regulation of BCR-ABL stability, this research potentially uncovers avenues for the development of targeted therapies, making a significant contribution to the field.
Two areas of the study could benefit from additional validation and exploration:
(1) The authors propose that targeting RAPSYN in Ph+ leukemia could have a high therapeutic index, suggesting that inhibition of RAPSYN may lead to cytotoxicity in Ph+ leukemia with high specificity and minimal side effects. The authors now include data showing RAPSYN knockdown in HS-5 cells does not affect cell growth (Figure 1C), supporting this assertion. This observation presents a contrast to DepMap data (https://depmap.org/), where RNAi and CRISPR-mediated RAPSYN depletion across hundreds of cell lines does not exhibit obvious differential effects on cell viability compared to Ph+ leukemia cell lines. Therefore, while the current results are promising, they call for additional validation by future studies to confirm RAPSYN as a viable therapeutic target in this context.
(2) A particularly notable yet underexplored aspect of this study is the observed disparity between RAPSYN protein and mRNA levels in Ph+ patient samples and cell lines. There is a marked enrichment of RAPSYN protein levels (Figure 1A, B) despite seemingly unchanged mRNA levels (Supplementary Figure 1 A-C). The authors convincingly demonstrate that RAPSYN stabilizes BCR-ABL, while SRC-mediated phosphorylation in turn stabilizes RAPSYN. This points to a specific, SRC-driven stabilization mechanism of RAPSYN in the Ph+ leukemia context. Consequently, the question arises whether BCR-ABL (through activation of SRC) reciprocally stabilize RAPSYN? Exploring the effects of BCR-ABL depletion on RAPSYN levels could shed light on this potential two-way stabilization mechanism, offering deeper insight into the complex molecular dynamics of RAPSYN and BCR-ABL in Ph+ leukemias.
In conclusion, this study represents a pivotal advancement in our understanding of Philadelphia chromosome-positive leukemias. It uniquely positions RAPSYN, a protein previously not associated with leukemogenesis, as a key regulator of BCR-ABL stability. Future research is essential to establish RAPSYN's potential as a therapeutic target and to more comprehensively understand its role in this context.
Comments on revised version:
I acknowledge and appreciate the author responses. Below are our comments on each reply:
Reply 1: Your response and the inclusion of data regarding RAPSYN knockdown in HS-5 cells adequately address the concerns.
Reply 2: The issue of the disparity between RAPSYN protein and mRNA levels in Ph+ leukemias has not sufficiently been resolved. Refer to point 2 in the revised review for more details. If conducting the proposed experiment is not feasible, I recommend a more thorough discussion in the manuscript to address and hypothesize about the causes of this discrepancy between protein and mRNA levels.
Reply 3: Your rationale for not performing additional assays with inactive mutants is satisfactory.
Reply 4: The clarification provided in your revision of the method section and the reorganization of Figure 6 successfully resolve the previously noted discrepancies. However, to ensure consistency and clarity across the paper, I recommend that you also specify the batches of constructs/viruses used in other relevant figures, such as Figure 1E.
Reply 5: The clarification provided on the immunoblots sufficiently addresses the concern raised.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Bae and colleagues substantially improved the data quality and revised their manuscript "Single cell transcriptome analysis of cavernous tissues reveals the key roles of pericytes in diabetic erectile dysfunction". While these revisions clarify some of the concerns raised, others remain. In my view, the following question must be addressed:
In my prior question on #3, I completely disagree with the statement that "identified cells with pericyte-like characteristics in the walls of large blood vessels". The staining that authors provided for LBH, was clearly stained for SMCs, not pericytes. Per Fig 2E, the authors are correct that LBH is colocalized with SMA+ cells( SMCs). However, the red signal from LBH clearly stains endothelial cells. In the rest of 2E and 2D, LBH is CD31- and their location suggests LBH stained for SMCs in the Aorta, Kidney vasculature, Dorsal vein, and Dorsal Artery.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Watanabe, Takashi, et al. investigated the use of the Golden Gate dual-expression vector system to enhance the modern standard for rapid screening of recombinant monoclonal antibodies. The presented data builds upon modern techniques that currently use multiple expression vectors to express heavy and light chain pairs. In a single vector, they express the linked heavy and light chain variable genes with a membrane-bound Ig which allows for rapid and more affordable cell-based screening. The final validation of H1 and H2 strain influenza screening resulted in 81 "H1+", 48 "H2+", and 9 "cross" reactive clones. The kinetics of some of the soluble antibodies were tested via SPR and validated with a competitive inhibition with classical well-characterized neutralizing clones.
Strengths:
In this study, Watanabe, Takashi, et al. further develop and refine the methodologies for the discovery of monoclonal antibodies. They elegantly merge newer technologies to speed up turnaround time and reduce the cost of antibody discovery. Their data supports the feasibility of their technique.
This study will have an impact on pandemic preparedness and antibody-based therapies.
Weaknesses:
A His tagged antigen was used for immunization and H1-his was used in all assays. Either the removal of His specific clones needs to be done before selection, or a different tag needs to be used in the subsequent assays.
This assay doesn't directly test the neutralization of influenza but rather equates viral clearance to competitive inhibition. The results would be strengthened with the demonstration of a functional antibody in vivo with viral clearance.
Limitations of this new technique are as follows: there is a significant loss of cells during FACs, transfection and cloning efficiency are critical to success, and well-based systems limit the number of possible clones (as the author discussed in the conclusions). Early enrichment of the B cells could improve efficiency, such as selection for memory B cells.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This is a straightforward manuscript assessing the specificity and efficiency of transgene expression in marmoset primary visual cortex (V1), for 4 different AAV vectors known to target transgene expression to either inhibitory cortical neurons (3 serotypes of AAV-h56D-tdTomato) or parvalbumin (PV)+ inhibitory cortical neurons in mice. Vectors are injected into the marmoset cortex and then postmortem tissue is analyzed following antibody labeling against GABA and PV. It is reported that: "in marmoset V1 AAV-h56D induces transgene expression in GABAergic cells with up to 91-94% specificity and 80% efficiency, depending on viral serotype and cortical layer. AAV-PHP.eB-S5E2 induces transgene expression in PV cells across all cortical layers with up to 98% specificity and 86-90% efficiency."
These claims are largely supported but slightly exaggerated relative to the actual values in the results presented. In particular, the overall efficiency for the best h56D vectors described in the results is: "Overall, across all layers, AAV9 and AAV1 showed significantly higher coverage (66.1{plus minus}3.9 and 64.9%{plus minus}3.7)". The highest coverage observed is just in middle layers and is also less than 80%: "(AAV9: 78.5%{plus minus}9.1; AAV1: 76.9%{plus minus}7.4)". For the AAV-PHP.eB-S5E2 the efficiency reported in the abstract ("86-90%) is also slightly exaggerated relative to the results: "Overall, across all layers coverage ranged from 78%{plus minus}1.9 for injection volumes >300nl to 81.6%{plus minus}1.8 for injection volumes of 100nl."
These data will be useful to others who might be interested in targeting transgene expression in these cell types in monkeys. Suggestions for improvement are to include more details about the vectors injected and to delete some comments about results that are not documented based on vectors that are not described (see below).
Major comments:
Details provided about the AAV vectors used with the h56D enhancer are not sufficient to allow assessment of their potential utility relative to the results presented. All that is provided is: "The fourth animal received 3 injections, each of a different AAV serotype (1, 7, and 9) of the AAV-h56D-tdTomato (Mehta et al., 2019), obtained from the Zemelman laboratory (UT Austin)." At a minimum, it is necessary to provide the titers of each of the vectors. It would also be helpful to provide more information about viral preparation for both these vectors and the AAVPHP.eB-S5E2.tdTomato. Notably, what purification methods were used, and what specific methods were used to measure the titers?
The first paragraph of the results includes brief anecdotal claims without any data to support them and without any details about the relevant vectors that would allow any data that might have been collected to be critically assessed. These statements should be deleted. Specifically, delete: "as well as 3 different kinds of PV-specific AAVs, specifically a mixture of AAV1-PaqR4-Flp and AAV1-h56D-mCherry-FRT (Mehta et al., 2019), an AAV1-PV1-ChR2-eYFP (donated by G. Horwitz, University of Washington)," and delete "Here we report results only from those vectors that were deemed to be most promising for use in primate cortex, based on infectivity and specificity. These were the 3 serotypes of the GABA-specific pAAV-h56D-tdTomato, and the PV-specific AAVPHP.eB-S5E2.tdTomato." These tools might in fact be just as useful or even better than what is actually tested and reported here, but maybe the viral titer was too low to expect any expression.
Based on the description in the Methods it seems that no antibody labeling against TdTomato was used to amplify the detection of the transgenes expressed from the AAV vectors. It should be verified that this is the case - a statement could be added to the Methods.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors used a combination of anchored hybrid enrichment and Sanger sequencing to construct a phylogenomic data set for the weevil family Belidae. Using evidence from fossils and previous studies they can estimate a phylogenetic tree with a range of dates for each node - a time tree. They use this to reconstruct the history of the belids' geographic distributions and associations with their host plants. They infer that the belids' association with conifers pre-dates the rise of the angiosperms. They offer an interpretation of belid history in terms of the breakup of Gondwanaland but acknowledge that they cannot rule out alternative interpretations that invoke dispersal.
Strengths:
The strength of any molecular-phylogenetic study hinges on four things: the extent of the sampling of taxa; the extent of the sampling of loci (DNA sequences) per genome; the quality of the analysis; and - most subjectively - the importance and interest of the evolutionary questions the study allows the authors to address. The first two of these, sampling of taxa and loci, impose a tradeoff: with finite resources, do you add more taxa or more loci? The authors follow a reasonable compromise here, obtaining a solid anchored-enrichment phylogenomic data set (423 genes, >97 kpb) for 33 taxa, but also doing additional analyses that included 13 additional taxa from which only Sanger sequencing data from 4 genes was available. The taxon sampling was pretty solid, including all 7 tribes and a majority of genera in the group. The analyses also seemed to be solid - exemplary, even, given the data available.
This leaves the subjective question of how interesting the results are. The very scale of the task that faces systematists in general, and beetle systematists in particular, presents a daunting challenge to the reader's attention: there are so many taxa, and even a sophisticated reader may never have heard of any of them. Thus it's often the case that such studies are ignored by virtually everyone outside a tiny cadre of fellow specialists. The authors of the present study make an unusually strong case for the broader interest and importance of their investigation and its focal taxon, the belid weevils.
The belids are of special interest because - in a world churning with change and upheaval, geologically and evolutionarily - relatively little seems to have been going on with them, at least with some of them, for the last hundred million years or so. The authors make a good case that the Araucaria-feeding belid lineages found in present-day Australasia and South America have been feeding on Araucaria continuously since the days when it was a dominant tree taxon nearly worldwide before it was largely replaced by angiosperms. Thus these lineages plausibly offer a modern glimpse of an ancient ecological community.
Weaknesses:
I didn't find the biogeographical analysis particularly compelling. The promise of vicariance biogeography for understanding Gondwanan taxa seems to have peaked about 3 or 4 decades ago, and since then almost every classic case has been falsified by improved phylogenetic and fossil evidence. I was hopeful, early in my reading of this article, that it would be a counterexample, showing that yes, vicariance really does explain the history of *something*. But the authors don't make a particularly strong claim for their preferred minimum-dispersal scenario; also they don't deal with the fact that the range of Araucaria was vastly greater in the past and included places like North America. Were there belids in what is now Arizona's petrified forest? It seems likely. Ignoring all of that is methodologically reasonable but doesn't yield anything particularly persuasive.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The authors report a quantitative comparative study regarding hind limb evolution among titanosaurs. I find the conclusions and findings of the manuscript interesting and relevant. The strength of the paper would be increased if the authors were to improve their reporting of taxon sampling and their discussion of age estimation and the potential implications that uncertainty in these estimates would have for their conclusions regarding gigantism (vs. ontogenetic patterns).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, Liu et al. identified an important pathway regulating the nuclear translocation of the key transcriptional factor FOG1 during human hematopoiesis. The authors show that heat shock cognate B (HSCB) can interact with and promote the proteasomal degradation of TACC3, and this function is independent of its role in iron-sulfur cluster biogenesis. TACC3 represses the activity of FOG1 by sequestering it in the cytoplasm. Therefore, HSCB can promote the nuclear translocation of FOG1 through down-regulating TACC3. The authors further show that the phosphorylation of HSCB by PI3K downstream of the EPO signaling pathway is important for its role in regulating the nuclear translocation of FOG1. The data are solid and the manuscript is overall well written. The findings of this manuscript provide important new knowledge to the fields of hematopoiesis and cell biology.
Strengths:
(1) This study uses a multi-pronged approach that combines techniques from a number of fields to convincingly demonstrate the pathway regulating the nuclear translocation of FOG1 during hematopoiesis. The proposed role of each component in the pathway is well supported by solid data.
(2) This work provides important new insights into the function of HSCB, which was known to be an iron-sulfur cluster assembly protein. This study identifies a new role of HSCB and shows that HSCB can regulate the stability of the TACC3 protein, and this cytoplasmic function of HSCB is regulated by protein phosphorylation by PI3K.
(3) The findings of this work open up new directions for research in hematopoiesis and related fields. For example, are there any other TACC3-binding proteins whose subcellular localization are regulated by the presence or absence of TACC3? What is the E3 ligase responsible for the degradation of TACC3? Does this identified mechanism contribute to the sideroblastic anemias observed in HSCB human patients and animal models?
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this study by Bendzunas et al, the authors show that the formation of intra-molecular disulfide bonds involving a pair of Cys residues near the catalytic HRD motif and a highly conserved T-Loop Cys with a BRSK-specific Cys at an unusual CPE motif at the end of the activation segment function as repressive regulatory mechanisms in BSK1 and 2. They observed that mutation of the CPE-Cys only, contrary to the double mutation of the pair, increases catalytic activity in vitro and drives phosphorylation of the BRSK substrate Tau in cells. Molecular modeling and molecular dynamics simulations indicate that oxidation of the CPE-Cys destabilizes a conserved salt bridge network critical for allosteric activation. The occurrence of spatially proximal Cys amino acids in diverse Ser/Thr protein kinase families suggests that disulfide-mediated control of catalytic activity may be a prevalent mechanism for regulation within the broader AMPK family. Understanding the molecular mechanisms underlying kinase regulation by redox-active Cys residues is fundamental as it appears to be widespread in signaling proteins and provides new opportunities to develop specific covalent compounds for the targeted modulation of protein kinases.
The authors demonstrate that intramolecular cysteine disulfide bonding between conserved cysteines can function as a repressing mechanism as indicated by the effect of DTT and the consequent increase in activity by BSK-1 and -2 (WT). The cause-effect relationship of why mutation of the CPE-Cys only increases catalytic activity in vitro and drives phosphorylation of the BRSK substrate Tau in cells is not clear to me. The explanation given by the authors based on molecular modeling and molecular dynamics simulations is that oxidation of the CPE-Cys (that will favor disulfide bonding) destabilizes a conserved salt bridge network critical for allosteric activation. However, no functional evidence of the impact of the salt-bridge network is provided. If you mutated the two main Cys-pairs (aE-CHRD and A-loop T+2-CPE) you lose the effect of DTT, as the disulfide pairs cannot be formed, hence no repression mechanisms take place, however when looking at individual residues I do not understand why mutating the CPE only results in the opposite effect unless it is independent of its connection with the T+2residue on the A-loop.
Strengths:
This is an important and interesting study providing new knowledge in the protein kinase field with important therapeutic implications for the rationale design and development of next-generation inhibitors.
Comments on revised version:
The authors have satisfactorily addressed my concerns.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Longhurst et al. assessed cell cycle regulators using a chemogenetic CRISPR-Cas9 screen in haploid human cell line HAP1. Besides known cell cycle regulators they identified the PRC2.1 subcomplex to be specifically involved in G1 progression, given that the absence of members of the complex makes the cells resistant to Palbociclib. They further showed that in HAP1 cells the PRC2.1, but not the PRC2.2 complex is important to repress the cyclins CCND1 and CCND2. This can explain the enhanced resistance to Palbociclib, a CDK4/6-Inhibitor, after PRC2.1 deletion.
Strengths:
The initial CRISPR screen is very interesting because it uses three distinct chemicals that disturb the cell cycle at various stages. This screen mostly identified known cell cycle regulators, which demonstrates the validity of the approach. The results can be used as a resource for future research.
The most interesting outcome of the experiment is the finding that knockouts of the PRC2.1 complex make the cell resistant to Palbociclib. In a further experiment, the authors focused on MTF2 and JARID2 as the main components of PRC2.1 and PRC2.2, respectively. Via extensive analyses, including genome-wide experiments, they confirmed that MTF2 is particularly important to repress the cyclins CCND1 and CCND2. The absence of MTF2 therefore leads to increased expression of these genes, sufficient to make the cell resistant to palociclib. This result will likely be of wide interest to the community.
Weaknesses:
The main weakness of the manuscript is that the experiments were performed in only one cell line. To draw more general conclusions, it would be essential to confirm some of the results in other cell lines.<br /> In addition, some of the findings, such as the results from the CRISPR screen as well as the stronger impact of the MTF2 KO on H3K27me3 and gene expression (compared to JARID2 KO), are not unexpected, given that similar results were already obtained before by other labs.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this study, the authors set out to use an unbiased CRISPR/Cas9 screen in CHO cells to identify genes encoding proteins that either increase or repress ATF6 signaling in CHO cells.
Strengths:
The strengths of the paper include the thoroughness of the screens, the use of a novel, double ATF6/IRE1 UPR reporter cell line, and follow-up detailed experiments on two of the findings in the screens, i.e. FURIN and CRT, to test the validity of involvement of each as direct regulators of ATF6 signaling. Additional strengths are the control experiments that validate the ATF6 specificity of the screens, as well as, for CRT, the finding of focus, determining roles for the glycosylation and cysteines in ATF6 as mechanistically involved in how CRT represses ATF6, at least in CHO cells.
Weaknesses:
The weaknesses of the paper are that the authors did not describe why they focused only on the top 100 proteins in each list of ATF6 activators and repressors. Additionally, there were a few methodology items missing, such as the nature of where the insertion site in the CHO cell genome of the XBP1::mCherry reporter. Since the authors go to great lengths to insert the other reporter for ATF6 activation in a "safe harbor" location, it leads to questions about whether the XBP1::mCherry reporter insertion is truly innocuous. An additional weakness is that the evidence for the physical interaction between ATF6LD and CRT is not strong, being dependent mainly on a single IP/IB experiment in Figure 4C that comprises only 1 lane on the gel for each of the test cases. Moreover, while that figure suggests that the interaction between CRT and ATF6 is decreased by mutating out the glycosylation sites in the ATF6LD, the BLI experiment in the same figure, 4B, suggests that there are no differences in the affinities of CRT for ATF6LD WT, deltaGly and deltaCys. An additional detail is that I found Figure 6A to be difficult to interpret, and that 6B was required in order for me to best evaluate the points being made by the authors in this figure.
Overall, I believe that this work will positively impact the field as it provides a list of potential regulators of ATF6 activation and repression that others will be able to use as a launch point for discovering such interactions in cells and tissues or interest beyond CHO cells. However, I agree with the authors that these findings were in CHO cell lines and that it is possible, if not likely, that some of the interactions they found will be cell type/line specific.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The PAClight1 sensor was developed using an approach successful for the development of other fluorescence-based GPCR sensors, which is the complete replacement of the third intracellular loop of the receptor with a circularly-permuted green fluorescent protein. When expressed in HEK cells, this sensor showed good expression and a weak but measurable response to the extracellular presence of PACAP1-38 (a F/Fo of 43%). Additional mutation near the site of insertion of the linearized GPF, at the C-terminus of the receptor, and within the second intracellular loop produced a final optimized sensor with F/Fo of >1000%. Finally, screening of mutational libraries that also included alterations in the extracellular ligand-binding domain of the receptor yielded a molecule, PAClight1P78A, that exhibited a high ligand-dependent fluorescence response combined with a high differential sensitivity to PACAP (EC50 30 nM based on cytometric sorting of stably transfected HEK293 cells) compared to its congener VIP, (with which PACAP shares two highly related receptors, VPAC1 and VPAC2) as well as several unrelated neuropeptides, and significantly slowed activation kinetics by PACAP in the presence of a 10-fold molar excess of the PAC1 antagonist PACAP6-38. A structurally highly similar control construct, PAClight1P78Actl, showed correspondingly similar basal expression in HEK293 cells, but no PACAP-dependent enhancement in fluorescent properties.
PAClight1P78A was expressed in neurons of the mouse cortex via AAV9.hSyn-mediated gene transduction. Slices taken from PAClight1P78A-transfected cortex, but not slices taken from PAClight1P78Actl-transfected cortex exhibited prompt and persistent elevation of F/Fo after 2 minutes of perfusion with PACAP1-38 which persisted for up to 14 minutes and was statistically significant after perfusion with 3000, but not 300 or 30 nM, of peptide. Likewise, microinfusion of 200 nL of 300 uM PACAP1-38 into the cortex of optical fiber-implanted freely moving mice elicited a F/Fo (%) of greater than 15, and significantly higher than that elicited by application of similar concentrations of VIP, CRF, or enkephalin, or vehicle alone. In vivo experiments were carried out in zebrafish larvae by the introduction of PAClight1P78A into single-cell stage Danio rerio embryos using a Tol2 transposase-based plasmid with a UAS promoter via injection (of plasmid and transposase mRNA), and sorting of post-fertilization embryos using a marker for transgenesis carried in the UAS : PAClight1P78A construct. Expression of PAClight1P78A was directed to cells in the olfactory bulb which express the fish paralog of the human PAC1 receptor by using the Tg(GnRH3:gal4ff) line, and fluorescent signals were elicited by intracerebroventricular administration of PACAP1-38 at a single concentration (1 mM), which were specific to PACAP and to the presence of PAClight1P78A per se, as controlled by parallel experiments in which PAClight1P78Actl instead of PAClight1P78A was contained in the transgenic plasmid.
Major strengths and weaknesses of the methods and results:
The report represents a rigorous demonstration of the elicitation of fluorescent signals upon pharmacological exposure to PACAP in nervous system tissue expressing PAClight1P78A in both mammals (mice) and fish (zebrafish larvae). Figure 4d shows a change in GFP fluorescence activation by PACAP occurring several seconds after the cessation of PACAP perfusion over a two-minute period, and its persistence for several minutes following. One wonders if one is apprehending the graphical presentation of the data incorrectly, or if the activation of fluorescence efficiency by ligand presentation is irreversible in this context, in which case the utility of the probe as a real-time indicator, in vivo, of released peptide might be diminished.
Appraisal of achievement of aims, and data support of conclusions:
Small cavils with controls are omitted for clarity; the larger issue of appraisal of results based on the scope of the designed experiments is discussed in the section below. An interesting question related to the time dependence of the PACAP-elicited activation of PAClight1P87A is its onset and reversibility, and additional data related to this would be welcome.
Discussion of the impact of the work, and utility of the methods and data:
Increasingly, neurotransmitter function may be observed in vivo, rather than by inferring in vivo function from in vitro, in cellular, or ex vivo experimentation. This very valuable report discloses the invention of a genetically encoded sensor for the class B1 GPCR PAC1. PAC1 is the major receptor for the neuropeptide PACAP, which in turn is a major neurotransmitter involved in brain response to psychogenic stress, or threat, in vertebrates as diverse as mammals and fishes. If this sensor possesses the sensitivity to detect endogenously released PACAP in vivo it will indeed be an impactful tool for understanding PACAP neurotransmission (and indeed PACAP action in general, in immune and endocrine compartments as well) in future experiments.
However, the sensor has not yet been used to detect endogenously released PACAP. Until this has been done, one cannot answer the question as to whether the levels of exogenously perfused/administered PACAP used here merely to calibrate the sensor's sensitivity are indeed unphysiologically high. If endogenous PACAP levels don't get that high, then the sensor will not be useful for its intended purpose. The authors should address this issue and allude to what kind of experiments would need to be done in order to detect endogenous PACAP release in living tissue in intact animals. The authors could comment upon the success of other GPCR sensors that have been used to observe endogenous ligand release, and where along the pathway to becoming a truly useful reagent this particular sensor is.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This manuscript combines live yeast cell imaging and other genomic approaches to study how transcription factor (TF) condensates might help organize and enhance the transcription of the target genes in the methionine starvation response pathway. The authors show that the TFs in this response can form phase-separated condensates through their intrinsically disordered regions (IDRs), and mediate the spatial clustering of the related endogenous genes as well as reporter inserted near the endogenous target loci.
Strengths:
This work uses rigorous experimental approaches, such as imaging of endogenously labeled TFs, determining expression and clustering of endogenous target genes, and reporter integration near the endogenous target loci. The importance of TFs is shown by rapid degradation. Single-cell data are combined with genomic sequencing-based assays. Control loci engineered in the same way are usually included. Some of these controls are very helpful in showing the pathway-specific effect of the TF condensates in enhancing transcription.
Weaknesses:
Perhaps the biggest weakness of this work is that the role of IDR and phase separation in mediating the target gene clustering is unclear. This is an important question. TF IDRs may have many functions including mediating phase separation and binding to other transcriptional molecules (not limited to proteins and may even include RNAs). The effect of IDR deletion on reduced Fano number in cells could come from reduced binding with other molecules. This should be tested on phase separation of the purified protein after IDR deletion. Also, the authors have not shown IDR deletion affects the clustering of the target genes, so IDR deletion may affect the binding of other molecules (not the general transcription machinery) that are specifically important for target gene transcription. If the self-association of the IDR is the main driving force of the clustering and target gene transcription enhancement, can one replace this IDR with totally unrelated IDRs that have been shown to mediate phase separation in non-transcription systems and still see the gene clustering and transcription enhancement effects? This work has all the setup to test this hypothesis.
The Met4 protein was tagged with MBP but Met 32 was not. MBP tag is well known to enhance protein solubility and prevent phase separation. This made the comparison of their in vitro phase behavior very different and led the authors to think that maybe Met32 is the scaffold in the co-condensates. If MBP was necessary to increase yield and solubility during expression and purification, it should be cleaved (a protease cleavage site should be engineered) to allow phase separation in vitro.
Are ATG36 and LDS2 also supposed to be induced by -met? This should be explained clearly. The signals are high at -met.
Figure 6B, the Met4-GFP seems to form condensates at all three loci without a very obvious difference, though 6C shows a difference. 6C is from only one picture each. The authors should probably quantify the signals from a large number of randomly selected pictures (cells) and do statistics.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors have performed a detailed analysis of the complex transcriptional status of numerous cell types present in wounded tissue, including keratinocytes, fibroblasts, macrophages, neutrophils, and endothelial cells. The comparison between infected and uninfected wounds is interesting and the analysis suggests possible explanations for why infected wounds are delayed in their healing response.
Strengths:
The paper presents a thorough and detailed analysis of the scRNAseq data. The paper is clearly written and the conclusions drawn from the analysis are appropriately cautious. The results provide an important foundation for future work on the healing of infected and uninfected wounds.
Weaknesses:
The analysis is purely descriptive and no attempt is made to validate whether any of the factors identified are playing functional roles in wound healing. Such experiments would be appropriate for followup work. The experimental setup is analyzing a single time point and does not include a comparison to unwounded skin. Nevertheless, the present data do provide a useful point of comparison for the field.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The widely distributed pannexin 1 (PANX1) is an ATP-permeable channel that plays an important role in intercellular communication and has been implicated in various pathophysiological processes and diseases. Previous studies have demonstrated that PANX1 can be phosphorylated at two molecular sites via the non-receptor kinase Src, thereby leading to channel opening and ATP release. In this paper, the authors used a variety of methods to detect tyrosine phosphorylation modification of PANX1 channel protein, however, their results showed that commercially available antibodies against the two phosphorylation sites used in previous studies did not work well, in other words, phosphorylation changes in PANX1 could not be detected by those antibodies. Therefore, the authors call for the re-examination and evaluation of previous research results.
In general, this is a meticulous study, using different detection methods and different expression systems.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors developed a 3D multi-cellular platform mimicking the complex interplays involved in the pathogenesis of NAFLD/NASH by employing hiPSCs-derived parenchymal and non-parenchymal cells in combination of organoids obtained from primary human cholangiocytes and the human hepatic stellate cell line LX2. They show that hiPSC-derived hepatocyte are able to accumulate intracellular lipids in fashion similar to human NAFLD and that prolonged accumulation leads to activation of inflammatory and fibrogenic pathways.
Strengths:
This is an original attempt to create a 3D all-human multicellular cellular platform recapitulating human NAFLD/NASH. The results are very encouraging. It is of particular note the fact that fibrogenic markers in the 3D system are not extremely (artificially) activated as in the classic 2D system. This makes the proposed platform more realistic.
Weaknesses:
The mixture of hiPSC-derived cells and primary or cell-line cells is understandable although potentially adding some variability to the system. The only unclear aspect is the characteristic of the collagen used to create the 3D system. Which type of collagen? Human? Which stiffness?
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors utilized (permeabilized) fibers from muscle samples obtained from brown and black bears, squirrels, and Garden dormice, to provide interesting and valuable data regarding changes in myosin conformational states and energetics during hibernation and different types of activity in summer and winter. Assuming that myosin structure is similar between species then its role as a regulator of metabolism would be similar and not different, yet the data reveal some interesting and perplexing differences between the selected hibernating species.
Strengths:
The experiments on the permeabilized fibers are complementary, sophisticated, and well-performed, providing new information regarding the characteristics of skeletal muscle fibers between selected hibernating mammalian species under different conditions (summer, interarousal, and winter).
The studies involve complementary assessments of muscle fiber biochemistry, sarcomeric structure using X-ray diffraction, and proteomic analyses of posttranslational modifications.
Weaknesses:
It would be helpful to put these findings on permeabilized fibers into context with the other anatomical/metabolic differences between the species to determine the relative contribution of myosin energetics (with these other contributors) to overall metabolism in these different species, including factors such as fat volume/distribution.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The study by Mowla et al analysed seminal microbiome together with semen quality parameters in fertile men and men from infertile couples with different infertility diagnoses. The study is of potential interest, with solid study design and methodology, nevertheless, the statistical analysis approach is not fully justified.
-The patient groups have different diagnoses and should be handled as different groups, and not fused into one 'patient' group in analyses.<br /> Why are the data in tables presented as controls and cases? I would consider men from couples with recurrent pregnancy loss, unexplained infertility, and male factor infertility to have different seminal parameters (not to fuse them into one group). This means, that the statistical analyses should be performed considering each group separately, and not to fuse 3 different infertility diagnoses into one patient group.
-Were any covariables included in the statistical analyses, e.g. age, BMI, smoking, time of sexual abstinence, etc?
-Furthermore, it is known that 16S rRNA gene analysis does not provide sensitive enough detection of bacteria on the species level. How much do the authors trust their results on the species level?
-Were the analyses of bacterial genera and species abundances with seminal quality parameters controlled for diagnosis and other confounders?
Strengths:
The cohort of participants seems to be homogenous in the sense of ethnicity and location.
The authors stress that their study is the biggest on the microbiome in semen. However, when considering that the study consists of 4 groups (with n=46-63), it does not stand out from previous studies.
Weaknesses:
There is a lack of paired seminal/urinal samples.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this study, the authors performed a screening for PDXP inhibitors to identify compounds that could increase levels of pyridoxal 5'- phosphate (PLP), the co-enzymatically active form of vitamin B6. For the screening of inhibitors, they first evaluated a library of about 42,000 compounds for activators and inhibitors of PDXP and secondly, they validated the inhibitor compounds with a counter-screening against PGP, a close PDXP relative. The final narrowing down to 7,8-DHF was done using PLP as a substrate and confirmed the efficacy of this flavonoid as an inhibitor of PDXP function. Physiologically, the authors show that, by acutely treating isolated wild-type hippocampal neurons with 7,8-DHF they could detect an increase in the ratio of PLP/PL compared to control cultures. This effect was not seen in PDXP KO neurons.
Strengths:
The screening and validation of the PDXP inhibitors have been done very well because the authors have performed crystallographic analysis, a counter screening, and mutation analysis. This is very important because such rigor has not been applied to the original report of 7,8 DHF as an agonist for TrkB. Which is why there is so much controversy on this finding.
Weaknesses:
As mentioned in the summary report the study may benefit from some in vivo analysis of PLP levels following 7,8-DHF treatment, although I acknowledge that it may be challenging because of the working out of the dosage and timing of the procedure.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Summary:
This manuscript reports the novel observation of alterations in the nuclear pore (NUP) components and the function of the nuclear envelope in knock-in models of APP and presenilin mutations. The data show that loss of NUP immunoreactivity (IR) and pore density are observed at times prior to plaque deposition in this model. The loss of NUP IR is correlated with an increase in intraneuronal Abeta IR with two monoclonal antibodies that react with the N-terminus of Abeta. Similar results are observed in cultured neurons from APP-KI and Wt mice where further results with cultured neurons indicate that Abeta "drives" this process: incubation of neurons with oligomeric, but not monomeric or fibrillar Abeta causes loss of NUP IR, incubation with conditioned media from KI cells but not wt cells also causes loss of NUP IR and treatment with the gamma secretase inhibitor, NAPT partially blocks the loss of NUP IR. Further data show that nuclear envelope function is altered in KI cells and KI cells are more sensitive to TNFalpha-induced necroptosis. This is potentially an important and significant report, but how this fits within the larger picture of what is known about amyloid aggregation and accumulation and pathogenesis in neurons needs to be clarified. The results from mouse brains are strong, while the results from cultured cells are in some instances are of a lower magnitude, less convincing, ambiguous, and sometimes over-interpreted.
Comments on revised version:
I am disappointed in the responses submitted in the revised manuscript. Although there are two new supplemental figures shown, there is no new data that would be needed to address the points raised by myself and the other reviewers. For example, I asked the authors to provide data to place their observations on lower levels of NUPs and mislocalization of nuclear proteins in the context of previously published reports of nuclear amyloid pathology in APP mouse models reported by Pensalfini et al 2014 and Lee et al, 2022 who report amyloid fibrils in some neuronal nuclei along with rosettes of perinuclear autophagic vacuoles containing Abeta immunoreactive material that also stains with amyloid fibril-specific antibodies. In response the authors state: "We have devoted a section of the discussion to highlight some of these findings in the context of Pensalfini et al. 2014 and Lee et al. 2022. Lee et al. tested multiple animal strains to observe the Panthos structures but did not use the App KI mouse model. Since none of our experiments directly tested their observations (e.g. perinuclear fibrils or acidity of autophagic vesicles) in App KI, we decided to take a more conservative approach in our interpretations by framing the NPC deficits without specifying the nature of the intracellular Aβ. We note in discussion that it is entirely possible that App KI animals also show the same Panthos phenotypes and the perinuclear accumulation of Aβ which results in damaged NUPs. To do that, the Panthos phenotype must first be established in App KI mice. "
But the "discussion" is just a couple of sentences that misrepresents the findings of the previous publications and excuses for not doing experiments that the authors should do, like examining whether neurons with intranuclear amyloid and perinuclear autophagic vacuoles occur in the mouse model they use. They are experiments that they should do, and it would be easy to do. Is not an imposition to ask for this data because they presumably have the mouse brain tissue, so they could cut more brain sections and co-stain them with NUP antibodies and the antibodies against fibrillar Abeta and autophagic vesicle markers.
This is just one of many comments where new data is needed but not provided. Disappointing that the revised manuscript is not significantly improved.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors identified that two of the placental CALHM orthologs, CALHM2 and CALHM4 can form heterooligomeric channels that are stable following detergent solubilization. By adding fiducial markers that specifically recognize either CALHM2 or CALHM4, the authors determine a cryo-EM density map of heterooligomeric CALHM2/CALHM4 from which they can determine how the channel in assembled. Surprisingly, the two orthologs segregate into two distinct segments of the channel. This segregation enables the interfacial subunits to ease the transition between the preferred conformations of each ortholog, which are similar to the confirmation that each ortholog adopts in homooligomeric channels.
Strengths:
Through the use of fiducial markers, the authors can clearly distinguish between the CALHM2 and CALHM4 promoters in the heterooligomeric channels, strengthening their assignment of most of the promoters. The authors take appropriate caution in identifying two subunits that are likely a mix of the two orthologs in the channel.
Weaknesses:
Despite the authors' efforts, no currents could be observed that corresponded to CALHM2/CALHM4 channels and thus the functional effect of their interaction is not known.
-
- Apr 2024
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The study by Yeo and co-authors addresses a long-lasting issue about botulinum neurotoxin (BoNT) intoxication. The current view is that the toxin binds to its receptors at the axon terminus by its HCc domain and is internalized in recycled neuromediator vesicles just after release of the neuromediators. Then, the HCn domain assists the translocation of the catalytic light chain (LC) of the toxin through the membrane of these endocytic vesicles into the cytosol of the axon terminus. There, the LC cleaves its SNARE substrate and blocks neurosecretion. However, other views involving kinetic aspects of intoxication suggest that the toxin follows the retrograde axonal transport up to the nerve cell body and then back to the nerve terminus before cleaving its substrate.
In the current study, the authors claim that the BoNT/A (isotype A of BoNT) not only progresses to the cell body but once there, follows the retrograde transport trafficking pathway in a retromer-dependent fashion, through the Golgi apparatus, until reaching the endoplasmic reticulum. Next, the LC dissociates from the HC (a process not studied here) and uses the translocon Sec61 machinery to retro-translocate into the cytosol. Only then, the LC traffics back to the nerve terminus following the anterograde axonal transport. Once there, LC cleaves its SNARE substrate (SNAP25 in the case of BoTN/A) and blocks neurosecretion.
To reach their conclusion, Yeo and co-authors use a combination of engineered tools: a cell line able to differentiate into neurons (ReNcell VN), a reporter dual fluorescent protein derived from SNAP25, the substrate of BoNT/A (called SNAPR), the use of either native BoNT/A or a toxin to which three fragment 11 of the reporter fluorescent protein Neon Green (mNG) are fused to the N-terminus of the LC (BoNT/A-mNG11x3), and finally ReNcell VN transfected with mNG1-10 (a protein consisting of the first 10 beta strands of the mNG).
SNAPR is stably expressed all over in the ReNcell VN. SNAPR is yellow (red and green) when intact and becomes red only when cleaved by BoNT/A LC, the green tip being degraded by the cell. When the LC of BoNT/A-mNG11x3 reaches the cytosol in ReNcell VN transfected by mNG1-10, the complete mNG is reconstituted and emits a green fluorescence.
In the first experiment, the authors show that the catalytic activity of the LC appears first in the cell body of neurons where SNAPR is cleaved first. This phenomenon starts 24 h after intoxication and progresses along the axon towards the nerve terminus during an additional 24 h. In a second experiment, the authors intoxicate the ReNcell VN transfected by mNG1-10 using the BoNT/A-mNG11x3. The fluorescence appears also first in the soma of neurons, then diffuses in the neurites in 48 h. The conclusion of these two experiments is that translocation occurs first in the cell body and that the LC diffuses in the cytosol of the axon in an anterograde fashion.
In the second part of the study, the authors perform a siRNA screen to identify regulators of BoNT/A intoxication. Their aim is to identify genes involved in intracellular trafficking of the toxin and translocation of the LC. Interestingly, they found positive and negative regulators of intoxication. Regulators could be regrouped according to the sequential events of intoxication. Genes affecting binding to the cell-surface receptor (SV2) and internalization. Genes involved in intracellular trafficking. Genes involved in translocation such as reduction of the disulfide bond linking the LC to the HC and refolding in the cytosol. Genes involved in signaling such as tyrosine kinases and phosphatases. All these groups of genes may be consistent with the current view of BoNT intoxication within the nerve terminus. However, two sets of genes were particularly significant to reach the main conclusion of the work and definitely constitute an original finding important to the field. One set of genes consists in those of the retromer, the other relates to the Sec61 translocon. This should indicate that once endocytosed, the BoNT traffics from the endosomes to Golgi apparatus, then to the ER. Ultimately, the LC should translocate from the ER lumen to the cytosol using the Sec61 translocon. The authors further control that the SV2 receptor for the BoNT/A traffics along the axon in a retromer-dependent fashion and that BoNT/A-mNG11x3 traverses the Golgi apparatus by fusing the mNG1-10 to a Golgi resident protein.
Strengths:
The findings in this work are convincing. The experiments are carefully done and are properly controlled. In the first part of the study, both the activity of the LC is monitored together with the physical presence of the toxin. In the second part of the work, the most relevant genes that came out of the siRNA screen are checked individually in the ReNcell VN / BoNT/A reporter system to confirm their role in BoNT/A trafficking and retro-translocation.<br /> These findings are important to the fields of toxinology and medical treatment of neuromuscular diseases by BoNTs. They may explain some aspects of intoxication such as slow symptom onset, aggravation and appearance of central effects.
Weaknesses:
The findings antagonize the current view of the intoxication pathway that is sustained by a vast amount of observations. The findings are certainly valid, but their generalization as the sole mechanism of BoNT intoxication should be tempered. These observations are restricted to one particular neuronal model and engineered protein tools. Other models such as isolated nerve/muscle preparations display nerve terminus paralysis within minutes rather than days. Also, the tetanus neurotoxin (TeNT), which mechanism of action involving axonal transport to the posterior ganglia in the spinal cord is well described, takes between 5 and 15 days. It is thus possible that different intoxication mechanisms co-exist for BoNTs or even vary depending on the type of neurons.
Although the siRNA experiments are convincing, it would be nice to reach the same observations with drugs affecting the endocytic to Golgi to ER transport (such as Retro-2, golgicide or brefeldin A) and the Sec61 retrotranslocation (such as mycolactone). Then, it would be nice to check other neuronal systems for the same observations.
-
-
-
Reviewer #2 (Public Review):
In the presented manuscript, the authors first use structured microfluidic devices with gliding filamentous cyanobacteria inside in combination with micropipette force measurements to measure the bending rigidity of the filaments. The distribution of bending rigidities is very broad.
Next, they use triangular structures to trap the bacteria with the front against an obstacle. Depending on the length and rigidity, the filaments buckle under the propulsive force of the cells. The authors use theoretical expressions for the buckling threshold to infer propulsive force, given the measured length and (mean-) stiffnesses. They find nearly identical values for both species, 𝑓 ∼ (1.0 {plus minus} 0.6) nN∕µm, nearly independent of the velocity. These measurements have to be taken with additional care, as then inferred forces depend strongly on the bending rigidity, which already shows a broad distribution.
Finally, they measure the shape of the filament dynamically to infer friction coefficients via Kirchhoff theory. In this section they report a strong correlation with velocity and report propulsive forces that vary over two orders of magnitude.
From a theoretical perspective, not many new results are presented. The authors repeat the the well-known calculation for filaments buckling under propulsive load and arrive at the literature result of buckling when the dimensionless number (f L^3/B) is larger than 30.6 as previously derived by Sekimoto et al in 1995. In my humble opinion, the "buckling theory" section belongs to methods.<br /> Finally, the Authors use molecular dynamics type simulations similar to other models to reproduce the buckling dynamics from the experiments.
Data and source code are available via trusted institutional or third-party repositories that adhere to policies that make data discoverable, accessible and usable.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Yi-Ting Tsai and colleagues conducted a systematic analysis of the correlation between the expression of retrotransposable elements (RTEs) and aging, using publicly available transcriptional and methylome microarray datasets of blood cells from large human cohorts, as well as single-cell transcriptomics. Although DNA hypomethylation was associated with chronological age across all RTE biotypes, the authors did not find a correlation between the levels of RTE expression and chronological age. However, expression levels of LINEs and LTRs positively correlated with DNA demethylation, and inflammatory and senescence gene signatures, indicative of "biological age". Gene set variation analysis showed that the inflammatory response is enriched in the samples expressing high levels of LINEs and LTRs. In summary, the study demonstrates that RTE expression correlates with "biological" rather than "chronological" aging.
Strengths:
The question the authors address is both relevant and important to the fields of aging and transposon biology.
Weaknesses:
The choice of methodology does not fully support the primary claims. Although microarrays can detect certain intergenic transposon sequences, the authors themselves acknowledge in the Discussion section that this method's resolution is limited. More critical considerations, however, should be addressed when interpreting the results. The coverage of transposon sequences by microarrays is not only very limited (232 unique probes) but also predetermined. This implies that any potential age-related overexpression of RTEs located outside of the microarray-associated regions, or of polymorphic intact transposons, may go undetected. Therefore, the authors should be more careful while generalising their conclusions.
Additionally, for some analyses, the authors pool signals from RTEs by class or family, despite the fact that these groups include subfamilies and members with very different properties and harmful potentials. For example, while sequences of older subfamilies might be passively expressed through readthrough transcription, intact members of younger groups could be autonomously reactivated and cause inflammation. The aggregation of signals by the largest group may obscure the potential reactivation of smaller subgroups. I recommend grouping by subfamily or, if not possible due to the low expression scores, by subgroup. For example, all HERV subfamilies are from the ERVL family.
Next, Illumina arrays might not accurately represent the true abundance of TEs due to non-specific hybridization of genomic transposons. Standard RNA preparations always contain traces of abundant genomic SINEs unless DNA elimination is specifically thorough. The problem of such noise should be addressed.
Lastly, scRNAseq was conducted using 10x Genomics technology. However, quantifying transposons in 10x sequencing datasets presents major challenges due to sparse signals. Smart-seq single-cell technology is better suited to this particular purpose. Anyway, it would be more convincing if the authors demonstrated TE expression across different clusters of immune cells using standard scRNAseq UMAP plots instead of boxplots.
I recommend validating the data by RNAseq, even on small cohorts. Given that the connection between RTE overexpression and inflammation has been previously established, the authors should consider better integrating their observations into the existing knowledge.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors investigate the influence of serotonin on feeding behavior and electrophysiological responses in the antennal lobe of locusts. They find that serotonin injection changes behavior in an odor-specific way. In physiology experiments, they can show that projection neurons in the antennal lobe generally increase their baseline firing and odor responses upon serotonin injection. Using a modeling approach the authors propose a framework on how a general increase in antennal lobe output can lead to odor-specific changes in behavior.
Strengths:
This study shows that serotonin affects feeding behavior and odor processing in the antennal lobe of locusts, as serotonin injection increases activity levels of projection neurons. This study provides another piece of evidence that serotonin is a general neuromodulator within the early olfactory processing system across insects and even phyla.
Weaknesses:
I still have several concerns regarding the generalizability of the model and interpretation of results. The authors cannot provide evidence that serotonin modulation of projection neurons impacts behavior.
The authors show that odor identity is maintained after 5-HT injection, however, the authors do not show if PN responses to different odors were differently affected after serotonin exposure.
Regarding the model, the authors show that the model works for odors with non-overlapping PN activation. However, only one appetitive, one neutral, and one aversive odor has been tested and modeled here. Can the fixed-weight model also hold for other appetitive and aversive odors that might share more overlap between active PNs? How could the model generate BZA attraction in 5-HT exposed animals (as seen in behavior data in Figure 1) if the same PNs just get activated more?
The authors should still not exclude the possibility that serotonin injections could affect behavior via modulation of other cell types than projection neurons. This should still be discussed, serotonin might rather shut down baseline activation of local inhibitory neurons - and thus lead to the interesting bursting phenotypes, which can also be seen in the baseline response, due to local PN-to-LN feedback.
The authors did not fully tone down their claims regarding causality between serotonin and starved state behavioral responses.<br /> There is no proof that serotonin injection mimics starved behavioral responses.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Pyoverdines, siderophores produced by many Pseudomonads, are one of the most diverse groups of specialized metabolites and are frequently used as model systems. Thousands of Pseudomonas genomes are available, but large-scale analyses of pyoverdines are hampered by the biosynthetic gene clusters (BGCs) being spread across multiple genomic loci and existing tools' inability to accurately predict amino acid substrates of the biosynthetic adenylation (A) domains. The authors present a bioinformatics pipeline that identifies pyoverdine BGCs and predicts the A domain substrates with high accuracy. They tackled a second challenging problem by developing an algorithm to differentiate between outer membrane receptor selectivity for pyoverdines versus other siderophores and substrates. The authors applied their dataset to thousands of Pseudomonas strains, producing the first comprehensive overview of pyoverdines and their receptors and predicting many new structural variants.
The A domain substrate prediction is impressive, including the correction of entries in the MIBiG database. Their high accuracy came from a relatively small training dataset of A domains from 13 pyoverdine BGCs. The authors acknowledge that this small dataset does not include all substrates, and correctly point out that new sequence/structure pairs can be added to the training set to refine the prediction algorithm. The authors could have been more comprehensive in finding their training set data. For instance, the authors claim that histidine "had not been previously documented in pyoverdines", but the sequenced strain P. entomophila L48, incorporates His (10.1007/s10534-009-9247-y). The workflow cannot differentiate between different variants of Asp and OHOrn, and it's not clear if this is a limitation of the workflow, the training data, or both. The prediction workflow holds up well in Burkholderiales A domains, however, they fail to mention in the main text that they achieved these numbers by adding more A domains to their training set.
To validate their predictions, they elucidated structures of several new pyoverdines, and their predictions performed well. However, the authors did not include their MS/MS data, making it impossible to validate their structures. In general, the biggest limitation of the submitted manuscript is the near-empty methods section, which does not include any experimental details for the 20 strains or details of the annotation pipeline (such as "Phydist" and "Syndist"). The source code also does not contain the requisite information to replicate the results or re-use the pipeline, such as the antiSMASH version and required flags. That said, skimming through the source code and data (kindly provided upon request) suggests that the workflow itself is sound and a clear improvement over existing tools for pyoverdine BGC annotation.
Predicting outer membrane receptor specificity is likewise a challenging problem and the authors have made a promising achievement by finding specific gene regions that differentiate the pyoverdine receptor FpvA from FpvB and other receptor families. Their predictions were not tested experimentally, but the finding that only predicted FpvA receptors were proximate to the biosynthesis genes lends credence to the predictive power of the workflow. The authors find predicted pyoverdine receptors across an impressive 468 genera, an exciting finding for expanding the role of pyoverdines as public goods beyond Pseudomonas. However, whether or not these receptors can recognize pyoverdines (and if so, which structures!) remains to be investigated.
In all, the authors have assembled a rich dataset that will enable large-scale comparative genomic analyses. This dataset could be used by a variety of researchers, including those studying natural product evolution, public good eco/evo dynamics, and NRPS engineering.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This study by Ngo et al. uses mostly high-speed AFM to estimate conformational changes within actin filaments, as they get decorated by cofilin. The authors build on their earlier study (Ngo et al. eLife 2015) where they used the same technique to monitor the expansion of cofilin clusters on actin filaments, and the propagation of the associated conformational changes in the filament (reduction of the helical pitch). Here, they propose a higher-resolution description of the binding of cofilin to actin filaments.
Strengths:
The high speed AFM technique used here is quite original to address this question, compared to classical light and electron microscopy techniques. It can certainly bring valuable information as it provides a high spatial resolution while monitoring live events. Also, in this paper, a nice effort was made to make the 3D structures and conformational changes clear and understandable.
Weaknesses:
The paper also has a number of limitations, which I detail below.
In addition to AFM, the authors also propose a Principal Component Analysis (PCA) of exisiting structural data on actin protomers. However, this part seems very similar to another published work by others (Oda et al. JMB 2019), which is not even cited.
The asymmetrical growth of cofilin clusters has so far only been seen using AFM, by the same authors (Ngo et al. eLife 2015). Using fluorescent microscopy, others have reported a very symmetrical expansion of cofilin clusters (Wioland et al. Curr Biol 2017). This is not mentioned at all, here. It should be discussed, and explanations for this discrepancy could be proposed.
Regarding the AFM technique, I have the following concerns.
The filaments appear densely packed on the surface, and even clearly in register in some images (if not most images, e.g., Figs 3A, 4BC, 5A). Why is that? Isn't there a risk that this could affect the result? This suggests there is some interaction between the filaments.
The properties of the lipid layer and its interaction with the actin filaments are not clear at all. A poor control of these interactions is a problem if one aims to measure conformational changes at high resolution. The strength of the interaction appears tuned by the ratio of lipids put on the surface to change its electrostatic charge. A strong attachement likely does more than suppress torsional motion (as claimed in Fig 8A). It may also hinder cofilin binding in several ways (lower availability of binding sites on the filament facing the surface, electrostatic interactions between cofilin and the surface, etc.)
How do we know that the variations over time are not mostly experimental noise, i.e. variations between repeats of the same measurement? As shown in Fig 3, correlation is mostly lost from one image to the next, and rather stable after that.
The identification of cofilactin regions relies on the additional height of the "peaks", due to the presence of cofilin. It thus seems that cofilin is detected every half helical pitch (HHP), but not in between, thereby setting the resolution for the localization of cluster borders to one HHP. It thus seems difficult to claim that there is a change in helicity without cofilin decoration over this distance. In Fig 7, the change in helicity could be due to cofilin decoration that is undetected because cofilins have not yet reached the next peak.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
Transmembrane signaling in plants is crucial for homeostasis. In this study, the authors set out to understand to what extent catalytic activity in the EFR tyrosine kinase is required in order to transmit a signal. This work was driven by mounting data that suggest many eukaryotic kinases do not rely on catalysis for signal transduction, relying instead on conformational switching to relay information. The crucial findings reported here involve the realisation that a kinase-inactive EFR can still activate (ie lead to downstream phosphorylation) of its partner protein BAK1. Using a convincing set of biochemical, mass spectrometric (HD-exchange) and in vivo assays, the team suggest a model in which EFR is likely phosphorylated in the canonical activation segment (where two Ser residues are present), which is sufficient to generate a conformation that can activate BAK1 through dimersation. A model is put forward involving C-helix positioning in BAK1, and the model extended to other 'non-RD' kinases in Arabidopsis kinases that likely do not require kinase activity for signaling.
Strengths:
The work uses logical and well-controlled approaches throughout, and is clear and convincing in most areas, linking data from IPs, kinase assays (including clear 32P-based biochemistry), HD-MX data (from non-phosphorylated EFR) structural biology, oxidative burst data and infectivity assays. Repetitions and statistical analysis all appear appropriate.<br /> Overall, the work builds a convincing story and the discussion does a clear job of explaining the potential impact of these findings (and perhaps an explanation of why so many Arabidopsis kinases are 'pseudokinases', including XPS1 and XIIa6, where this is shown explicitly).
Weaknesses:
No major weaknesses are noted from reviewing the data and the paper follows a logical course built on solid foundations; the use of Tables to explain various experimental data pertinent to the reported studies is appreciated.
(1) The use of a, b,c, d in Figures 2C and 3C etc is confusing to this referee, and is now addressed in the latest version<br /> (2) The debate about kinase v pseudokinases is well over a decade old. For non-experts, the kinase alignments/issues raised are in PMID: 23863165 and might prove useful if cited.<br /> (3) Early on in the paper, the concept of kinases and pseudokinases related to R-spine (and extended R-spine) stability and regulation really needs to be more adequately introduced to explain what comes next; e.g. some of the key work in this area for RAF and Tyr kinases where mutual F-helix Phe amino acid changes are evaluated (conceptually similar to this study of the E-helix Tyr to Phe changes in EFR) should be cited (PMID: 17095602, 24567368 and 26925779).<br /> (4) In my version, some of the experimental text is also currently in the wrong order (and no page numbers, so hard for me to state exactly where in the manuscript); However, I am certain that Figure 2C is mentioned in the text when the data are actually shown in Figure 3C for the EFR-SSAA protein.<br /> (5) Tyr 156 in PKA is not shown in Supplement 1, 2A as suggested in the text; for readers, it will be important to show the alignment of the Tyr residue in other kinases; this has been updated in the second version. Although it is clearly challenging to generate phosphorylated EFR (seemingly through Codon-expansion here?), it appears unlikely that a phosphorylated EFR protein, even semi-pure, couldn't have been assayed to test the idea that the phosphorylation drives/supports downstream signaling. What about a DD or EE mutation, as commonly used (perhaps over-used) in MEK-type studies?
Impact:
The work is an important new step in the huge amount of follow-up work needed to examine how kinases and pseudokinases 'talk' to each other in (especially) the plant kingdom, where significant genetic expansions have occurred. The broader impact is that we might understand better how to manipulate signaling for the benefit of plants and mankind; as the authors suggest, their study is a natural progression both of their own work, and the kingdom-wide study of the Kannan group.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The study employs quantitative metabolomic and lipidomic analyses to scrutinize tumor interstitial fluid (TIF), adjacent normal kidney interstitial fluid (KIF), and plasma samples from renal cell carcinoma (RCC) patients. The authors delve into the intricate world of renal cell carcinoma and its tumor microenvironment, shedding light on the factors that shape nutrient availability in both cancerous and adjacent normal tissues. The authors prove that non-cancer-driven tissue factors play a dominant role in shaping nutrient availability in RCC. This finding opens up new avenues for research, suggesting that the tumor microenvironment is profoundly influenced by factors beyond the presence of cancer cells. This study not only contributes valuable insights into RCC metabolism but also prompts a reevaluation of the factors governing nutrient availability in tumor microenvironments more broadly. Overall, it represents a significant step forward in our understanding of the intricate interplay between cancer and its surrounding milieu.
The study is overall well-constructed, including appropriate analysis. Likewise, the manuscript is written clearly and supported by high-quality figures. Since the authors exclusively employed samples from RCC patients and did not include kidney interstitial fluid and plasma samples from healthy individuals, we cannot accurately assess the true significance and applicability of the results until the role of cancer cells in reshaping KIF is understood. In essence, some metabolite levels in the tumor interstitial fluid did not show an increase or decrease compared to the adjacent normal kidney interstitial fluid. However, the levels of these metabolites in both TIF and KIF might be higher or lower than those in kidney interstitial fluid from healthy individuals, and the roles of these metabolites should not be overlooked. Similar concerns extend to plasma levels, emphasizing the importance of metabolites that synchronously change in RCC TIF, KIF, and plasma-whether elevated or reduced.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, the authors investigated how partial loss of SynGap1 affects inhibitory neurons derived from the MGE in the auditory cortex, focusing on their synaptic inputs and excitability. While haplo-insufficiently of SynGap1 is known to lead to intellectual disabilities, the underlying mechanisms remain unclear.
Strengths:
The questions are novel
Weaknesses:
Despite the interesting and novel questions, there are significant concerns regarding the experimental design and data quality, as well as potential misinterpretations of key findings. Consequently, the current manuscript fails to contribute substantially to our understanding of SynGap1 loss mechanisms and may even provoke unnecessary controversies.
Major issues:
(1) One major concern is the inconsistency and confusion in the intermediate conclusions drawn from the results. For instance, while the sEPSC data indicates decreased amplitude in PV+ and SOM+ cells in cHet animals, the frequency of events remains unchanged. In contrast, the mEPSC data shows no change in amplitudes in PV+ cells, but a significant decrease in event frequency. The authors conclude that the former observation implies decreased excitability. However, traditionally, such observations on mEPSC parameters are considered indicative of presynaptic mechanisms rather than changes of network activity. The subsequent synapse counting experiments align more closely with the traditional conclusions. This issue can be resolved by rephrasing the text. However, it would remain unexplained why the sEPSC frequency shows no significant difference. If the majority of sEPSC events were indeed mediated by spiking (which is blocked by TTX), the average amplitudes and frequency of mEPSCs should be substantially lower than those of sEPSCs. Yet, they fall within a very similar range, suggesting that most sEPSCs may actually be independent of action potentials. But if that was indeed the case, the changes of purported sEPSC and mEPSC results should have been similar.
(2) Another significant concern is the quality of synapse counting experiments. The authors attempted to colocalize pre- and postsynaptic markers Vglut1 and PSD95 with PV labelling. However, several issues arise. Firstly, the PV labelling seems confined to soma regions, with no visible dendrites. Given that the perisomatic region only receives a minor fraction of excitatory synapses, this labeling might not accurately represent the input coverage of PV cells.<br /> Secondly, the resolution of the images is insufficient to support clear colocalization of the synaptic markers. Thirdly, the staining patterns are peculiar, with PSD95 puncta appearing within regions clearly identified as somas by Vglut1, hinting at possible intracellular signals. Furthermore, PSD95 seems to delineate potential apical dendrites of pyramidal cells passing through the region, yet Vglut1+ partners are absent in these segments, which are expected to be the marker of these synapses here.<br /> Additionally, the cumulative density of Vglut2 and Vglut1 puncta exceeds expectations, and it's surprising that subcortical fibers labeled by Vglut2 are comparable in number to intracortical Vglut1+ axon terminals. Ideally, N(Vglut1)+N(Vglut2) should be equal or less than N(PSD95), but this is not the case here. Consequently, these results cannot be considered reliable due to these issues.
(3) One observation from the minimal stimulation experiment was concluded by an unsupported statement. Namely, the change in the onset delay cannot be attributed to a deficit in the recruitment of PV+ cells, but it may suggest a change in the excitability of TC axons.
(4) The conclusions drawn from the stimulation experiments are also disconnected from the actual data. To make conclusions about TC release, the authors should have tested release probability using established methods, such as paired-pulse changes. Instead, the only observation here is a change in the AMPA components, which remained unexplained.
(5) The sampling rate of CC recordings is insufficient to resolve the temporal properties of the APs. Therefore, the phase-plots cannot be interpreted (e.g. axonal and somatic AP components are not clearly separated), raising questions about how AP threshold and peak were measured. The low sampling rate also masks the real derivative of the AP signals, making them apparently faster.<br /> A related issue is that the Methods section lacks essential details about the recording conditions, such as bridge balance and capacitance neutralization.
(6) Interpretation issue: One of the most fundamental measures of cellular excitability, the rheobase, was differentially affected by cHet in BCshort and BCbroad. Yet, the authors concluded that the cHet-induced changes in the two subpopulations are common.
(7) Design issue:<br /> The Kv1 blockade experiments are disconnected from the main manuscript. There is no experiment that shows the causal relationship between changes in DTX and cHet cells. It is only an interesting observation on AP halfwidth and threshold. However, how they affect rheobase, EPSCs, and other topics of the manuscript are not addressed in DTX experiments.<br /> Furthermore, Kv1 currents were never measured in this work, nor was the channel density tested. Thus, the DTX effects are not necessarily related to changes in PV cells, which can potentially generate controversies.
(8) Writing issues:<br /> Abstract:<br /> The auditory system is not mentioned in the abstract.<br /> One statement in the abstract is unclear. What is meant by "targeting Kv1 family of voltage-gated potassium channels was sufficient..."? "Targeting" could refer to altered subcellular targeting of the channels, simple overexpression/deletion in the target cell population, or targeted mutation of the channel, etc. Only the final part of the Results revealed that none of the above, but these channels were blocked selectively.<br /> Introduction:<br /> There is a contradiction in the introduction. The second paragraph describes in detail the distinct contribution of PV and SST neurons to auditory processing. But at the end, the authors state that "relatively few reports on PV+ and SST+ cell-intrinsic and synaptic properties in adult auditory cortex". Please be more specific about the unknown properties.
(9) The introduction emphasizes the heterogeneity of PV neurons, which certainly influences the interpretation of the results of the current manuscript. However, the initial experiments did not consider this and handled all PV cell data as a pooled population.
(10) The interpretation of the results strongly depends on unpublished work, which potentially provide the physiological and behavioral contexts about the role of GABAergic neurons in SynGap-haploinsufficiency. The authors cite their own unpublished work, without explaining the specific findings and relation to this manuscript.
(11) The introduction of Scholl analysis experiments mentions SOM staining, however, there is no such data about this cell type in the manuscript.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
This work brings important information regarding the composition of interneurons in the mammalian spinal cord, with a developmental perspective. Indeed, for the past decades, tools inspired from developmental biology have opened up promising avenues for challenging the functional heterogeneity in the spinal cord. They rely on the fact that neurons sharing similar mature properties also share a largely similar history of expression of specific transcription factor (TF) genes during embryogenic and postnatal development. For instance, neurons originating from p1 progenitors and expressing the TF Engrailed-1, form the V1 neuronal class. While such "cardinal" neuronal classes defined by one single RF indeed share numerous features - e.g., for the case of V1 neurons, a ventral positioning, an inhibitory nature and ipsilatetal projections - there is accumulating evidence for a finer-grained diversity and specialization in each class which is still largely obscure. The present work studies the heterogeneity of V1 interneurons and describes multiple classes based on their birthdate, final positioning, and expression of additional TF. It brings in particular a solid characterization of the Foxp2-expressing V1 interneurons for which authors also delve into the connectivity, and hence, possible functional implication. The work will be of interest to developmental biologists and those interested in the organization of the locomotor spinal network.
Strengths:
This study has deeply analyzed the diversity of V1 neurons by intersecting multiple criteria: TF expression, birthdate, location in the spinal cord, diversity along the rostro-caudal axis, and for some subsets, connectivity. This illustrates and exemplifies the absolute need to not consider cardinal classes, defined by one single TF, as homogeneous. Rather, it highlights the limits of single-TF classification, and exemplifies the existence of further diversity within cardinal class.
Experiments are generally well performed with a satisfactory number of animals and adequate statistical tests.
Authors have also paid strong attention to potential differences in cell-type classification when considering neurons currently expressing of a given TF (e.g., using antibodies), from those defined as having once expressed that TF (e.g., defined by a lineage-tracing strategy). This ambiguity is a frequent source of discrepancy of findings across studies.
Furthermore, there is a risk in developmental studies to overlook the fact that the spinal cord is functionally specialized rostro-caudally, and to generalize features that may only be applicable to a specific segment and hence to a specific motor pool. While motoneurons share the same dorso-ventral origin and appear homogenous on a ChAT staining, specific clusters are dedicated to specific muscle groups, e.g., axial, hypaxial or limb muscles. Here, the authors make the important distinction between different lumbar levels and detail the location and connectivity of their neurons of interest with respect to specific clusters of MN.
Finally, the authors are fully transparent on inter-animal variability in their representation and quantification. This is crucial to avoid the overgeneralization of findings but to rather provide a nuanced understanding of the complexities of spinal circuits.
Weaknesses:
The current version of the paper is VERY hard to read. It is often extremely difficult to "see the forest for the trees" and the reader is often drowned in methodological details that provide only minor additions to the scientific message. Non-specialists in developmental biology, but still interested in the spinal cord organization, especially students, might find this article challenging to digest and there is a high risk that they will be inclined to abandon reading it. The diversity of developmental stages studied (with possible mistakes between text and figures) adds a substantial complexity in the reading. It is also not clear at all why authors choose to focus on the Foxp2 V1 from page 9. Naively, the Pou6f2 might have been equally interesting. Finally, numerous discrepancies in the referencing of figures must also be fixed. I strongly recommend an in-depth streamlining and proofreading, and possibly moving some material to supplement (e.g. page 8, and elsewhere).
Second, and although the different V1 populations have been investigated in detail regarding their development and positioning, their functional ambition is not directly investigated through gain or loss of function experiments. For the Foxp2-V1, the developmental and anatomical mapping is complemented by a connectivity mapping (Fig 6s, 8), but the latter is fairly superficial compared to the former. Synapses (Fig 6) are counted on a relatively small number of motoneurons per animal, that may, or may not, be representative of the population. Likewise, putative synaptic inputs are only counted on neuronal somata. Motoneurons that lack of axono-somatic contacts may still be contacted distally. Hence, while this data is still suggestive of differences between V1 pools, it is only little predictive of function.
Third, I suggest taking with caution the rabies labelling (Figure 8). It is known that this type of Rabies vectors, when delivered from the periphery, might also label sensory afferents and their post-synaptic targets in the cord through anterograde transport and transneuronal spread (e.g., Pimpinella et al., 2022). Yet I am not sure authors have made all controls to exclude that labelled neurons, presumed here to be premotoneurons, could rather be anterogradely labelled from sensory afferents.
Fourth, the ambition to differentiate neuronal birthdate at a half-day resolution (e.g., E10 vs E10.5) is interesting but must be considered with caution. As the author explains in their methods, animals are caged at 7pm, and the plug is checked the next morning at 7 am. There is hence a potential error of 12h.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors have developed a novel bimanual task that allows them to study how the sensorimotor control system deals with redundancy within our body. Specifically, the two hands control two robot handles that control the position and orientation of a virtual stick, where the end of the stick is moved into a target. This task has infinite solutions to any movement, where the two hands influence both tip-movement direction and stick-tilt angle. When moving to different targets in the baseline phase, participants change the tilt angle of the stick in a specific pattern that produces close to the minimum movement of the two hands to produce the task. In a series of experiments, the authors then apply perturbations to the stick angle and stick movement direction to examine how either tip-movement (task-relevant) or stick-angle (task-irrelevant) perturbations affect adaptation. Both types of perturbations affect adaptation, but this adaptation follows the baseline pattern of tip-movement and stick angle relation such that even task-irrelevant perturbations drive adaptation in a manner that results in task-relevant errors. Overall, the authors suggest that these baseline relations affect how we adapt to changes in our tasks. This work provides an important demonstration that underlying solutions/relations can affect the manner in which we adapt. I think one major contribution of this work will also be the task itself, which provides a very fruitful and important framework for studying more complex motor control tasks.
Strengths:
Overall, I find this a very interesting and well-written paper. Beyond providing a new motor task that could be influential in the field, I think it also contributes to studying a very important question - how we can solve redundancy in the sensorimotor control system, as there are many possible mechanisms or methods that could be used - each of which produces different solutions and might affect the manner in which we adapt.
Weaknesses:
I would like to see further discussion of what the particular chosen solution implies in terms of optimality.
The underlying baseline strategy used by the participants appears to match the path of minimum movement of the two hands. This suggests that participants are simultaneously optimizing accuracy and minimizing some metabolic cost or effort to solve the redundancy problem. However, once the perturbations are applied, participants still use this strategy for driving adaptation. I assume that this means that the solution that participants end up with after adaptation actually produces larger movements of the two hands than required. That is - they no longer fall onto the minimum hand movement strategy - which was used to solve the problem. Can the authors demonstrate that this is either the case or not clearly? These two possibilities produce very different implications in terms of the results.
If my interpretation is correct, such a result (using a previously found solution that no longer is optimal) reminds me of the work of Selinger et al., 2015 (Current Biology), where participants continue to walk at a non-optimal speed after perturbations unless they get trained on multiple conditions to learn the new landscape of solutions. Perhaps the authors could discuss their work within this kind of interpretation. Do the authors predict that this relation would change with extensive practice either within the current conditions or with further exploration of the new task landscape? For example, if more than one target was used in the adaptation phase of the experiment?
On the other hand, if the adaptation follows the solution of minimum hand movement and therefore potentially effort, this provides a completely different interpretation.
Overall, I would find the results even more compelling if the same perturbations applied to movements to all of the targets and produced similar adaptation profiles. The question is to what degree the results derive from only providing a small subset of the environment to explore.
-
-
osf.io osf.io
-
Reviewer #2 (Public Review):
Summary:
The study focuses on how relatedness with existing memories affects the formation and retention of new memories. Of core interest were the conditions that determine when prior memories facilitate new learning or interfere with it. Across a set of experiments that varied the degree of relatedness across memories as well as retention interval, the study compellingly shows that relatedness typically leads to proactive facilitation of new learning, with interference only observed under specific conditions and immediate test and being thus an exception rather than a rule.
Strengths:
The study uses a well-established word-pair learning paradigm to study interference and facilitation of overlapping memories. However it goes more in-depth than a typical interference study in the systematic variation of several factors: (1) which elements of an association are overlapping and which are altered (change target, change cue, change both, change neither); (2) how much the changed element differs from the original (word relatedness, with two ranges of relatedness considered); (3) retention period (immediate test, 2-day delay). Furthermore, each experiment has a large N sample size, so both significant effects as well as null effects are robust and informative.
The results show the benefits of relatedness, but also replicate interference effects in the "change target" condition when the new target is not related to the old target and when the test is immediate. This provides a reconciliation of some existing seemingly contradictory results on the effect of overlap on memory. Here, the whole range of conditions is mapped to convincingly show how the direction of the effect can flip across the surface of relatedness values.
Additional strength comes from supporting analyses, such as analyses of learning data, demonstrating that relatedness leads to both better final memory and also faster initial learning.<br /> More broadly, the study informs our understanding of memory integration, demonstrating how the interdependence of memory for related information increases with relatedness. Together with a prior study or retroactive interference and facilitation, the results provide new insights into the role of reminding in memory formation.
In summary, this is a highly rigorous body of work that sets a great model for future studies and improves our understanding of memory organization.
Weaknesses:
The evidence for the proactive facilitation driven by relatedness is very convincing. However, in the finer scale results, the continuous relationship between the degree of relatedness and the degree of proactive facilitation/interference is less clear. This could be improved with some additional analyses and/or context and discussion. In the narrower range, the measure used was AS, with values ranging from 0.03-0.98, where even 0.03 still denotes clearly related words (pious - holy). Within this range from "related" to "related a lot", no relationship to the degree of facilitation was found. The wider range results are reported using a different scale, GloVe, with values from -0.14 to 0.95, where the lower end includes unrelated words (sap - laugh). It is possible that any results of facilitation/interference observed in the wider range may be better understood as a somewhat binary effect of relatedness (yes or no) rather than the degree of relatedness, given the results from the narrower condition. These two options could be more explicitly discussed. The report would benefit from providing clearer information about these measures and their range and how they relate to each other (e.g., not a linear transformation). It would be also helpful to know how the values reported on the AS scale would end up if expressed in the GloVe scale (and potentially vice-versa) and how that affects the results. Currently, it is difficult to assess whether the relationship between relatedness and memory is qualitative or quantitative. This is less of a problem with interdependence analyses where the results converge across a narrow and wider range.
A smaller weakness is generalizability beyond the word set used here. Using a carefully crafted stimulus set and repeating the same word pairings across participants and conditions was important for memorability calculations and some of the other analyses. However, highlighting the inherently noisy item-by-item results, especially in the Osgood-style surface figures, makes it challenging to imagine how the results would generalize to new stimuli, even within the same relatedness ranges as the current stimulus sets.
Tags
Annotators
URL
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In the manuscript "Metabolic heterogeneity of colorectal cancer as a prognostic factor: insights gained from fluorescence lifetime imaging" by Komarova et al., the authors used fluorescence lifetime imaging and quantitative analysis to assess the metabolic heterogeneity of colorectal cancer. Generally, this work is logically well-designed, including in vitro and in vivo animal models and ex vivo patient samples. However, since the key parameter presented in this study, the BI index, is already published in a previous paper by this group (Shirshin et al., 2022), and the quantification method of metabolic heterogeneity has already been well (and even better) described in previous studies (such as the one by Heaster et al., 2019), the novelty of this study is doubted. Moreover, I am afraid that the way of data analysis and presentation in this study is not well done, which will be mentioned in detail in the following sections.
Strengths:
(1) Solid experiments are performed and well-organized, including in vitro and in vivo animal models and ex vivo patient samples.
(2) Attempt and efforts to build the association between the metabolic heterogeneity and prognosis for colorectal cancer.
Weaknesses:
(1) The human sample number (from 21 patients) is very limited. I wonder how the limited patient number could lead to reliable diagnosis and prognosis;.
(2) The BI index or similar optical metrics have been well established by this and other groups; therefore, the novelty of this study is doubted.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The authors report the discovery of a population of gingival fibroblasts displaying the expression of cellular senescence markers P21 and P16 in human periodontitis samples and a murine ligature-induced periodontitis (LIP) model. They support this finding in the murine model through bulk RNA-sequencing and show that differentially expressed genes are significantly enriched in the SenMayo cellular senescence in an aging dataset. They then show that Ligature-Induced Periodontitis (LIP) mice treated with the senomorphic drug metformin display overall diminished bone damage, reduced histomorphic alterations, and a reduction in P21 and P16 immunostaining signal. To explore the cell types expressing cellular senescence markers in periodontitis, the authors make use of a combination of bioinformatic analyses on publicly available scRNA-seq data, immunostainings on patient samples and their LIP model; as well as in vitro culture of healthy human gingival fibroblasts treated with LPS. They found that fibroblasts are a cell population expressing P16 in periodontitis which are also enriched for SenMayo genes, suggesting they have a senescent phenotype. They then point to a subgroup of fibroblasts expressing CD81+ with the highest enrichment for a SASP geneset in periodontitis. They also show that treatment of LIP mice and human LPS-treated gingival fibroblasts with metformin leads to a reduction of P21 and P16-positive cells, as well as the senescence-associated beta-galactosidase (SA-beta-gal) marker. Finally, they show evidence suggesting that CD81+ senescent fibroblasts are the source of C3 complement protein, which they stipulate signals through the C3AR1 receptor present in neutrophils in periodontitis. The authors observed that both CD81+ fibroblast and C3AR1+ neutrophil populations are expanded in periodontitis, that both populations appear to be in close contact, and that treatment with metformin reduced both C3 and the neutrophil marker MPO in their mouse LIP model.
Strengths:
The study implements several different techniques and tools on human samples, mouse models, fibroblast cultures, and publicly available data to support their conclusions. In summary, the evidence suggests that in the context of periodontitis, there is an expansion of cells expressing senescence markers P21, and P16, as well as members of the SASP, and that this includes CD81+ fibroblasts.
Weaknesses:
The manuscript appears to use as synonyms the terms "senescent cells" and "aging cells", as well as "senescence" and "aging", or "accelerated senescence" and "accelerated aging". This choice of words makes it difficult to understand the objectives of the study and the interpretations the authors are deriving from their results. The current understanding of the role of cellular senescence is that it is only one of the multiple biological aspects that characterize physiological aging. Although deeply intertwined, aging and cellular senescence are widely considered distinct phenomena, but the difference between these concepts seems blurry to me within the manuscript.
After reading the manuscript, my understanding is that the authors are comparing the process of periodontitis to a form of accelerated aging, in which senescent cells are potential drivers or contributors. I believe this to be an interesting point of view. As the authors mention, periodontitis is more common in the elderly, and senescence is strongly implicated in aging. However, I am not entirely sure if the authors were trying to address such a question, and more importantly, the experiments conducted here cannot address the relationships between cellular senescence in periodontitis and aging as (1) they do not conduct an expanded analysis of molecular and cellular features of aging in the oral epithelium beyond cellular senescence, (2) they do not test this hypothesis in vitro and in vivo using models of accelerated or delayed aging (or publicly available datasets of such models), and (3) interpretations regarding the aging process are hindered by the fact that all human healthy patients were young adults, while all human periodontitis patients were middle-aged, while the mouse model did not include different age groups.
The authors also refer to metformin as an "anti-aging" drug. Therefore, to me, it is not clear if the authors intended to use metformin as a senotherapeutic agent to show a correlation between senescence markers and the severity of periodontitis, or if they conceived their experiments and interpreted their results as "delaying the aging process". The latter would be more difficult to determine as cellular senescence is only one of the several aspects of the aging process in tissues. As none of the other molecular and cellular hallmarks that characterize the process of aging (epigenetic alterations, telomere shortening, immunosenescence, mitochondrial dysfunction, stem cell depletion, genomic instability, loss of proteostasis, nutrient sensing disruption, etc.) were studied, I believe this might be just a matter of semantics and rephrasing.
On the other hand, and assuming the authors were only seeking to explore the role of cellular senescence in periodontitis (irrespective of the aging process), I have the following concerns:
Major concerns:
(1) A majority of the bioinformatic analyses regarding cellular senescence were conducted using only the SenMayo geneset reported by Dominik Saul et al. That geneset was developed by literature searching for genes associated with cellular senescence that had been studied in the context of human aging (in bone marrow). Thus, my understanding is that it is not an "aging" gene set as the authors describe it (and possibly interpret it) throughout the manuscript but a gene set of cellular senescence-associated genes that are overrepresented in aging tissues.
The SenMayo geneset specifically excludes important genes like P21, P16, and RELA as they were used for validating that dataset against other datasets. Additionally, most of the genes that comprise SenMayo are cytokines and growth factors. This includes part of the SASP (and the authors also show enrichment for some SASP factors using the Coppé dataset in Figure 5) but excludes many of the core important processes that are known to define cellular senescence, including cell cycle inhibition, lack of cell proliferation, accumulation of DNA damage, activation of the lysosomal compartment and disruption of the nuclear envelope, among others. As the SenMayo geneset was developed for studying senescence in the context of aging, I believe it is important to conduct a more extensive analysis with other published gene sets of cellular senescence. Examples include the cellular senescence and SASP REACTOME pathways, the KEGG cellular senescence pathway, the cellular senescence GO term, the Fridman dataset, SeneQuest, CSGene, CellAge, etc. Most importantly, it will be important to show the enrichment of pathways related to hallmark pathways underlying cellular senescence such as cell cycle inhibition, the DNA damage response and repair pathways, NF-kB signaling, MTOR, and autophagy signaling, etc. Showing the enrichment level of these pathways in the CD81+ fibroblasts in periodontitis would be of utmost importance for backing up the conclusions of this study.
(2) The most important aspect of the definition of cellular senescence is the absence of cell proliferation. Beyond the expression of the p21, p16, and SASP markers, any evidence showing that CD81+ fibroblasts are not proliferating in vivo in humans and mice, and in vitro in the case of LPS experiments, would be of great importance for defining these cells as senescent. Otherwise, conclusions should be toned down to refer to the expression of senescence markers or cells having a "senescent-like" phenotype.
(3) The use of a "relative optic density" metric instead of positive cell counts as a measure for quantifying IHC stainings might pose challenges in reproducing these results, especially for the P21 and P16 stainings which are proteins that despite being possibly also being found in the cytoplasm, should be clearly present in the nucleus of positive cells. The quantification of the levels of these markers is of great importance for the conclusions of this study but I am concerned they would be too difficult to reproduce. In my opinion, cell counts and % of positive cells should be used, with a clear description of the total number of cells counted in the methodology. Otherwise, a strong justification for using OD in the methodology is required in addition to considering the following comments:
a. There is no description in the methodology describing how this relative OD is measured and calculated. It is not clear if the data points shown in the graphs are biological replicates or OD means measured in different histological sections from the same sample.
b. The images of P16 and P21 stainings in Figures 2E and 2F do not appear to resemble the differences in OD between conditions shown in the graphs of Figures 2Gd and 2Ge.
c. The stainings shown for p16 in Figure 2E seem considerably different from those shown in Figure 1D. Additionally, the relative OD of P16 at 14D is around 0.08 in Figure 1D, while the mean for the control appears to be around 0.015 at 14D in Figure 2Gd. The use of OD as a measure is again worrying as this could be impacting interpretations: the difference between the ODs of LIP+MET (around 0.08) and LIP+ddH2O (around 0.015) is reported as significant but the difference in OD between LIP14D in Figure 1D (around 0.07) and LIP+ddH2o in Figure 2Gd (around 0.015) should not be significant as they are supposed to similar control conditions.
d. Irrespective of the measure used, the authors should state exact means and standard deviations, as well as exact P values, the statistical test used, and the number of biological replicates per group in parenthesis in the main text and figure legend.
(4) The conclusions derived from experiments with metformin in mice and cell cultures are not fully supported by the evidence.
First, metformin has multiple molecular targets, as well as multiple organ and tissue targets. The experiments presented in mice do not consider/evaluate the systemic effects of metformin nor local effects in other gingival cell types and this should be discussed.
As mentioned before, these experiments cannot be interpreted as testing metformin in the context of "anti-aging", as this study addresses cellular senescence in periodontitis. However, the results are still relevant as there is considerable evidence showing that metformin has senomorphic activity. In this regard, the authors could make use of a compound that has been more extensively characterized as a senolytic such as ABT-737, ABT-263 (Navitoclax), or the combination of Dasatinib + Quercetin, to show the effect of eliminating senescent cells in their LPS induction fibroblast model.
They could also show the effect of metformin on the activation of other hallmark senescence pathways such as (the NF-kB pathway or the DNA damage response) and in the expression of SASP factors they identified as overexpressed in CD81+ fibroblasts through their analysis against the SenMayo dataset (e.g., IL6, CXCL1, CXCL12). This could be done in their samples from metformin-treated mouse experiments and in their LPS induction fibroblast model.
(5) For the data produced on the authors' human samples, the difference in the age of patient groups is a significant confounding factor. This is because all their periodontitis patient samples came from middle-aged individuals (mean age above 50 years), while all healthy samples were obtained from young adults (mean age 25 years). The authors should justify this selection of age groups and justify why differences in the age of each experimental group could impact the validity of their results regarding the accumulation of senescent cells. Showing the level of P21 and P16 positive cell accumulation in samples from healthy patients from a similar age group (middle-aged) is of great importance to support the validity of their results in humans.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The authors frame the MS-spectrum-based prediction of antimicrobial resistance prediction as a drug recommendation task. Weis et al introduced the dataset this model is tested on and benchmark models which take as input a single species and are trained to predict resistance to a single drug. Instead here, a pair of drug and spectrum are fed to 2 neural network models to predict a resistance probability. In this manner, knowledge from different drugs and species can be shared through the model parameters. Three questions are asked: 1. what is the best way to encode the drugs? 2. does the dual NN outperform the single-spectrum drug?
Overall the paper is well-written and structured. It presents a novel framework for a relevant problem. The work would benefit from more work on evaluation.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In the paper, the authors use a cellular Potts model to investigate muscle regeneration. The model is an attempt to combine many contributors to muscle regeneration into one coherent framework. I believe the resulting model has the potential to be very useful in investigating the complex interplay of multiple actors contributing to muscle regeneration.
Strengths:
The manuscript identified relevant model parameters from a long list of biological studies. This collation of a large amount of literature into one framework has the potential to be very useful to other authors. The mathematical methods used for parameterization and validation are transparent.
Comments on revised version:
The authors have satisfactorily addressed my previous comments.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
The manuscript by Chen et al from the group of Helen Miranda aims to describe an improved neuromuscular junction (NMJ) model to study synaptic dysfunction in several cases of familial ALS. Overall, the system described in the paper appears as a valid platform to study disease phenotypes with exciting results showing specific effects of GDNF on non-SOD1 ALS patient lines. The strength of the paper lies in the use of myotubes, and motor neurons derived from the same donor. However, the current study: (1) lacks a clear comparison of the current system with numerous previously described systems; (2) is limited by the number of repeat experiments in the study and (3) has no description of the synaptic phenotype observed in the study. These major points are discussed in more detail below.
Major points:
(1) In the introduction the authors state (p. 4): "Finally, recent human NMJ models have been established from PSCs by differentiating these cells into both skeletal muscles and motor neurons in 2D and 3D formats. These previous systems present a remarkable advancement to the studies of human NMJs, however, they require long NMJ formation and maturation time (40 to 60 days), which, restricts their sensitivity and scalability [42]"
In fact, a number of studies have described various in-vitro NMJ systems, with the same timeframes for NMJ formation. For example, in studies by Osaki et al, 2018, Sci Adv; Bellmann et al, 2019, Biomat; Demestre et al, 2015, Stem Cell Res; Badu-Mensah et al, 2022, Biomat (this is just an exemplar selection of the papers); NMJ formation was observed as early as 14 d in culture, in line with or at least slightly longer than reported by Chen et al. With the exception of the study by Osaki et al, all co-culture systems cited above are 2D-based. The authors need to expand on this further or provide a quantitative assessment of why their system is better compared to previously published models.
(2) Further, when comparing their results with other work it is hard to claim how the current system is (p. 5) "more reproducible, and offers a 6-fold increase in scalability compared to previous models [40-43]". The authors need to expand on this further.
(3) Although mentioned, there were no examples of the modularity of the system, which of course would strengthen the paper and help to uncover ALS mechanisms of synaptic formation, for example by combining WT myotubes and fALS motor neurons (see point 4 below). The authors should show how they would adapt to 96 well plate format to showcase the scalability of the system. Based on their data on the efficacy of synaptic formation (60 per 0.7 cm2 area), is further miniaturization allowed?
(4) A lot of a-bungarotoxin staining corresponds to AChR clusters that do not seem to be associated with muscle and do not form normal rings of clustering (pretzel-like) associated with the NMJ in vivo. This is seen clearly in Figure 3B and Figure 5B. Figures 3B and 5B only show low-magnification images which makes it difficult to assess the specificity of localization of the pre-/post-synaptic markers. The authors should clearly show the morphologies of the NMJs formed in WT and fALS lines at high magnification. In addition, the authors should show co-localization images for a-bungarotoxin and myosin-heavy chains to confirm the localization of the bungarotoxin signal on the myotubes.
In addition to that, the authors report that the number of functional synapses formed on a plate varies from 30 (fASL) to 60 (Ctrl) per 10,000 neurons spread over the 0.7 cm2 area (0.6%). How do the authors explain an extensive loss of a-bungarotoxin signal in Figure 5B the majority of which likely corresponds to AChR clusters that are formed outside of neuronal connections? Such clustering can be usually observed in immature co-cultures and in vivo prior to the innervation of myotubes. One explanation could be that myotubes derived from fALS PSC are less capable of synaptic formation. Noteworthy, a study of PSC-derived myotubes and motor neurons from PSC lines with various SOD1 mutations has already been published, but not cited by Chen et al (Badu-Mensah et al). Given the importance of those confounding factors, the authors should test cell-intrinsic (motor neuron-related) vs non-cell-intrinsic mechanisms by co-culturing healthy myotubes with fALS-derived motor neurons followed by NMJ quantification.
(5) The authors present the advantage of optogenetic stimulation, but they only show the proof-of-principle and never really apply it to their studies. Specifically, with regard to Figure 6, are motor units derived from fALS PSCs incapable of being ontogenetically activated to the same extent as control motor units? Does the dysfunction stem from fALS motor neurons or fALS myotubes?
(6) Figures 6 B and C appear to be identical except for the addition of the GDNF effect on the fALS lines. This should all be put in one figure. The authors should also show whether GDNF-induced functional recovery is associated with recovery in the number of motor units or with merely synaptic function by quantifying the NMJ number in the presence of GDNF.
(7) Figure 5 and Figure 6. The authors only use one line per fALS mutation and their corresponding isogenic controls. They state that the n=6 for these experiments represents the technical replication of the experiment. These experiments should be performed at least n=3 times starting from neuronal differentiation, and not by seeding replicate wells representing a true replication of each experiment. This would significantly strengthen their argument that their method is robust and the results are easily reproducible.
(8) In the discussion the authors may want to mention that the lack of function of GDNF on the SOD1 lines may relate to the fact that SOD1 mutations do not lead to TDP43 pathology. Although speculative this suggests that in cases with TDP43 mutations (their data) or sporadic disease GDNF may be effective.
(9) Although beyond the scope of this paper, it would of course be interesting to see if sporadic forms of ALS had this same phenotype.
-
-
docdrop.org docdrop.org
-
second piece of advice is darkness
for - sleep hygiene - 2 - darkness
advice - sleep hygiene - 2 - darkness - one hour before sleeping, turn lights down to lowest level possible
-
regularity
for - sleep hygiene - 1 - regularity
advice - sleep hygiene - 1 - regularity - try to make weekend sleep times same as weekday - don't deviate if possible
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this contribution, the authors investigate the degree of alternative splicing across the evolutionary tree and identify a trend of increasing alternative splicing as you move from the base of the tree (here, only prokaryotes are considered) towards the tips of the tree. In particular, the authors investigate how the degree of alternative splicing (roughly speaking, the number of different proteins made from a single ORF (open reading frame) via alternative splicing) relates to three genomic variables: the genome size, the gene content (meaning the fraction of the genome composed of ORFs), and finally, the coding percentage of ORFs, meaning the ratio between exons and total DNA in the ORF. When correlating the degree of alternative splicing with these three variables, they find that the different taxonomic groups have a different correlation coefficient, and identify a "progressive pattern" among metazoan groups, namely that the correlation coefficient mostly increases when moving from flowering plants to arthropods, fish, birds, and finally mammals. They conclude that therefore the amount of splicing that is performed by an organismal group could be used as a measure of its complexity.
Weaknesses:
While I find the analysis of alternative splicing interesting, I also find that it is a very imperfect measure of organismal complexity and that the manuscript as a whole is filled with unsupported statements. First, I think it is clear to anyone studying evolution over the tree of life that it is the complexity of gene regulation that is at the origin of much of organismal structural and behavioral complexity. Arguably, creating different isoforms out of a single ORF is just one example of complex gene regulation. However, the complexity of gene regulation is barely mentioned by the authors. Further, it is clear that none of their correlation coefficients actually show a simple trend (see Table 3). According to these coefficients, birds are more complex than mammals for 3 out of 4 measures. It is also not clear why the correlation coefficient between alternative splicing ratio and genome length, gene content, and coding percentage should display such a trend, rather than the absolute value. There are only vague mechanistic arguments.
Much more troubling, however, is the statement that the data supports "lineage-specific trends" (lines 299-300). Either this is just an ambiguous formulation, or the authors claim that you can see trends *within* lineages. The latter is clearly not the case. In fact, within each lineage, there is a tremendous amount of variation, to such an extent that many of the coefficients given in Table 3 are close to meaningless. Note that no error bars or p-values are presented for the values shown in Table 3. Figure 2 shows the actual correlation, and the coefficient for flowering plants there is given as 0.151, with a p-value of 0.193. Table 3 seems to quote r=0.174 instead. It should be clear that a correlation within a lineage or species is not a sign of a trend.
There are several wrong or unsupported statements in the manuscript. Early on, the authors state that the alternative splicing ratio (a number greater or equal to one that can be roughly understood as the number of different isoforms per ORF) "quantifies the number of different isoforms that can be transcribed using the same amount of information" (lines 51-52). But in many cases, this is incorrect, because the same sequence can represent different amounts of information depending on the context. So, if a changed context gives rise to a different alternative splice, it is because the genetic sequence has a different meaning in the changed context: the information has changed. In line 149, the authors state that "the energetic cost of having large genomes is high". No citation is given, and while such a statement seems logical, it does not have very solid support. If there was indeed a strong selective force to reduce genome size, we would not see the stunning diversity of genome sizes even within lineages. This statement is repeated (without support) several times in the manuscript, apparently in support of the idea that mammals had "no choice" to increase complexity via alternative splicing because they can't increase it by having longer genomes. I don't think this reasoning can be supported. Even more problematic is the statement that "the amount of protein-coding DNA seems to be limited to a size of about 10MB" (line 219). There is no evidence whatsoever for this statement. The reference that is cited (Choi et al 2020) suggests that there is a maximum of 150GB in total genome size due to physiological constraints. In lines 257-258, the authors write that "plants are less restricted in terms of storing DNA sequences compared to animals" (without providing evidence or a citation). I believe this statement is made due to the observation that plants tend to have large intergenic regions. But without examining the functionality of these interagency regions (they might host long non-coding RNA stretches that are used to regulate the expression of other genes, for example) it is quite adventurous to use such a simple measure as being evidence that plants "are less restricted in terms of storing DNA sequences", whatever that even means. I do not think the authors mean that plants have better access to -80 freezers. The authors conclude that "plant's primary mechanism of genome evolution is by expanding their genome". This statement itself is empty: we know that plants are prone to whole genome duplication, but this duplication is not, as far as we understand, contributing to complexity. It is not a "primary mechanism of genome evolution". In lines 293-294, the authors claim that "alternative splicing is maximized in mammalian genomes". There is no evidence that this ratio cannot be increased. So, to conclude (on lines 302-303) that alternative splicing ratios are "a potential candidate to quantify organismal complexity" seems, based on this evidence, both far-fetched and weak at the same time.
I am also not very comfortable with the data analysis. The authors, for example, say that they have eliminated from their analysis a number of "outlier species". They mention one: Emmer wheat because it has a genome size of 900 Mb (line 367). Since 900MB does not appear to be extreme, perhaps the authors meant to write 900 Gb. When I consulted the paper that sequenced Triticum dicoccoides, they noted that 14 chromosomes are about 10GB. Even a tetraploid species would then not be near 900Gb. But more importantly, such a study needs to state precisely which species were left out, and what the criteria are for leaving out data, lest they be accused of selecting data to fit their hypothesis.
I understand that Methods are often put at the end of a manuscript, but the measures discussed here are so fundamental to the analysis that a brief description of what the different measures are (in particular, the "alternative splicing ratio") should be in the main text, even when the mathematical definition can remain in the Methods.
Finally, a few words on presentation. I understand that the following comments might read differently after the authors change their presentation. This manuscript was at the border of being comprehensible. In many cases, I could discern the meaning of words and sentences in contexts but sometimes even that failed (as an example above, about "species-specific trends", illustrates). The authors introduced jargon that does not have any meaning in the English language, and they do this over and over again.
Note that I completely agree with all the comments by the other reviewer, who alerted me to problems I did not catch, including the possible correlation with effective population size: a possible non-adaptive explanation for the results.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, the authors provide a method aiming to accurately reflect the individual deviation of longitudinal/temporal change compared to the normal temporal change characterized based on pre-trained population normative model (i.e., a Bayesian linear regression normative model), which was built based on cross-sectional data. This manuscript aims to solve a recently identified problem of using normative models based on cross-sectional data to make inferences about longitudinal change.
Although the proposed method was implemented with real data and shown to be more sensitive in capturing the differences confirmed by previous studies than conventional methods, there is still a lack of simulation studies to formally evaluate the performance of the proposed method in making accurate estimations and inferences about the longitudinal changes.
Strengths:
The efforts of this work make a good contribution to addressing an important question of normative modeling. With the greater availability of cross-sectional studies for normative modeling than longitudinal studies, and the inappropriateness of making inferences about longitudinal subject-specific changes using these cross-sectional data-based normative models, it's meaningful to try to address this gap from the perspective of methodological development.
Weaknesses:
• The organization and clarity of this manuscript need enhancement for better comprehension and flow. For example, in the first few paragraphs of the introduction, the wording is quite vague. A lot of information was scattered and repeated in the latter part of the introduction, and the actual challenges/motivation of this work were not introduced until the 5th paragraph.
• There are no simulation studies to evaluate whether the adjustment of the cross-sectional normative model to longitudinal data can make accurate estimations and inferences regarding the longitudinal changes. Also, there are some assumptions involved in the modeling procedure, for example, the deviation of a healthy control from the population over time is purely caused by noise and constant variability of error/noise across x_n, and these seem to be quite strong assumptions. The presentation of this work's method development would be strengthened if the authors can conduct a formal simulation study to evaluate the method's performance when such assumptions are violated, and, ideally, propose some methods to check these assumptions before performing the analyses.
• The proposed "z-diff score" still falls in the common form of z-score to describe the individual deviation from the population/reference level, but now is just specifically used to quantify the deviation of individual temporal change from the population level. The authors need to further highlight the difference between the "z-score" and "z-diff score", ideally at its first mention, in case readers get confused (I was confused at first until I reached the latter part of the manuscript). The z-score can also be called a measure of "standardized difference" which kind of collides with what "z-diff" implies by its name.
• Explaining that one component of the variance is related to the estimation of the model and the other is due to prediction would be helpful for non-statistical readers.
• It would be easier for the non-statistical reader if the authors consistently used precision or variance for all variance parameters. Probably variance would be more accessible.
• The functions psi were never explicitly described. This would be helpful to have in the supplement with a reference to that in the paper.
• What is the goal of equations (13) and (14)? The authors should clarify what the point of writing these equations is prior to showing the math. It seems like it is to obtain an estimate of \sigma_{\ksi}^2, which the reader only learns at the end.
• What is the definition of "adaption" as used to describe equation (15)? In this equation, I think norm on subsample was not defined.
• "(the sandwich part with A)" - maybe call this an inner product so that it is not confused with a sandwich variance estimator. This is a bit unclear. Equation (8) does have the inner product involving A and \beta^{-1} does include variability of \eta. It seems like you mean that equation (8) incorrectly includes variability of \eta and does not have the right term vector component of the inner product involving A, but this needs clarifying.
• One challenge with the z-diff score is that it does not account for whether a person sits above or below zero at the first time point. It might make it difficult to interpret the results, as the results for a particular pathology could change depending on what stage of the lifespan a person is in. I am not sure how the authors would address those challenges.
-
-
www.medrxiv.org www.medrxiv.org
-
Reviewer #2 (Public Review):
HIV infection is characterized by viral integration into permissive host cells - an event that occurs very early in viral-host encounter. This constitutes the HIV proviral reservoir and is a feature of HIV infection that provides the greatest challenge for eradicating HIV-1 infection once an individual is infected.
This study looks at how starting HIV treatment very early after infection, which substantially reduces the peak viral load detectable (compared to untreated infection), affects the amount and characteristics of the viral reservoir. The authors studied 35 women in South Africa who were at high risk of getting HIV. Some of these women started HIV treatment very soon after getting infected, while others started later. This study is well-designed and has as its focus a very well-characterized cohort. Comparison groups are appropriately selected to address reservoir characterization and dynamics in the context of acute and chronic treated HIV-1. The amount of HIV and various characteristics of the genetic makeup of the virus (intact/defective proviral reservoir) were evaluated over one year of treatment. Methods employed for reservoir characterization are state-of-the-art and provide in-depth insights into the reservoir in peripheral blood.
While starting treatment early didn't reduce the amount of HIV DNA at the outset, it did lead to a gradual decrease in total HIV DNA quantity over time. In contrast, those who started treatment later didn't see much change in this parameter. Starting treatment early led to a faster decrease in intact provirus (a measure of replication-competence), compared to starting treatment later. Additionally, early treatment reduced the genetic diversity of the viral DNA and resulted in fewer immune escape variants within intact genomes. This suggests that collectively having a smaller intact replication-competent reservoir, less viral variability, and less opportunity for the virus to evade the immune system - are all features that are likely to facilitate more effective clearance of viral reservoir, especially when combined with other intervention strategies.
Major strengths of the study include the cohort of very early treated persons with HIV and the depth of study. These are important findings, particularly as the study was conducted in HIV-1 subtype C infected women (more cure studies have focussed on men and with subtype B infection)- and in populations most affected by HIV and in need of HIV cure interventions. This is highly relevant because it cannot be assumed that any interventions employed for reducing/clearing the HIV reservoir would perform similarly in men and women or across different populations. Other factors also deserve consideration and include age, and environment (e.g. other comorbidities and coinfections).
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #3 (Public Review):
Summary:
Bartolome et al. present ProteasomeID, a novel method to identify components, interactors, and (potentially) substrates of the proteasome in cell lines and mouse models. As a major protein degradation machine that is highly conserved across eukaryotes, the proteasome has historically been assumed to be relatively homogeneous across biological scales (with few notable exceptions, e.g., immunoproteasomes and thymoproteasomes). However, a growing body of evidence suggests that there is some degree of heterogeneity in the composition of proteasomes across cell tissues, and can be highly dynamic in response to physiologic and pathologic stimuli. This work provides a methodological framework for investigating such sources of variation. The authors start by adapting the increasingly popular biotin ligation strategy for labelling proteins coming into close proximity with one of three different subunits of the proteasome, before proceeding with PSMA4 for further development and analysis based on their preliminary labelling data. In a series of well-constructed and convincing validation experiments, the authors go on to show that the tagged PSMA4 construct can be incorporated into functional proteasomes, and is able to label a broad set of known proteasome components and interacting proteins in HEK293T cells. They also attempt to identify novel proteasomal degradation substrates with ProteasomeID; while this was convincing for known substrates with particularly short half-lives, the results for substrates with longer half-lives were less clear. One of the most compelling results was from a similar experiment to confirm proteasomal degradation induced by a BRD-targeting PROTAC, which I think is likely to be of keen interest to the targeted degradation community. Finally, the authors establish a ProteasomeID mouse model, and demonstrate its utility across several tissues.
Strengths:
(1) ProteasomeID itself is an important step forward for researchers with an interest in protein turnover across biological scales (e.g., in sub-cellular compartments, in cells, in tissues, and whole organisms). I especially see interest from two communities: those studying fundamental proteostasis in physiological and pathologic processes (e.g., ageing; tissue-specific protein aggregation diseases), and those developing targeted protein degradation modalities (e.g., PROTACs; molecular glues). All the datasets generated and deposited here are likely to provide a rich resource to both. The HEK293T cell line data are a valuable proof-of-concept to allow expansion into more biologically-relevant cell culture settings; however, I envision the greatest innovation here to be the mouse model. For example, in the targeted protein degradation space, two major hurdles in early-stage pre-clinical development are (i) evaluation of degradation efficacy across disease-relevant tissues, and (ii) toxicity and safety implications caused by off-target degradation, e.g., of newly-identified molecular glues and/or in particularly-sensitive tissues. The ProteasomeID mouse allows early in vivo assessment of both these questions. The results of the BRD PROTAC experiment in 293T cells provides an excellent in vitro proof-of-concept for this approach.
(2) The mass-spectrometry-based proteomics workflows used and presented throughout the manuscript are robust, rigorous, and convincing. For example, the algorithm the authors use for defining enrichment score cut-offs are logical and based on rational models, rather than on arbitrary cut-offs that are common for similar proteomics studies. The construction (and subsequent validation) of both BirA*- and miniTurbo- tagged PSMA4 variants also increases the utility of the method, allowing researchers to choose the variant with the labelling time-scale required for their particular research question.
(3) The optimised BioID and TurboID protocol the authors develop (summarised in Fig. S2A) and validate (Fig. S2B-D) is likely to be of broad interest to cell and molecular biologists beyond the protein degradation field, given that proximity labelling is a current gold-standard in global protein:protein interaction profiling.
Limitations:
I think the authors do an excellent job in highlighting the limitations of ProteasomeID throughout the Results and Discussion. I do have some specific comments that might provide additional context for the reader.
(1) The authors do a good job in showing that a substantial proportion of PSMA4-BirA* is incorporated into functional proteasome particles; however, it is not immediately clear to me how much background (false-positive IDs) might be contributed by the ~40 % of PSMA4-BirA* that is not incorporated into the mature core particle (based on the BirA* SEC-MS traces in Fig. 2b and S3b, i.e., the large peak ~ fraction 20). Are there any bands lower down in the native gel shown in Fig. 2c, i.e., corresponding to lower molecular weight complexes or monomeric PSMA4-BirA*? The enrichment of proteasome assembly factors in all the ProteasomeID experiments might suggest the presence of assembly intermediates, which might themselves become substrates for proteasomal degradation (as has been shown for other incompletely-assembled protein complexes, e.g., the ribosome, TRiC/CCT).
(2) Although the authors attempt to show that BirA* tagging of PSMA4 does not interfere with proteasome activity (Fig. 2e-f), I think the experimental evidence for this is incomplete. They show that the overall chymotrypsin-like activity (attributable to PSMB5) in cells expressing PSMA4-BirA* is not markedly reduced compared with control BirA*-expressing cells. However, they do not show that the activity of the specific proteasome sub-population that contains PSMA4-BirA* is unaffected (e.g., by purifying this sub-population via the Flag tag). The proteasome activity of the sub-population of wild-type proteasome complexes that do not contain the PSMA4-BirA* (~50%, based on the earlier immunoblots) could account for the entire chymotrypsin-like activity-especially in the context of HEK293T cells, where steady-state proteasome levels are unlikely to be limiting. It would also be useful to assess any changes in tryspin- and caspase- like activities, especially as tagging of PSMA4 could conceivably interfere with the activity of some PSMB subunits, but not others.
(3) I was left slightly unsure as to the general utility of ProteasomeID for identifying novel proteasomal substrates in homeostatic conditions--especially for proteins with longer half-lives. The cycloheximide chases in Fig. 4g/S4j are clear for MYC and TIGD5 (which have short half-lives), but are not so clear for ARMC6 and BRAT1: the reduction in the bands are modest, and might have been clearer with longer "chase" time-points. Furthermore, classifying candidates based on enrichment following proteasome inhibition with MG-132 have the potential to lead to a high number of false positives. ProteasomeID's utility in identifying potential substrates in more targeted settings (e.g., molecular glues, off-target PROTAC substrates) is far more apparent.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this paper, the authors wanted to show that capsaicin can disrupt the interaction between Keap1 and Nrf2 by directly binding to Keap1 at an allosteric site. The resulting stabilization of Nrf2 would protect CAP-treated gastric cells from alcohol-induced redox stress and damage as well as inflammation (both in vitro and in vivo).
Strengths:
One major strength of the study is the use of multiple methods (CoIP, SPR, BLI, deuterium exchange MS, CETSA, MS simulations, target gene expression) that consistently show for the first time that capsaicin can disrupt the Nrf2/Keap1 interaction at an allosteric site and lead to stabilization and nuclear translocation of Nrf2.
Weaknesses:
One major weakness of the study is that plausibility is taken as proof for causality. The finding that capsaicin directly binds to Keap1 and releases Nrf2 from its fate of degradation (in vitro) is taken for granted as the sole explanation for the observed improved gastric health upon alcohol exposure (in vivo). There is no consideration or exclusion of any potential unrelated off-target effect of capsaicin, or proteins other than Nrf2 that are also controlled by Keap1.
Another point that hampers full appreciation of the capsaicin effect in cells is that capsaicin is not investigated alone, but mostly in combination with alcohol only.
Bottom Line:
Overall, the authors are convincing that capsaicin (although weakly) binds to Keap1 and releases Nrf2 from degradation. With this, the authors provide a significant finding with marked relevance for the redox/Nrf2 as well as natural products /hit discovery communities. Moreover, the employed toolbox of different complementary methodologies can set the bar for future PPI inhibitor studies. The translation of this novel finding in a biological setting (alcohol-stressed gastric cells) still leaves some open questions and concerns. These concerns mainly arise from lacking control experiments and/or somewhat biased conclusions from the obtained data sets.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
The paper addresses several factors that influence daily changes in concentration of metabolites in the Drosophila melanogaster gut. The authors describe metabolomes extracted from fly guts at four time-points during a 24-hour period, comparing profiles of primary metabolites, lipids, and biogenic amines. The study elucidates that the percentage of metabolites that exhibit a circadian cycle, peak phases of their increased appearance, and the cycling amplitude depends on the combination of factors (microbiome status, composition or timing of the diet, circadian clock genotype). Multiple general conclusions based on the data obtained with modern metabolomics techniques are provided in each part of the article. Descriptive analysis of the data supports the finding that microbiome increases the number of metabolites for which concentration oscillates during the day period. Results of the experiments show that timed feeding significantly enhanced metabolite cycling and changed its phase regardless of the presence of a microbiome. The authors suggest that the host circadian rhythm modifies both metabolite cycling period and the number of such metabolites.
Strengths:
The obvious strength of the study is the data on circadian cycling of the detected 843, 4510, and 4330 total primary metabolites, lipids, and biogenic amines respectively in iso31 flies and 623, 2245, and 2791 respective metabolites in per01 mutants. The comparison of the abundance of these metabolites, their cycling phase, and the ratio of cycling/non-cycling metabolites is well described and illustrated. The conditions tested represent significant experimental interest for contemporary chronobiology: with/without microbiota, wild-type/mutant period gene, ad libitum/time-restricted feeding, and high-sugar/high-protein diet. The authors conclude that the complex interaction between these factors exists, and some metabolic implications of combinations of these factors can be perceived as reminiscent of metabolic implications of another combination ("...the microbiome and time-restricted feeding paradigms can compensate for each other, suggesting that different strategies can be leveraged to serve organismal health"). Enrichment analysis of cycling metabolites leads to an interesting suggestion that oscillation of metabolites related to amino acids is promoted by the absence of microbiota, alteration of circadian clock, and time-restricted feeding. In contrast, association with microbiota induces oscillation of alpha-linolenic acid-related metabolites. These results provide the initial step for hypothesising about functional explanations of the uncovered observations.
Weaknesses:
Among the weaknesses of the study, one might point out too generalist interpretations of the results, which propose hypothetical conclusions without their mechanistic proof. The quantitative indices analysed are obviously of particular interest, however are not self-explaining and exhaustive. More specific biological examples would add valuable insights into the results of this study, making conclusions clearer. More specific comments on the weaknesses are listed below:
(1) The criterion of the percentage of cycling metabolites used for comparisons has its own limitations. It is not clear, whether the cycling metabolites are the same in the guts with/without microbiota, or whether there are totally different groups of metabolites that cycle in each condition. GO enrichment analysis gives only a partial assessment, but is still not quantitative enough.
(2) The period of cycling data is based on only 4 time points during 24 hours in 4 replicates (>200 guts per replicate) on the fifth day of the experiment (10-12-day-old adults). It does not convincingly prove that these metabolites cycle the following days or more finely within the day. Moreover, it is not clear how peaks in polar histogram plots were detected in between the timepoints of ZT0, ZT6, ZT12, and ZT18.
(3) Average expression and amplitude are analysed for groups of many metabolites, whereas the data on distinct metabolites is hidden behind these general comparisons. This kind of loss of information can be misleading, making interpretation of the mentioned parameters quite complicated for authors and their readers. Probably more particular datasets can be extracted to be discussed more thoroughly, rather than those general descriptions.
(4) The metabolites' preservation is crucial for this type of analysis, and both proper sampling plus normalisation require more attention. More details about measures taken to avoid different degradation rates, different sizes of intestines, and different amounts of microbes inside them will be beneficial for data interpretation.
(5) The data in the article describes formal phenomena, not directly connected with organism physiology. The parameters discussed obviously depend on the behaviour of flies. Food consumption, sleep, and locomotor activity could be additionally taken into account.
(6) Division of metabolites into three classes limits functional discussion of found differences. Since the enrichment analysis provided insights into groups of metabolites of particular interest (for example, amino acid metabolism), a comparison of their cycling characteristics can be shown separately and discussed.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In this manuscript, Wang and co-workets employ single molecule light microscopy (SMLM) to detect Nipah virus Fusion protein (NiV-F) in the surface of cells. They corroborate that these glycoproteins form microclusters (previously seen and characterized together with the NiV-G and Nipah Matrix protein by Liu and co-workers (2018) also with super-resolution light microscopy). Also seen by Liu and coworkers the authors show that the level of expression of NiV-F does not alter the identity of these microclusters nor endosomal cleavage. Moreover, mutations and the transmembrane domain or the hexamer-of-trimer interface seem to have a mild effect on the size of the clusters that the authors quantified. Importantly, it has also been shown that these particles tend to cluster in Nipah VLPs.
Strengths:
The authors have tried to perform SMLM in single VLPs and have shown partially the importance of NiV-F clustering.
Weaknesses:
The labelling strategy for the NiV-F is not sufficiently explained. The use of a FLAG tag in the extracellular domain should be validated and compared with the unlabelled WT NiV-F when expressed in functional pseudoviruses (for example HIV-1 based particles decorated with NiV-F). This experiment should also be carried out for both infection and fusion (including BlaM-Vpr as a readout for fusion). I would also suggest to run a time-of-addition BlaM experiment to understand how this particular labelling strategy affects single virion fusion as compared to the the WT. It would also be very important to compare the FLAG labelling approach with recent advances in the field (for instance incorporating noncanonical amino acids (ncAAs) into NiV-F by amber stop-codon suppression, followed by click chemistry).
The correlation between the existence of microclusters of a particular size and their functionality is missing. Only cell-cell fusion assays are shown in supplementary figures and clearly, single virus entry and fusion cannot be compared with the biophysics of cell-cell fusion. Not only the environment is completely different, membrane curvature and the number of NiV-F drastically varies also. Therefore, specific fusion assays (either single virus tracking and/or time-of-addition BlaM kinetics with functional pseudoviruses) are needed to substantiate this claim.
The authors also claim they could not characterize the number of NiV-F particles per cluster. Another technique such as number and brightness (Digman et al., 2008) could support current SMLM data and identify the number of single molecules per cluster. Also, this technology does not require complex microscopy apparatus. I suggest they perform either confocal fluorescence fluctuation spectroscopy or TIRF-based nandb to validate the clusters and identify how many molecule are present in these clusters. Also, it is not clear how many cells the authors employ for their statistics (at least 30-50 cells should be employed and not consider the number of events blinking events). I hope the authors are not considering only a single cell to run their stats... The differences between the mutants and the NiV-F is minor even if their statistical analyses give a difference (they should average the number and size of the clusters per cell for a total of 30-50 cells with experiments performed at least in three different cells following the same protocol). They should also compare the level of expression (with the number of molecules per cell provided by number and brightness) with the total number of clusters. Overall, it seems that the authors have only evaluated a very low number of cells.
The same applies to the VLP assay. I assume the authors have only taken VLPs expressing both NiV-M and NiV-F (and NiV-G). But even if this is not clearly stated I would urge the authors to show how many viruses were compared per condition (normally I would expect 300 particles per condition coming from three independent experiments). As a negative control to evaluate the cluster effect I would mix the different conditions. Clearly you have clusters with all conditions and the differences in clustering depending on each condition are minimal. Therefore you need to increase the n for all experiments.
-
-
www.researchsquare.com www.researchsquare.com
-
Reviewer #2 (Public Review):
Summary:
Environmental influences on development are ubiquitous, affecting many phenotypes in organisms. However molecular genetic and cellular mechanisms transducing environmental signals are still only barely understood. This study examines part of one such intracellular mechanism in a polyphenic (or dimorphic) aphid.
Strengths:
While other published reports have linked phenotypic plasticity to RNA editing before, this study reports such an interaction in insects. The study uses a wide array of molecular tools to identify connections upstream and downstream of the RNA editing to elucidate the regulatory mechanism, which is illuminating.
Weaknesses:
While this system is intriguing, this report does not foster confidence in its conclusions. Many of the analyses seem based on very small sample sizes. It is itself problematic that sample sizes are not obvious in most figures, although based on Methods section covering RNAseq, they seem to be either 3, 6 or 9, depending on whether stages were pooled, but that point is not made clear. With such small sample sizes, statistical tests of any kind are unreliable. Besides the ambiguity on sample sizes, it's unclear what error bars or whiskers show in plots throughout this study. When sample sizes are small estimates of variance are not reliable. Student's t-test is not appropriate for comparisons with such small sample sizes. Presently, it is not possible to replicate the tests shown in Figures 3, 4 and 6. (Besides the HT-seq reads, other data should also be made publicly available, following the journal's recommendations.) Regardless, effect sizes in some comparisons (Fig 3J, 4A-C, 6E,H) are clearly not large, making confidence in conclusions low. The authors should be cautious about over-interpreting these data.
Tags
Annotators
URL
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
Summary:
In their manuscript, Zhao et al. describe a link between JAK-STAT pathway activation in nephrocytes on a high-fat diet. Nephrocytes are the homologs to mammalian podocytes and it has been previously shown, that metabolic syndrome and obesity are associated with worse outcomes for chronic kidney disease. A study from 2021 (Lubojemska et al.) could already confirm a severe nephrocyte phenotype upon feeding Drosophila a high-fat diet and also linking lipid overflow by expressing adipose triglyceride lipase in the fat body to nephrocyte dysfunction. In this study, the authors identified a second pathway and mechanism, how lipid dysregulation impact on nephrocyte function. In detail, they show activation of JAK-STAT signaling in nephrocytes upon feeding them a high-fat diet, which was induced by Upd2 expression (a leptin-like hormone) in the fat body, and the adipose tissue in Drosophila. Further, they could show genetic and pharmacological interventions can reduce JAK-STAT activation and thereby prevent the nephrocyte phenotype in the high-fat diet model.
Strengths:
The strength of this study is the combination of genetic tools and pharmacological intervention to confirm a mechanistic link between the fat body/adipose tissue and nephrocytes. Inter-organ communication is crucial in the development of several diseases, but the underlying mechanisms are only poorly understood. Using Drosophila, it is possible to investigate several players of one pathway, here JAK-STAT. This was done, by investigating the functional role of Hop, Socs36E, and Stat92E in nephrocytes and has also been combined with feeding a high-fat diet, to assess restoration of nephrocyte function by inhibiting JAK-STAT signaling. Adding a translational approach was done by inhibiting JAK-STAT signaling with methotrexate, which also resulted in attenuated nephrocyte dysfunction. Expression of the leptin-like hormone upd2 in the fat body is a good approach to studying inter-organ communication and the impact of other organs/tissue on nephrocyte function and expands their findings from nephrocyte function towards whole animal physiology.
Weaknesses:
Although the general findings of this study are of great interest, there are some weaknesses in the study, which should be addressed. Overall, the number of flies investigated for the majority of the experiments is very low (6 flies) and it is not clear whether the flies used, are from independent experiments to exclude problems with food/diet. For the analysis, the mean values of flies should be calculated, as one fly can be considered a biological replicate, but not all individual cells. By increasing the number of flies investigated, statistical analysis will become more solid. In addition, the morphological assessment is rather preliminary, by only using a Pyd antibody. Duf or Sns should be visualized as well, also the investigation of the different transgenic fly strains studying the importance of JAK-STAT signaling in nephrocytes needs to include a morphological assessment. Moreover, the expected effect of feeding a high-fat diet on nephrocytes needs to be shown (e.g. by lipid droplet formation) and whether upd2 is actually increased here should also be assessed. The time points of assessment vary between 1, 3, and 7 days and should be consistent throughout the study or the authors should describe why they use different time points.
-
-
-
Reviewer #2 (Public Review):
This work deals with a very difficult physical problem: relating the assembly of building blocks on a molecular scale to the appearance of large, macroscopic assemblies. This problem is particularly difficult to treat, because of the large number of units involved, and of the complex way in which these units-monomers-interact with each other and with the solvent. In order to make the problem treatable, the authors recur to a number of approximations: Among these, there is the assumption that the system is spatially homogeneous, i.e., its features are the same in all regions of space. In particular, the homogeneity assumption may not hold in biologically relevant systems such as cells, where the behavior close to the cell membrane may strongly differ from the one in the bulk. As a result, this hypothesis calls for a cautious consideration and interpretation of the results of this work. Another notable simplification introduced by the authors is the assumption that the system can only follow two possible behaviors: In the first, each monomer interacts equally with the solvent; no matter the size of the cluster of which it is part. In the second case, monomers in the bulk of a cluster and monomers at the assembly boundary interact with the solvent in a different way. These two cases are considered not only because they simplify the problem, but also because they are inspired by biologically relevant proteins.
With these simplifications, the authors trace the phase diagram of the system, characterizing its phases for different fractions of the volume occupied by the monomers and solvent, and for different values of the temperature. The results qualitatively reproduce some features observed in recent experiments, such as an anomalous distribution of cluster sizes below the system saturation threshold, and the gelation of condensed phases above such threshold.
-
-
www.biorxiv.org www.biorxiv.org
-
Reviewer #2 (Public Review):
This concise and focused study by Lowry and colleagues identifies a motif in the pores of three families of channel/scramblase proteins that regulate exclusive ion permeation and lipid transport. These three ion channel families, which include the TMEM16s, the plant-expressed and stress-gated cation channel OSCA, and the mammalian homolog and mechanosensitive cation channel, TMEM63 share low sequence similarity between them and have seemingly differing functions, as anion (TMEM16s), or stress-activated cation channels (OSCA/TMEM63). The study finds that in all three families, mutating a single hydrophobic residue in the ion permeation pathway of the channels confers lipid transport through the pores of the channels, indicating that TMEM16 and the related OSCA and TMEM63 channels have a conserved potential for both ion and lipid permeation. The authors interpret the findings as revealing that these channel/scramblase proteins have a relatively low "energetic barrier for scramblase" activity. The experiments themselves seem to be done with a high level of rigor and the paper is well written. A weakness is the limited scope of the experiments which, if fixed, could open up a new line of inquiry.
-